| Revision as of 00:06, 20 June 2010 view sourceBubba73 (talk | contribs)Autopatrolled, Extended confirmed users, Pending changes reviewers, Rollbackers93,168 editsm →History of DNA research: link to Gosling in text← Previous edit | Latest revision as of 18:18, 22 December 2024 view source Wcamp9 (talk | contribs)Extended confirmed users1,419 editsNo edit summaryTag: Visual edit: Switched | ||
| Line 1: | Line 1: | ||
| {{Short description|Molecule that carries genetic information}} | |||
| {{For|a non-technical introduction to the topic|Introduction to genetics}} | |||
| {{pp-vandalism|small=yes}} | |||
| {{Other uses}} | |||
| {{pp-move}} | |||
| {{For-multi|a non-technical introduction to the topic|Introduction to genetics|other uses}} | |||
| {{Featured article}} | |||
| {{Use dmy dates|date=March 2022}} | |||
| {{Chromosome}} | |||
| ] (type ]). The ]s in the structure are colour-coded by ] and the detailed structures of two ]s are shown in the bottom right.]]] | |||
| {{Genetics sidebar}} | |||
| '''Deoxyribonucleic acid''' ({{IPAc-en|audio=en-us-Deoxyribonucleic acid.ogg|d|iː|ˈ|ɒ|k|s|ᵻ|ˌ|r|aɪ|b|oʊ|nj|uː|ˌ|k|l|iː|ᵻ|k|,_|-|ˌ|k|l|eɪ|-}};<ref>{{MerriamWebsterDictionary|deoxyribonucleic acid}}</ref> '''DNA''') is a ] composed of two ] chains that coil around each other to form a ]. The polymer carries ] instructions for the development, functioning, growth and ] of all known ]s and many ]es. DNA and ] (RNA) are ]s. Alongside ]s, ] and complex carbohydrates (]s), nucleic acids are one of the four major types of ]s that are essential for all known forms of ]. | |||
| ]]] | |||
| The two DNA strands are known as polynucleotides as they are composed of simpler ]ic units called ]s.<ref>{{Cite book |vauthors= Alberts B, Johnson A, Lewis J, Raff M, Roberts K, Walter P |title= Molecular Biology of the Cell |edition= 6th |publisher= Garland |year= 2014 |url= http://www.garlandscience.com/product/isbn/9780815344322 |page= Chapter 4: DNA, Chromosomes and Genomes |isbn= 978-0-8153-4432-2 |url-status=live |archive-url= https://web.archive.org/web/20140714210549/http://www.garlandscience.com/product/isbn/9780815344322 |archive-date= 14 July 2014 |df= dmy-all }}</ref><ref>{{cite web | vauthors = Purcell A |title=DNA |url=http://basicbiology.net/micro/genetics/dna|website=Basic Biology |url-status=live |archive-url=https://web.archive.org/web/20170105045651/http://basicbiology.net/micro/genetics/dna/ |archive-date=5 January 2017}}</ref> Each nucleotide is composed of one of four ] ]s (] , ] , ] or ] ), a ] called ], and a ]. The nucleotides are joined to one another in a chain by ]s (known as the ]) between the sugar of one nucleotide and the phosphate of the next, resulting in an alternating ]. The nitrogenous bases of the two separate polynucleotide strands are bound together, according to ]ing rules (A with T and C with G), with ]s to make double-stranded DNA. The complementary nitrogenous bases are divided into two groups, the single-ringed ]s and the double-ringed ]s. In DNA, the pyrimidines are thymine and cytosine; the purines are adenine and guanine. | |||
| {{pp-move-indef}} | |||
| {{pp-semi-vandalism|small=yes}} | |||
| Both strands of double-stranded DNA store the same ]. This information is ] when the two strands separate. A large part of DNA (more than 98% for humans) is ], meaning that these sections do not serve as patterns for ]. The two strands of DNA run in opposite directions to each other and are thus ]. Attached to each sugar is one of four types of nucleobases (or ''bases''). It is the ] of these four nucleobases along the backbone that encodes genetic information. ] strands are created using DNA strands as a template in a process called ], where DNA bases are exchanged for their corresponding bases except in the case of thymine (T), for which RNA substitutes ] (U).<ref>{{Cite web|url=https://www.genome.gov/genetics-glossary/Uracil|title=Uracil|website=Genome.gov|language=en|access-date=21 November 2019}}</ref> Under the ], these RNA strands specify the sequence of ]s within proteins in a process called ]. | |||
| '''Deoxyribonucleic acid''' ({{Audio-IPA|en-us-Deoxyribonucleic_acid.ogg|/diːˌɒksɨˌraɪbɵ.n(j)uːˈkleɪ.ɪk ˈæsɪd/}}) ('''DNA''') is a ] that contains the ] instructions used in the development and functioning of all known living ]s and some ]es. The main role of DNA ]s is the long-term storage of ]. DNA is often compared to a set of ]s or a recipe, or a ], since it contains the instructions needed to construct other components of ], such as ]s and ] molecules. The DNA segments that carry this genetic information are called ]s, but other DNA sequences have structural purposes, or are involved in regulating the use of this genetic information. | |||
| Within eukaryotic cells, DNA is organized into long structures called ]s. Before typical ], these chromosomes are duplicated in the process of DNA replication, providing a complete set of chromosomes for each daughter cell. ] (]s, ]s, ] and ]s) store most of their DNA inside the ] as ], and some in the ] as ] or in ]s as ].<ref>{{cite book | vauthors = Russell P | title= iGenetics |url= https://archive.org/details/igenetics0000russ_v6o1 |url-access= registration |publisher= Benjamin Cummings |location= New York |year= 2001 |isbn= 0-8053-4553-1}}</ref> In contrast, ]s (] and ]) store their DNA only in the ], in ]s. Within eukaryotic chromosomes, ] proteins, such as ]s, compact and organize DNA. These compacting structures guide the interactions between DNA and other proteins, helping control which parts of the DNA are transcribed. | |||
| Chemically, DNA consists of two long ] of simple units called ]s, with ]s made of ]s and ] groups joined by ] bonds. These two strands run in opposite directions to each other and are therefore ]. Attached to each sugar is one of four types of molecules called ]. It is the sequence of these four bases along the backbone that encodes information. This information is read using the ], which specifies the sequence of the ]s within proteins. The code is read by copying stretches of DNA into the related nucleic acid RNA, in a process called ]. | |||
| {{TOC limit|3}} | |||
| Within cells, DNA is organized into long structures called ]s. These chromosomes are duplicated before cells ], in a process called ]. ] (]s, ]s, ], and ]s) store most of their DNA inside the ] and some of their DNA in ]s, such as ] or ].<ref>{{cite book | last = Russell | first = Peter | title = iGenetics | publisher = Benjamin Cummings | location = New York | year = 2001 | isbn = 0-805-34553-1 }}</ref> In contrast, ]s (] and ]) store their DNA only in the ]. Within the chromosomes, ] proteins such as ]s compact and organize DNA. These compact structures guide the interactions between DNA and other proteins, helping control which parts of the DNA are transcribed. | |||
| ==Properties== | == Properties == | ||
| ]s shown as dotted lines.]] | ]s shown as dotted lines. Each end of the double helix has an exposed ] phosphate on one strand and an exposed ] hydroxyl group (—OH) on the other.]] | ||
| DNA is a long ] made from repeating units called ]s.<ref>{{cite book | |
DNA is a long ] made from repeating units called ]s.<ref>{{cite book | vauthors = Saenger W |title= Principles of Nucleic Acid Structure |publisher= Springer-Verlag |location= New York |year= 1984 |isbn= 0-387-90762-9}}</ref><ref name="Alberts">{{cite book | vauthors = Alberts B, Johnson A, Lewis J, Raff M, Roberts K, Peter W | title = Molecular Biology of the Cell | edition = Fourth | publisher = Garland Science | year = 2002 | location = New York and London | isbn = 0-8153-3218-1 | oclc = 145080076 | url = https://www.ncbi.nlm.nih.gov/books/NBK21054/ | url-status=live | archive-url = https://web.archive.org/web/20161101022040/https://www.ncbi.nlm.nih.gov/books/NBK21054/ | archive-date = 1 November 2016 | df = dmy-all }}</ref> The structure of DNA is dynamic along its length, being capable of coiling into tight loops and other shapes.<ref>{{cite journal | vauthors = Irobalieva RN, Fogg JM, Catanese DJ, Catanese DJ, Sutthibutpong T, Chen M, Barker AK, Ludtke SJ, Harris SA, Schmid MF, Chiu W, Zechiedrich L | title = Structural diversity of supercoiled DNA | journal = Nature Communications | volume = 6 | pages = 8440 | date = October 2015 | pmid = 26455586 | pmc = 4608029 | doi = 10.1038/ncomms9440 | bibcode = 2015NatCo...6.8440I |issn=2041-1723 }}</ref> In all species it is composed of two helical chains, bound to each other by ]. Both chains are coiled around the same axis, and have the same ] of {{convert|34|Å|nm|lk=on}}. The pair of chains have a radius of {{cvt|10|Å|nm}}.<ref name="Watson-1953">{{cite journal | vauthors = Watson JD, Crick FH | title = Molecular structure of nucleic acids; a structure for deoxyribose nucleic acid | journal = Nature | volume = 171 | issue = 4356 | pages = 737–38 | date = April 1953 | pmid = 13054692 | doi = 10.1038/171737a0 | url = http://www.nature.com/nature/dna50/watsoncrick.pdf | bibcode = 1953Natur.171..737W | s2cid = 4253007 | url-status=live | archive-url = https://web.archive.org/web/20070204110320/http://www.nature.com/nature/dna50/watsoncrick.pdf | archive-date = 4 February 2007 | df = dmy-all |issn=0028-0836 }}</ref> According to another study, when measured in a different solution, the DNA chain measured {{cvt|22|-|26|Å|nm}} wide, and one nucleotide unit measured {{cvt|3.3|Å|nm}} long.<ref>{{cite journal | vauthors = Mandelkern M, Elias JG, Eden D, Crothers DM | title = The dimensions of DNA in solution | journal = Journal of Molecular Biology | volume = 152 | issue = 1 | pages = 153–61 | date = October 1981 | pmid = 7338906 | doi = 10.1016/0022-2836(81)90099-1|issn=0022-2836 }}</ref> The buoyant density of most DNA is 1.7g/cm<sup>3</sup>.<ref>{{cite journal |last1=Arrighi |first1=Frances E. |last2=Mandel |first2=Manley |last3=Bergendahl |first3=Janet |last4=Hsu |first4=T. C. |title=Buoyant densities of DNA of mammals |journal=Biochemical Genetics |date=June 1970 |volume=4 |issue=3 |pages=367–376 |doi=10.1007/BF00485753|pmid=4991030 |s2cid=27950750 |issn=0006-2928 }}</ref> | ||
| DNA does not usually exist as a single strand, but instead as a pair of strands that are held tightly together.<ref name="Watson-1953" /><ref name=berg>{{cite book | vauthors = Berg J, Tymoczko J, Stryer L | date = 2002 | title = Biochemistry | publisher = W.H. Freeman and Company | isbn = 0-7167-4955-6 }}</ref> These two long strands coil around each other, in the shape of a ]. The nucleotide contains both a segment of the ] of the molecule (which holds the chain together) and a ] (which interacts with the other DNA strand in the helix). A nucleobase linked to a sugar is called a ], and a base linked to a sugar and to one or more phosphate groups is called a ]. A ] comprising multiple linked nucleotides (as in DNA) is called a ].<ref name="IUPAC">{{cite journal | author = IUPAC-IUB Commission on Biochemical Nomenclature (CBN) | title = Abbreviations and Symbols for Nucleic Acids, Polynucleotides and their Constituents. Recommendations 1970 | journal = The Biochemical Journal | volume = 120 | issue = 3 | pages = 449–54 | date = December 1970 | pmid = 5499957 | pmc = 1179624 | doi = 10.1042/bj1200449 | url = http://www.chem.qmul.ac.uk/iupac/misc/naabb.html | archive-url = https://web.archive.org/web/20070205191106/http://www.chem.qmul.ac.uk/iupac/misc/naabb.html | url-status=dead | archive-date = 5 February 2007 |issn=0306-3283 }}</ref> | |||
| The backbone of the DNA strand is made from alternating ] and ] |
The backbone of the DNA strand is made from alternating ] and ] groups.<ref name=Ghosh>{{cite journal | vauthors = Ghosh A, Bansal M | title = A glossary of DNA structures from A to Z | journal = Acta Crystallographica Section D | volume = 59 | issue = Pt 4 | pages = 620–26 | date = April 2003 | pmid = 12657780 | doi = 10.1107/S0907444903003251| bibcode = 2003AcCrD..59..620G |issn=0907-4449 }}</ref> The sugar in DNA is ], which is a ] (five-]) sugar. The sugars are joined by phosphate groups that form ]s between the third and fifth carbon ]s of adjacent sugar rings. These are known as the ] (three prime end), and ] (five prime end) carbons, the prime symbol being used to distinguish these carbon atoms from those of the base to which the deoxyribose forms a ].<ref name="berg" /> | ||
| Therefore, any DNA strand normally has one end at which there is a phosphate group attached to the 5′ carbon of a ribose (the 5′ phosphoryl) and another end at which there is a free hydroxyl group attached to the 3′ carbon of a ribose (the 3′ hydroxyl). The orientation of the 3′ and 5′ carbons along the sugar-phosphate backbone confers ] (sometimes called polarity) to each DNA strand. In a ], the direction of the nucleotides in one strand is opposite to their direction in the other strand: the strands are ]. The asymmetric ends of DNA strands are said to have a directionality of five prime end (5′ ), and three prime end (3′), with the 5′ end having a terminal phosphate group and the 3′ end a terminal hydroxyl group. One major difference between DNA and ] is the sugar, with the 2-deoxyribose in DNA being replaced by the related pentose sugar ] in RNA.<ref name="berg" /> | |||
| </ref> Animated version at ].]] | |||
| ]).]] | |||
| The DNA double helix is stabilized by ]s between the bases attached to the two strands. The four bases found in DNA are ] (abbreviated A), ] (C), ] (G) and ] (T). These four bases are attached to the sugar/phosphate to form the complete nucleotide, as shown for ]. | |||
| The DNA double helix is stabilized primarily by two forces: ]s between nucleotides and ] interactions among ] nucleobases.<ref name="Yakovchuk2006">{{cite journal | vauthors = Yakovchuk P, Protozanova E, Frank-Kamenetskii MD | title = Base-stacking and base-pairing contributions into thermal stability of the DNA double helix | journal = Nucleic Acids Research | volume = 34 | issue = 2 | pages = 564–74 | year = 2006 | pmid = 16449200 | pmc = 1360284 | doi = 10.1093/nar/gkj454 |issn=0305-1048 }}</ref> The four bases found in DNA are ] ({{mono|A}}), ] ({{mono|C}}), ] ({{mono|G}}) and ] ({{mono|T}}). These four bases are attached to the sugar-phosphate to form the complete nucleotide, as shown for ]. Adenine pairs with thymine and guanine pairs with cytosine, forming {{mono|A-T}} and {{mono|G-C}} ]s.<ref>{{cite book | vauthors = Tropp BE | title = Molecular Biology | edition = 4th | year = 2012 | publisher = Jones and Barlett Learning | location = Sudbury, Mass. | isbn = 978-0-7637-8663-2 }}</ref><ref>{{cite web | url = https://www.mun.ca/biology/scarr/Watson-Crick_Model.html | title = Watson-Crick Structure of DNA | year = 1953 | vauthors = Carr S | publisher = Memorial University of Newfoundland | access-date=13 July 2016 | url-status=live | archive-url = https://web.archive.org/web/20160719095721/http://www.mun.ca/biology/scarr/Watson-Crick_Model.html | archive-date = 19 July 2016 | df = dmy-all }}</ref> | |||
| These bases are classified into two types; adenine and guanine are fused five- and six-membered ]s called ]s, while cytosine and thymine are six-membered rings called ]s.<ref name=berg/> A fifth pyrimidine base, called ] (U), usually takes the place of thymine in RNA and differs from thymine by lacking a ] on its ring. Uracil is not usually found in DNA, occurring only as a breakdown product of cytosine. In addition to RNA and DNA, a large number of artificial ] have also been created to study the proprieties of nucleic acids, or for use in biotechnology.<ref>{{cite journal |author=Verma S, Eckstein F |title=Modified oligonucleotides: synthesis and strategy for users |journal=Annu. Rev. Biochem. |volume=67 |issue= |pages=99–134 |year=1998 |pmid=9759484 |doi=10.1146/annurev.biochem.67.1.99}}</ref> | |||
| === Nucleobase classification === | |||
| ===Grooves=== | |||
| The nucleobases are classified into two types: the ]s, {{mono|A}} and {{mono|G}}, which are fused five- and six-membered ]s, and the ]s, the six-membered rings {{mono|C}} and {{mono|T}}.<ref name=berg /> A fifth pyrimidine nucleobase, ] ({{mono|U}}), usually takes the place of thymine in RNA and differs from thymine by lacking a ] on its ring. In addition to RNA and DNA, many artificial ]s have been created to study the properties of nucleic acids, or for use in biotechnology.<ref>{{cite journal | vauthors = Verma S, Eckstein F | title = Modified oligonucleotides: synthesis and strategy for users | journal = Annual Review of Biochemistry | volume = 67 | pages = 99–134 | year = 1998 | pmid = 9759484 |issn=0066-4154 | doi = 10.1146/annurev.biochem.67.1.99 | doi-access = free }}</ref> | |||
| === Non-canonical bases === | |||
| Twin helical strands form the DNA backbone. Another double helix may be found by tracing the spaces, or grooves, between the strands. These voids are adjacent to the base pairs and may provide a ]. As the strands are not directly opposite each other, the grooves are unequally sized. One groove, the major groove, is 22 Å wide and the other, the minor groove, is 12 Å wide.<ref>{{cite journal |author=Wing R, Drew H, Takano T, Broka C, Tanaka S, Itakura K, Dickerson R |title=Crystal structure analysis of a complete turn of B-DNA |journal=Nature |volume=287 |issue=5784 |pages=755–8 |year=1980 |pmid=7432492 |doi=10.1038/287755a0}}</ref> The narrowness of the minor groove means that the edges of the bases are more accessible in the major groove. As a result, proteins like ]s that can bind to specific sequences in double-stranded DNA usually make contacts to the sides of the bases exposed in the major groove.<ref>{{cite journal |author=Pabo C, Sauer R |title=Protein-DNA recognition |journal=Annu Rev Biochem |volume=53 |pages=293–321 |year=1984 |pmid=6236744 | doi = 10.1146/annurev.bi.53.070184.001453}}</ref> This situation varies in unusual conformations of DNA within the cell ''(see below)'', but the major and minor grooves are always named to reflect the differences in size that would be seen if the DNA is twisted back into the ordinary B form. | |||
| Modified bases occur in DNA. The first of these recognized was ], which was found in the ] of '']'' in 1925.<ref name=Johnson1925>{{cite journal | vauthors = Johnson TB, Coghill RD | year = 1925 | title = Pyrimidines. CIII. The discovery of 5-methylcytosine in tuberculinic acid, the nucleic acid of the tubercle bacillus. | journal = Journal of the American Chemical Society | volume = 47 | pages = 2838–44 | doi=10.1021/ja01688a030|issn=0002-7863}}</ref> The reason for the presence of these noncanonical bases in bacterial viruses (]s) is to avoid the ]s present in bacteria. This enzyme system acts at least in part as a molecular immune system protecting bacteria from infection by viruses.<ref name="pmid27319741">{{cite journal |vauthors=Weigele P, Raleigh EA |title=Biosynthesis and Function of Modified Bases in Bacteria and Their Viruses |journal=Chemical Reviews |volume=116 |issue=20 |pages=12655–12687 |date=October 2016 |pmid=27319741 |doi=10.1021/acs.chemrev.6b00114 |doi-access=free |issn=0009-2665 }}</ref> Modifications of the bases cytosine and adenine, the more common and modified DNA bases, play vital roles in the ] control of gene expression in plants and animals.<ref name="pmid30619465">{{cite journal |vauthors=Kumar S, Chinnusamy V, Mohapatra T |title=Epigenetics of Modified DNA Bases: 5-Methylcytosine and Beyond |journal=Frontiers in Genetics |volume=9 |pages=640 |date=2018 |pmid=30619465 |pmc=6305559 |doi=10.3389/fgene.2018.00640 |issn=1664-8021 |doi-access=free }}</ref> | |||
| ===Base pairing=== | |||
| {{Further|]}} | |||
| A number of noncanonical bases are known to occur in DNA.<ref name="pmid28941008">{{cite journal | vauthors = Carell T, Kurz MQ, Müller M, Rossa M, Spada F | title = Non-canonical Bases in the Genome: The Regulatory Information Layer in DNA | journal = Angewandte Chemie | volume = 57 | issue = 16 | pages = 4296–4312 | date = April 2018 | pmid = 28941008 | doi = 10.1002/anie.201708228 }}</ref> Most of these are modifications of the canonical bases plus uracil. | |||
| Each type of base on one strand forms a bond with just one type of base on the other strand. This is called complementary ]ing. Here, purines form ]s to pyrimidines, with A bonding only to T, and C bonding only to G. This arrangement of two nucleotides binding together across the double helix is called a base pair. As hydrogen bonds are not ], they can be broken and rejoined relatively easily. The two strands of DNA in a double helix can therefore be pulled apart like a zipper, either by a mechanical force or high ].<ref>{{cite journal |author=Clausen-Schaumann H, Rief M, Tolksdorf C, Gaub H |title=Mechanical stability of single DNA molecules |pmc=1300792 |journal=Biophys J |volume=78 |issue=4 |pages=1997–2007 |year=2000 |pmid=10733978 |doi=10.1016/S0006-3495(00)76747-6}}</ref> As a result of this complementarity, all the information in the double-stranded sequence of a DNA helix is duplicated on each strand, which is vital in DNA replication. Indeed, this reversible and specific interaction between complementary base pairs is critical for all the functions of DNA in living organisms.<ref name=Alberts/> | |||
| <div class="thumb tleft" style="background-color: #f9f9f9; border: 1px solid #CCCCCC; margin:0.5em;"> | |||
| * Modified '''Adenine''' | |||
| {|border="0" width=230 border="0" cellpadding="2" cellspacing="0" style="font-size: 85%; border: 1px solid #CCCCCC; margin: 0.3em;" | |||
| ** N6-carbamoyl-methyladenine | |||
| |] | |||
| ** N6-methyadenine | |||
| * Modified '''Guanine''' | |||
| ** 7-Deazaguanine | |||
| ** 7-Methylguanine | |||
| * Modified '''Cytosine''' | |||
| ** N4-Methylcytosine | |||
| ** 5-Carboxylcytosine | |||
| ** 5-Formylcytosine | |||
| ** 5-Glycosylhydroxymethylcytosine | |||
| ** 5-Hydroxycytosine | |||
| ** 5-Methylcytosine | |||
| * Modified '''Thymidine''' | |||
| ** α-Glutamythymidine | |||
| ** α-Putrescinylthymine | |||
| * '''Uracil''' and modifications | |||
| ** ] | |||
| ** Uracil | |||
| ** 5-Dihydroxypentauracil | |||
| ** 5-Hydroxymethyldeoxyuracil | |||
| * Others | |||
| ** Deoxyarchaeosine | |||
| ** 2,6-Diaminopurine (2-Aminoadenine) | |||
| === Grooves === | |||
| ] dye 33258.]] | |||
| Twin helical strands form the DNA backbone. Another double helix may be found tracing the spaces, or grooves, between the strands. These voids are adjacent to the base pairs and may provide a ]. As the strands are not symmetrically located with respect to each other, the grooves are unequally sized. The major groove is {{convert|22|Å|nm}} wide, while the minor groove is {{cvt|12|Å|nm}} in width.<ref>{{cite journal | vauthors = Wing R, Drew H, Takano T, Broka C, Tanaka S, Itakura K, Dickerson RE | title = Crystal structure analysis of a complete turn of B-DNA | journal = Nature | volume = 287 | issue = 5784 | pages = 755–58 | date = October 1980 | pmid = 7432492 | doi = 10.1038/287755a0 | bibcode = 1980Natur.287..755W | s2cid = 4315465 }}</ref> Due to the larger width of the major groove, the edges of the bases are more accessible in the major groove than in the minor groove. As a result, proteins such as ]s that can bind to specific sequences in double-stranded DNA usually make contact with the sides of the bases exposed in the major groove.<ref name="Pabo1984">{{cite journal | vauthors = Pabo CO, Sauer RT | title = Protein-DNA recognition | journal = Annual Review of Biochemistry | volume = 53 | pages = 293–321 | year = 1984 | pmid = 6236744 | doi = 10.1146/annurev.bi.53.070184.001453 }}</ref> This situation varies in unusual conformations of DNA within the cell ''(see below)'', but the major and minor grooves are always named to reflect the differences in width that would be seen if the DNA was twisted back into the ordinary ]. | |||
| === Base pairing === | |||
| {{further|Base pair}} | |||
| <div class="thumb tright" style="background:#f9f9f9; border:1px solid #ccc; margin:0.5em;"> | |||
| {| border="0" cellpadding="2" cellspacing="0" style="width:230px; font-size:85%; border:1px solid #ccc; margin:0.3em;" | |||
| |- | |||
| |] | |||
| |} | |} | ||
| {| |
{| border="0" cellpadding="2" cellspacing="0" style="width:230px; font-size:85%; border:1px solid #ccc; margin:0.3em;" | ||
| |- | |||
| |] | |||
| |] | |||
| |} | |} | ||
| <div style="border: none; width:282px;"><div class="thumbcaption">Top, a '''GC''' base pair with three ]s. Bottom, an '''AT''' base pair with two hydrogen bonds. Non-covalent hydrogen bonds between the pairs are shown as dashed lines.</div></div></div> | <div style="border: none; width:282px;"><div class="thumbcaption">Top, a '''{{mono|GC}}''' base pair with three ]s. Bottom, an '''{{mono|AT}}''' base pair with two hydrogen bonds. Non-covalent hydrogen bonds between the pairs are shown as dashed lines.</div></div></div> | ||
| The two types of base pairs form different numbers of hydrogen bonds, AT forming two hydrogen bonds, and GC forming three hydrogen bonds (see figures, left). | |||
| DNA with high ] is more stable than DNA with low GC-content, but contrary to popular belief, this is not due to the extra hydrogen bond of a GC base pair but rather the contribution of stacking interactions (hydrogen bonding merely provides specificity of the pairing, not stability).<ref name ="Yakovchuk2006">{{cite journal |author=Yakovchuk P, Protozanova E, Frank-Kamenetskii MD |title=Base-stacking and base-pairing contributions into thermal stability of the DNA double helix |journal=Nucleic Acids Res. |volume=34 |issue=2 |pages=564–74 |year=2006 |pmid=16449200 |pmc=1360284 |doi=10.1093/nar/gkj454 |url=http://nar.oxfordjournals.org/cgi/pmidlookup?view=long&pmid=16449200}}</ref> | |||
| As a result, it is both the percentage of GC base pairs and the overall length of a DNA double helix that determine the strength of the association between the two strands of DNA. Long DNA helices with a high GC content have stronger-interacting strands, while short helices with high AT content have weaker-interacting strands.<ref>{{cite journal |author=Chalikian T, Völker J, Plum G, Breslauer K |title=A more unified picture for the thermodynamics of nucleic acid duplex melting: a characterization by calorimetric and volumetric techniques |pmc=22151 |journal=Proc Natl Acad Sci USA |volume=96 |issue=14 |pages=7853–8 |year=1999 |pmid=10393911 |doi=10.1073/pnas.96.14.7853}}</ref> In biology, parts of the DNA double helix that need to separate easily, such as the TATAAT ] in some ]s, tend to have a high AT content, making the strands easier to pull apart.<ref>{{cite journal |author=deHaseth P, Helmann J |title=Open complex formation by Escherichia coli RNA polymerase: the mechanism of polymerase-induced strand separation of double helical DNA |journal=Mol Microbiol |volume=16 |issue=5 |pages=817–24 |year=1995 |pmid=7476180 |doi=10.1111/j.1365-2958.1995.tb02309.x}}</ref> In the laboratory, the strength of this interaction can be measured by finding the temperature required to break the hydrogen bonds, their ] (also called ''T<sub>m</sub>'' value). When all the base pairs in a DNA double helix melt, the strands separate and exist in solution as two entirely independent molecules. These {{Anchors|ssDNA}}single-stranded DNA molecules (''ssDNA'') have no single common shape, but some conformations are more stable than others.<ref>{{cite journal |author=Isaksson J, Acharya S, Barman J, Cheruku P, Chattopadhyaya J |title=Single-stranded adenine-rich DNA and RNA retain structural characteristics of their respective double-stranded conformations and show directional differences in stacking pattern |journal=Biochemistry |volume=43 |issue=51 |pages=15996–6010 |year=2004 |pmid=15609994 | doi = 10.1021/bi048221v}}</ref> | |||
| In a DNA double helix, each type of nucleobase on one strand bonds with just one type of nucleobase on the other strand. This is called ] ]ing. Purines form ]s to pyrimidines, with adenine bonding only to thymine in two hydrogen bonds, and cytosine bonding only to guanine in three hydrogen bonds. This arrangement of two nucleotides binding together across the double helix (from six-carbon ring to six-carbon ring) is called a Watson-Crick base pair. DNA with high ] is more stable than DNA with low {{mono|GC}}-content. A ] (hydrogen bonding the 6-carbon ring to the 5-carbon ring) is a rare variation of base-pairing.<ref name="pmid23818176">{{cite journal |vauthors=Nikolova EN, Zhou H, Gottardo FL, Alvey HS, Kimsey IJ, Al-Hashimi HM |title=A historical account of Hoogsteen base-pairs in duplex DNA |journal=Biopolymers |volume=99 |issue=12 |pages=955–68 |year=2013 |pmid=23818176 |pmc=3844552 |doi=10.1002/bip.22334 }}</ref> As hydrogen bonds are not ], they can be broken and rejoined relatively easily. The two strands of DNA in a double helix can thus be pulled apart like a zipper, either by a mechanical force or high ].<ref>{{cite journal | vauthors = Clausen-Schaumann H, Rief M, Tolksdorf C, Gaub HE | title = Mechanical stability of single DNA molecules | journal = Biophysical Journal | volume = 78 | issue = 4 | pages = 1997–2007 | date = April 2000 | pmid = 10733978 | pmc = 1300792 | doi = 10.1016/S0006-3495(00)76747-6 | bibcode = 2000BpJ....78.1997C }}</ref> As a result of this base pair complementarity, all the information in the double-stranded sequence of a DNA helix is duplicated on each strand, which is vital in DNA replication. This reversible and specific interaction between complementary base pairs is critical for all the functions of DNA in organisms.<ref name=Alberts /> | |||
| ===Sense and antisense=== | |||
| {{Further|]}} | |||
| {{Anchor|ssDNA}} | |||
| A DNA sequence is called "sense" if its sequence is the same as that of a ] copy that is translated into protein.<ref> JCBN/NC-IUB Newsletter 1989, Accessed 07 May 2008</ref> The sequence on the opposite strand is called the "antisense" sequence. Both sense and antisense sequences can exist on different parts of the same strand of DNA (i.e. both strands contain both sense and antisense sequences). In both prokaryotes and eukaryotes, antisense RNA sequences are produced, but the functions of these RNAs are not entirely clear.<ref>{{cite journal |author=Hüttenhofer A, Schattner P, Polacek N |title=Non-coding RNAs: hope or hype? |journal=Trends Genet |volume=21 |issue=5 |pages=289–97 |year=2005 |pmid=15851066 |doi=10.1016/j.tig.2005.03.007}}</ref> One proposal is that antisense RNAs are involved in regulating ] through RNA-RNA base pairing.<ref>{{cite journal |author=Munroe S |title=Diversity of antisense regulation in eukaryotes: multiple mechanisms, emerging patterns |journal=J Cell Biochem |volume=93 |issue=4 |pages=664–71 |year=2004 |pmid=15389973 | doi = 10.1002/jcb.20252}}</ref> | |||
| ==== ssDNA vs. dsDNA ==== | |||
| A few DNA sequences in prokaryotes and eukaryotes, and more in ]s and ]es, blur the distinction between sense and antisense strands by having overlapping genes.<ref>{{cite journal |author=Makalowska I, Lin C, Makalowski W |title=Overlapping genes in vertebrate genomes |journal=Comput Biol Chem |volume=29 |issue=1 |pages=1–12 |year=2005 |pmid=15680581 |doi=10.1016/j.compbiolchem.2004.12.006}}</ref> In these cases, some DNA sequences do double duty, encoding one protein when read along one strand, and a second protein when read in the opposite direction along the other strand. In ], this overlap may be involved in the regulation of gene transcription,<ref>{{cite journal |author=Johnson Z, Chisholm S |title=Properties of overlapping genes are conserved across microbial genomes |journal=Genome Res |volume=14 |issue=11 |pages=2268–72 |year=2004 |pmid=15520290 | doi = 10.1101/gr.2433104 |pmc=525685}}</ref> while in viruses, overlapping genes increase the amount of information that can be encoded within the small viral genome.<ref>{{cite journal |author=Lamb R, Horvath C |title=Diversity of coding strategies in influenza viruses |journal=Trends Genet |volume=7 |issue=8 |pages=261–6 |year=1991 |pmid=1771674}}</ref> | |||
| Most DNA molecules are actually two polymer strands, bound together in a helical fashion by noncovalent bonds; this double-stranded (dsDNA) structure is maintained largely by the intrastrand base stacking interactions, which are strongest for {{mono|G,C}} stacks. The two strands can come apart—a process known as melting—to form two single-stranded DNA (ssDNA) molecules. Melting occurs at high temperatures, low salt and high ] (low pH also melts DNA, but since DNA is unstable due to acid depurination, low pH is rarely used). | |||
| The stability of the dsDNA form depends not only on the {{mono|GC}}-content (% {{mono|G,C}} basepairs) but also on sequence (since stacking is sequence specific) and also length (longer molecules are more stable). The stability can be measured in various ways; a common way is the ] (also called ''T<sub>m</sub>'' value), which is the temperature at which 50% of the double-strand molecules are converted to single-strand molecules; melting temperature is dependent on ionic strength and the concentration of DNA. As a result, it is both the percentage of {{mono|GC}} base pairs and the overall length of a DNA double helix that determines the strength of the association between the two strands of DNA. Long DNA helices with a high {{mono|GC}}-content have more strongly interacting strands, while short helices with high {{mono|AT}} content have more weakly interacting strands.<ref>{{cite journal | vauthors = Chalikian TV, Völker J, Plum GE, Breslauer KJ | title = A more unified picture for the thermodynamics of nucleic acid duplex melting: a characterization by calorimetric and volumetric techniques | journal = Proceedings of the National Academy of Sciences of the United States of America | volume = 96 | issue = 14 | pages = 7853–58 | date = July 1999 | pmid = 10393911 | pmc = 22151 | doi = 10.1073/pnas.96.14.7853 | bibcode = 1999PNAS...96.7853C | doi-access = free }}</ref> In biology, parts of the DNA double helix that need to separate easily, such as the {{mono|TATAAT}} ] in some ], tend to have a high {{mono|AT}} content, making the strands easier to pull apart.<ref>{{cite journal | vauthors = deHaseth PL, Helmann JD | title = Open complex formation by Escherichia coli RNA polymerase: the mechanism of polymerase-induced strand separation of double helical DNA | journal = Molecular Microbiology | volume = 16 | issue = 5 | pages = 817–24 | date = June 1995 | pmid = 7476180 | doi = 10.1111/j.1365-2958.1995.tb02309.x | s2cid = 24479358 }}</ref> | |||
| ===Supercoiling=== | |||
| {{Further|]}} | |||
| DNA can be twisted like a rope in a process called ]ing. With DNA in its "relaxed" state, a strand usually circles the axis of the double helix once every 10.4 base pairs, but if the DNA is twisted the strands become more tightly or more loosely wound.<ref>{{cite journal |author=Benham C, Mielke S |title=DNA mechanics |journal= Annu Rev Biomed Eng |volume=7 |pages=21–53 |year=2005 |pmid=16004565 | doi = 10.1146/annurev.bioeng.6.062403.132016}}</ref> If the DNA is twisted in the direction of the helix, this is positive supercoiling, and the bases are held more tightly together. If they are twisted in the opposite direction, this is negative supercoiling, and the bases come apart more easily. In nature, most DNA has slight negative supercoiling that is introduced by ]s called ]s.<ref name=Champoux>{{cite journal |author=Champoux J |title=DNA topoisomerases: structure, function, and mechanism |journal=Annu Rev Biochem |volume=70 |pages=369–413 |year=2001 |pmid=11395412 | doi = 10.1146/annurev.biochem.70.1.369}}</ref> These enzymes are also needed to relieve the twisting stresses introduced into DNA strands during processes such as ] and ].<ref name=Wang>{{cite journal |author=Wang J |title=Cellular roles of DNA topoisomerases: a molecular perspective |journal=Nat Rev Mol Cell Biol |volume=3 |issue=6 |pages=430–40 |year=2002 |pmid=12042765 | doi = 10.1038/nrm831}}</ref> | |||
| ] | |||
| In the laboratory, the strength of this interaction can be measured by finding the melting temperature ''T<sub>m</sub>'' necessary to break half of the hydrogen bonds. When all the base pairs in a DNA double helix melt, the strands separate and exist in solution as two entirely independent molecules. These single-stranded DNA molecules have no single common shape, but some conformations are more stable than others.<ref>{{cite journal | vauthors = Isaksson J, Acharya S, Barman J, Cheruku P, Chattopadhyaya J | title = Single-stranded adenine-rich DNA and RNA retain structural characteristics of their respective double-stranded conformations and show directional differences in stacking pattern | journal = Biochemistry | volume = 43 | issue = 51 | pages = 15996–6010 | date = December 2004 | pmid = 15609994 | doi = 10.1021/bi048221v | url = http://www.boc.uu.se/boc14www/thesis/johan2005/Paper%20V/Paper%20V.pdf | url-status=live | archive-url = https://web.archive.org/web/20070610205112/http://www.boc.uu.se/boc14www/thesis/johan2005/Paper%20V/Paper%20V.pdf | archive-date = 10 June 2007 | df = dmy-all }}</ref> | |||
| ===Alternate DNA structures=== | |||
| {{Further|], * ], and ]}} | |||
| DNA exists in many possible ] that include ], B-DNA, and ] forms, although, only B-DNA and Z-DNA have been directly observed in functional organisms.<ref name=Ghosh/> The conformation that DNA adopts depends on the hydration level, DNA sequence, the amount and direction of supercoiling, chemical modifications of the bases, the type and concentration of metal ]s, as well as the presence of ]s in solution.<ref>{{cite journal |author=Basu H, Feuerstein B, Zarling D, Shafer R, Marton L |title=Recognition of Z-RNA and Z-DNA determinants by polyamines in solution: experimental and theoretical studies | journal=J Biomol Struct Dyn |volume=6 |issue=2 | pages=299–309 |year=1988 |pmid=2482766}}</ref> | |||
| === Amount === | |||
| The first published reports of A-DNA ]— and also B-DNA used analyses based on ] that provided only a limited amount of structural information for oriented fibers of DNA.<ref>{{cite journal |author=Franklin RE, Gosling RG |title=The Structure of Sodium Thymonucleate Fibres I. The Influence of Water Content |journal=Acta Cryst |volume=6 |issue=8-9 |pages=673–7 |date=6 March 1953 |doi=10.1107/S0365110X53001939 |url=http://hekto.med.unc.edu:8080/CARTER/carter_WWW/Bioch_134/PDF_files/Franklin_Gossling.pdf}}<br/>{{cite journal |author=Franklin RE, Gosling RG |title=The structure of sodium thymonucleate fibres. II. The cylindrically symmetrical Patterson function |journal=Acta Cryst |volume=6 |issue=8-9 |pages=678–85 |year=1953 |month=September |doi=10.1107/S0365110X53001940 }}</ref><ref name=NatFranGos>{{cite journal| title=Molecular Configuration in Sodium Thymonucleate. Franklin R. and Gosling R.G| journal=Nature | volume= 171 | pages= 740–1 | year=1953 | url=http://www.nature.com/nature/dna50/franklingosling.pdf | pmid=13054694 | doi= 10.1038/171740a0| author=Franklin, Rosalind and Gosling, Raymond |format=PDF| issue=4356}}</ref> An alternate analysis was then proposed by Wilkins ''et al.'', in 1953, for the ''in vivo'' B-DNA X-ray diffraction/scattering patterns of highly hydrated DNA fibers in terms of squares of ]s.<ref name=NatWilk>{{cite journal| title=Molecular Structure of Deoxypentose Nucleic Acids | author= Wilkins M.H.F., A.R. Stokes A.R. & Wilson, H.R. | journal=Nature | volume= 171 | pages= 738–740 | year=1953 | url=http://www.nature.com/nature/dna50/wilkins.pdf| pmid=13054693 | doi=10.1038/171738a0| format=PDF| issue=4356}}</ref> In the same journal, ] and ] presented their ] analysis of the DNA X-ray diffraction patterns to suggest that the structure was a double-helix.<ref name=FWPUB/> | |||
| ] of a human. It shows 22 ]s, both the female (XX) and male (XY) versions of the ] (bottom right), as well as the ] (to scale at bottom left). The blue scale to the left of each chromosome pair (and the mitochondrial genome) shows its length in terms of millions of DNA ]s.{{further|Karyotype}}]] | |||
| In humans, the total female ] ] per cell extends for 6.37 Gigabase pairs (Gbp), is 208.23 cm long and weighs 6.51 picograms (pg).<ref name="pmid30813969">{{cite journal| vauthors=Piovesan A, Pelleri MC, Antonaros F, Strippoli P, Caracausi M, Vitale L| title=On the length, weight and GC content of the human genome. | journal=BMC Res Notes | year= 2019 | volume= 12 | issue= 1 | pages= 106 | pmid=30813969 | doi=10.1186/s13104-019-4137-z | pmc=6391780 | doi-access=free }}</ref> Male values are 6.27 Gbp, 205.00 cm, 6.41 pg.<ref name="pmid30813969"/> Each DNA polymer can contain hundreds of millions of nucleotides, such as in ]. Chromosome 1 is the largest human ] with approximately 220 million ]s, and would be {{val|85|u=mm}} long if straightened.<ref name="Gregory_2006" /> | |||
| In ]s, in addition to ], there is also ] (mtDNA) which encodes certain proteins used by the mitochondria. The mtDNA is usually relatively small in comparison to the nuclear DNA. For example, the ] forms closed circular molecules, each of which contains 16,569<ref name="Anderson_1981">{{cite journal | vauthors = Anderson S, Bankier AT, Barrell BG, de Bruijn MH, Coulson AR, Drouin J, Eperon IC, Nierlich DP, Roe BA, Sanger F, Schreier PH, Smith AJ, Staden R, Young IG | display-authors = 6 | title = Sequence and organization of the human mitochondrial genome | journal = Nature | volume = 290 | issue = 5806 | pages = 457–465 | date = April 1981 | pmid = 7219534 | doi = 10.1038/290457a0 | s2cid = 4355527 | bibcode = 1981Natur.290..457A }}</ref><ref>{{Cite web |url=http://chemistry.umeche.maine.edu/CHY431/MitoDNA.html |title=Untitled |access-date=2012-06-13 |archive-url=https://web.archive.org/web/20110813123936/http://chemistry.umeche.maine.edu/CHY431/MitoDNA.html |archive-date=2011-08-13 |url-status=dead }}</ref> DNA base pairs,<ref name=Satoh1991>{{cite journal | vauthors = Satoh M, Kuroiwa T | title = Organization of multiple nucleoids and DNA molecules in mitochondria of a human cell | journal = Experimental Cell Research | volume = 196 | issue = 1 | pages = 137–140 | date = September 1991 | pmid = 1715276 | doi = 10.1016/0014-4827(91)90467-9 }}</ref> with each such molecule normally containing a full set of the mitochondrial genes. Each human mitochondrion contains, on average, approximately 5 such mtDNA molecules.<ref name=Satoh1991/> Each human ] contains approximately 100 mitochondria, giving a total number of mtDNA molecules per human cell of approximately 500.<ref name=Satoh1991/> However, the amount of mitochondria per cell also varies by cell type, and an ] can contain 100,000 mitochondria, corresponding to up to 1,500,000 copies of the mitochondrial genome (constituting up to 90% of the DNA of the cell).<ref name="pmid28721182">{{cite journal | vauthors = Zhang D, Keilty D, Zhang ZF, Chian RC | title = Mitochondria in oocyte aging: current understanding | journal = Facts, Views & Vision in ObGyn | volume = 9 | issue = 1 | pages = 29–38 | date = March 2017 | pmid = 28721182 | pmc = 5506767 }}</ref> | |||
| Although the `B-DNA form' is most common under the conditions found in cells,<ref>{{cite journal |author=Leslie AG, Arnott S, Chandrasekaran R, Ratliff RL |title=Polymorphism of DNA double helices |journal=J. Mol. Biol. |volume=143 |issue=1 |pages=49–72 |year=1980 |pmid=7441761 |doi=10.1016/0022-2836(80)90124-2}}</ref> it is not a well-defined conformation but a family of related DNA conformations<ref>{{cite journal |author=Baianu, I.C. |title=Structural Order and Partial Disorder in Biological systems|journal= Bull. Math. Biol. |volume= 42 |issue=4 |pages=137–141|year=1980}} http://cogprints.org/3822/</ref> that occur at the high hydration levels present in living cells. Their corresponding X-ray diffraction and scattering patterns are characteristic of molecular ] with a significant degree of disorder.<ref>Hosemann R., Bagchi R.N., ''Direct analysis of diffraction by matter'', North-Holland Publs., Amsterdam – New York, 1962.</ref><ref>{{cite journal|author=Baianu, I.C. |title=X-ray scattering by partially disordered membrane systems.|journal=Acta Cryst., |volume=A34 |issue=5 |pages=751–753|year=1978|doi=10.1107/S0567739478001540}}</ref> | |||
| === Sense and antisense === | |||
| Compared to B-DNA, the A-DNA form is a wider right-handed spiral, with a shallow, wide minor groove and a narrower, deeper major groove. The A form occurs under non-physiological conditions in partially dehydrated samples of DNA, while in the cell it may be produced in hybrid pairings of DNA and RNA strands, as well as in enzyme-DNA complexes.<ref>{{cite journal |author=Wahl M, Sundaralingam M |title=Crystal structures of A-DNA duplexes | journal=Biopolymers |volume=44 |issue=1 | pages=45–63 |year=1997 |pmid=9097733 | doi = 10.1002/(SICI)1097-0282(1997)44:1 |doi_brokendate=2009-03-14}}</ref><ref>{{cite journal |author=Lu XJ, Shakked Z, Olson WK |title=A-form conformational motifs in ligand-bound DNA structures |journal=J. Mol. Biol. |volume=300 |issue=4 |pages=819–40 |year=2000 |pmid=10891271 |doi=10.1006/jmbi.2000.3690}}</ref> Segments of DNA where the bases have been chemically modified by ] may undergo a larger change in conformation and adopt the ]. Here, the strands turn about the helical axis in a ] spiral, the opposite of the more common B form.<ref>{{cite journal |author=Rothenburg S, Koch-Nolte F, Haag F |title=DNA methylation and Z-DNA formation as mediators of quantitative differences in the expression of alleles | journal=Immunol Rev |volume=184 | pages=286–98 |year=2001|pmid=12086319 |doi=10.1034/j.1600-065x.2001.1840125.x}}</ref> These unusual structures can be recognized by specific Z-DNA binding proteins and may be involved in the regulation of transcription.<ref>{{cite journal |author=Oh D, Kim Y, Rich A |title=Z-DNA-binding proteins can act as potent effectors of gene expression in vivo |journal=Proc. Natl. Acad. Sci. U.S.A. |volume=99 |issue=26 |pages=16666–71 |year=2002 |pmid=12486233 |doi=10.1073/pnas.262672699 |pmc=139201 |url=http://www.pnas.org/cgi/pmidlookup?view=long&pmid=12486233}}</ref> | |||
| {{further|Sense (molecular biology)}} | |||
| A ] is called a "sense" sequence if it is the same as that of a ] copy that is translated into protein.<ref> {{Webarchive|url=https://web.archive.org/web/20080424015915/http://www.chem.qmul.ac.uk/iubmb/newsletter/misc/DNA.html |date=24 April 2008 }} JCBN/NC-IUB Newsletter 1989. Retrieved 7 May 2008</ref> The sequence on the opposite strand is called the "antisense" sequence. Both sense and antisense sequences can exist on different parts of the same strand of DNA (i.e. both strands can contain both sense and antisense sequences). In both prokaryotes and eukaryotes, antisense RNA sequences are produced, but the functions of these RNAs are not entirely clear.<ref>{{cite journal | vauthors = Hüttenhofer A, Schattner P, Polacek N | title = Non-coding RNAs: hope or hype? | journal = Trends in Genetics | volume = 21 | issue = 5 | pages = 289–97 | date = May 2005 | pmid = 15851066 | doi = 10.1016/j.tig.2005.03.007 }}</ref> One proposal is that antisense RNAs are involved in regulating ] through RNA-RNA base pairing.<ref>{{cite journal | vauthors = Munroe SH | title = Diversity of antisense regulation in eukaryotes: multiple mechanisms, emerging patterns | journal = Journal of Cellular Biochemistry | volume = 93 | issue = 4 | pages = 664–71 | date = November 2004 | pmid = 15389973 | doi = 10.1002/jcb.20252 | s2cid = 23748148 }}</ref> | |||
| ] repeats. The looped conformation of the DNA backbone is very different from the typical DNA helix.<ref>Created from </ref>]] | |||
| A few DNA sequences in prokaryotes and eukaryotes, and more in ]s and ]es, blur the distinction between sense and antisense strands by having ]s.<ref>{{cite journal | vauthors = Makalowska I, Lin CF, Makalowski W | title = Overlapping genes in vertebrate genomes | journal = Computational Biology and Chemistry | volume = 29 | issue = 1 | pages = 1–12 | date = February 2005 | pmid = 15680581 | doi = 10.1016/j.compbiolchem.2004.12.006 }}</ref> In these cases, some DNA sequences do double duty, encoding one protein when read along one strand, and a second protein when read in the opposite direction along the other strand. In ], this overlap may be involved in the regulation of gene transcription,<ref>{{cite journal | vauthors = Johnson ZI, Chisholm SW | title = Properties of overlapping genes are conserved across microbial genomes | journal = Genome Research | volume = 14 | issue = 11 | pages = 2268–72 | date = November 2004 | pmid = 15520290 | pmc = 525685 | doi = 10.1101/gr.2433104 }}</ref> while in viruses, overlapping genes increase the amount of information that can be encoded within the small viral genome.<ref>{{cite journal | vauthors = Lamb RA, Horvath CM | title = Diversity of coding strategies in influenza viruses | journal = Trends in Genetics | volume = 7 | issue = 8 | pages = 261–66 | date = August 1991 | pmid = 1771674 | doi = 10.1016/0168-9525(91)90326-L | pmc = 7173306 }}</ref> | |||
| ===Quadruplex structures=== | |||
| {{Further|]}} | |||
| === Supercoiling === | |||
| At the ends of the linear chromosomes are specialized regions of DNA called ]s. The main function of these regions is to allow the cell to replicate chromosome ends using the enzyme ], as the enzymes that normally replicate DNA cannot copy the extreme 3′ ends of chromosomes.<ref name=Greider>{{cite journal |author=Greider C, Blackburn E |title=Identification of a specific telomere terminal transferase activity in Tetrahymena extracts | journal=Cell |volume=43 |issue=2 Pt 1 | pages=405–13 |year=1985 |pmid=3907856 |doi=10.1016/0092-8674(85)90170-9}}</ref> These specialized chromosome caps also help protect the DNA ends, and stop the ] systems in the cell from treating them as damage to be corrected.<ref name=Nugent>{{cite journal |author=Nugent C, Lundblad V |title=The telomerase reverse transcriptase: components and regulation | url=http://www.genesdev.org/cgi/content/full/12/8/1073 | journal=Genes Dev |volume=12 |issue=8 | pages=1073–85 |year=1998 |pmid=9553037 |doi=10.1101/gad.12.8.1073}}</ref> In ], telomeres are usually lengths of single-stranded DNA containing several thousand repeats of a simple TTAGGG sequence.<ref>{{cite journal |author=Wright W, Tesmer V, Huffman K, Levene S, Shay J |title=Normal human chromosomes have long G-rich telomeric overhangs at one end | url=http://www.genesdev.org/cgi/content/full/11/21/2801 | journal=Genes Dev |volume=11 |issue=21 | pages=2801–9 |year=1997 |pmid=9353250 |doi=10.1101/gad.11.21.2801 |pmc=316649}}</ref> | |||
| {{further|DNA supercoil}} | |||
| DNA can be twisted like a rope in a process called ]ing. With DNA in its "relaxed" state, a strand usually circles the axis of the double helix once every 10.4 base pairs, but if the DNA is twisted the strands become more tightly or more loosely wound.<ref>{{cite journal | vauthors = Benham CJ, Mielke SP | s2cid = 1427671 | title = DNA mechanics | journal = Annual Review of Biomedical Engineering | volume = 7 | pages = 21–53 | year = 2005 | pmid = 16004565 | doi = 10.1146/annurev.bioeng.6.062403.132016 | url = http://pdfs.semanticscholar.org/ab63/d57290ebf9bc3536fd3f2257a2b509076fc1.pdf | archive-url = https://web.archive.org/web/20190301225243/http://pdfs.semanticscholar.org/ab63/d57290ebf9bc3536fd3f2257a2b509076fc1.pdf | url-status = dead | archive-date = 1 March 2019 }}</ref> If the DNA is twisted in the direction of the helix, this is positive supercoiling, and the bases are held more tightly together. If they are twisted in the opposite direction, this is negative supercoiling, and the bases come apart more easily. In nature, most DNA has slight negative supercoiling that is introduced by ]s called ]s.<ref name=Champoux>{{cite journal | vauthors = Champoux JJ | s2cid = 18144189 | title = DNA topoisomerases: structure, function, and mechanism | journal = Annual Review of Biochemistry | volume = 70 | pages = 369–413 | year = 2001 | pmid = 11395412 | doi = 10.1146/annurev.biochem.70.1.369 }}</ref> These enzymes are also needed to relieve the twisting stresses introduced into DNA strands during processes such as ] and ].<ref name=Wang>{{cite journal | vauthors = Wang JC | title = Cellular roles of DNA topoisomerases: a molecular perspective | journal = Nature Reviews Molecular Cell Biology | volume = 3 | issue = 6 | pages = 430–40 | date = June 2002 | pmid = 12042765 | doi = 10.1038/nrm831 | s2cid = 205496065 }}</ref> | |||
| These guanine-rich sequences may stabilize chromosome ends by forming structures of stacked sets of four-base units, rather than the usual base pairs found in other DNA molecules. Here, four guanine bases form a flat plate and these flat four-base units then stack on top of each other, to form a stable '']'' structure.<ref name=Burge>{{cite journal |author=Burge S, Parkinson G, Hazel P, Todd A, Neidle S |title=Quadruplex DNA: sequence, topology and structure | journal=Nucleic Acids Res |volume=34 |issue=19 | pages=5402–15 |year=2006 |pmid=17012276 |pmc=1636468 | doi = 10.1093/nar/gkl655 |url=http://nar.oxfordjournals.org/cgi/pmidlookup?view=long&pmid=17012276}}</ref> These structures are stabilized by hydrogen bonding between the edges of the bases and ] of a metal ion in the centre of each four-base unit.<ref>{{cite journal |author=Parkinson G, Lee M, Neidle S |title=Crystal structure of parallel quadruplexes from human telomeric DNA | journal=Nature |volume=417 |issue=6891 | pages=876–80 |year=2002 |pmid=12050675 | doi = 10.1038/nature755}}</ref> Other structures can also be formed, with the central set of four bases coming from either a single strand folded around the bases, or several different parallel strands, each contributing one base to the central structure. | |||
| === Alternative DNA structures === | |||
| In addition to these stacked structures, telomeres also form large loop structures called telomere loops, or T-loops. Here, the single-stranded DNA curls around in a long circle stabilized by telomere-binding proteins.<ref>{{cite journal |author=Griffith J, Comeau L, Rosenfield S, Stansel R, Bianchi A, Moss H, de Lange T |title=Mammalian telomeres end in a large duplex loop | journal=Cell |volume=97 |issue=4 | pages=503–14 |year=1999 |pmid=10338214 |doi=10.1016/S0092-8674(00)80760-6}}</ref> At the very end of the T-loop, the single-stranded telomere DNA is held onto a region of double-stranded DNA by the telomere strand disrupting the double-helical DNA and base pairing to one of the two strands. This ] structure is called a displacement loop or ].<ref name=Burge/> | |||
| {{further|Molecular Structure of Nucleic Acids: A Structure for Deoxyribose Nucleic Acid|Molecular models of DNA|DNA structure}} | |||
| ], ] and ]]] | |||
| DNA exists in many possible ] that include ], ], and ] forms, although only B-DNA and Z-DNA have been directly observed in functional organisms.<ref name=Ghosh /> The conformation that DNA adopts depends on the hydration level, DNA sequence, the amount and direction of supercoiling, chemical modifications of the bases, the type and concentration of metal ]s, and the presence of ]s in solution.<ref>{{cite journal | vauthors = Basu HS, Feuerstein BG, Zarling DA, Shafer RH, Marton LJ | title = Recognition of Z-RNA and Z-DNA determinants by polyamines in solution: experimental and theoretical studies | journal = Journal of Biomolecular Structure & Dynamics | volume = 6 | issue = 2 | pages = 299–309 | date = October 1988 | pmid = 2482766 | doi = 10.1080/07391102.1988.10507714 }}</ref> | |||
| <div class="thumb tright" style="background-color: #f9f9f9; border: 1px solid #CCCCCC; margin:0.5em;"> | |||
| {|border="0" width=200 border="0" cellpadding="2" cellspacing="0" style="font-size: 85%; border: 1px solid #CCCCCC; margin: 0.3em;" | |||
| The first published reports of A-DNA ]—and also B-DNA—used analyses based on ]s that provided only a limited amount of structural information for oriented fibers of DNA.<ref> | |||
| |] | |||
| * {{cite journal |vauthors=Franklin RE, Gosling RG |title=The Structure of Sodium Thymonucleate Fibres I. The Influence of Water Content |journal=Acta Crystallogr |volume=6 |issue=8–9 |pages=673–77 |date=6 March 1953 |doi=10.1107/S0365110X53001939 |bibcode=1953AcCry...6..673F |url=http://journals.iucr.org/q/issues/1953/08-09/00/a00979/a00979.pdf |url-status=live |archive-url=https://web.archive.org/web/20160109043915/http://journals.iucr.org/q/issues/1953/08-09/00/a00979/a00979.pdf |archive-date=9 January 2016 |doi-access=free }} | |||
| |] | |||
| * {{cite journal |vauthors=Franklin RE, Gosling RG |title=The structure of sodium thymonucleate fibres. II. The cylindrically symmetrical Patterson function |journal=Acta Crystallogr |volume=6 |issue=8–9 |pages=678–85 |year=1953|doi=10.1107/S0365110X53001940|bibcode=1953AcCry...6..678F |url=http://journals.iucr.org/q/issues/1953/08-09/00/a00980/a00980.pdf |archive-url=https://web.archive.org/web/20170629084321/http://journals.iucr.org/q/issues/1953/08-09/00/a00980/a00980.pdf |archive-date=2017-06-29 |url-status=live |doi-access=free }}</ref><ref name=NatFranGos>{{cite journal | vauthors = Franklin RE, Gosling RG | title = Molecular configuration in sodium thymonucleate | journal = Nature | volume = 171 | issue = 4356 | pages = 740–41 | date = April 1953 | pmid = 13054694 | doi = 10.1038/171740a0 | url = http://www.nature.com/nature/dna50/franklingosling.pdf | bibcode = 1953Natur.171..740F | s2cid = 4268222 | url-status=live | archive-url = https://web.archive.org/web/20110103160712/http://www.nature.com/nature/dna50/franklingosling.pdf | archive-date = 3 January 2011 | df = dmy-all }}</ref> An alternative analysis was proposed by Wilkins ''et al.'' in 1953 for the '']'' B-DNA X-ray diffraction-scattering patterns of highly hydrated DNA fibers in terms of squares of ]s.<ref name=NatWilk>{{cite journal | vauthors = Wilkins MH, Stokes AR, Wilson HR | title = Molecular structure of deoxypentose nucleic acids | journal = Nature | volume = 171 | issue = 4356 | pages = 738–40 | date = April 1953 | pmid = 13054693 | doi = 10.1038/171738a0 | url = http://www.nature.com/nature/dna50/wilkins.pdf | bibcode = 1953Natur.171..738W | s2cid = 4280080 | url-status=live | archive-url = https://web.archive.org/web/20110513234223/http://www.nature.com/nature/dna50/wilkins.pdf | archive-date = 13 May 2011 | df = dmy-all }}</ref> In the same journal, ] and ] presented their ] analysis of the DNA X-ray diffraction patterns to suggest that the structure was a double helix.<ref name="Watson-1953" /> | |||
| Although the ''B-DNA form'' is most common under the conditions found in cells,<ref>{{cite journal | vauthors = Leslie AG, Arnott S, Chandrasekaran R, Ratliff RL | title = Polymorphism of DNA double helices | journal = Journal of Molecular Biology | volume = 143 | issue = 1 | pages = 49–72 | date = October 1980 | pmid = 7441761 | doi = 10.1016/0022-2836(80)90124-2 }}</ref> it is not a well-defined conformation but a family of related DNA conformations<ref>{{cite journal|vauthors=Baianu IC|s2cid=189888972|year=1980|title=Structural Order and Partial Disorder in Biological systems|url=http://cogprints.org/3822/|journal=Bull. Math. Biol.|volume=42|issue=4|pages=137–41|doi=10.1007/BF02462372}}</ref> that occur at the high hydration levels present in cells. Their corresponding X-ray diffraction and scattering patterns are characteristic of molecular ] with a significant degree of disorder.<ref>{{cite book | vauthors = Hosemann R, Bagchi RN | title = Direct analysis of diffraction by matter | publisher = North-Holland Publishers | location = Amsterdam – New York | year = 1962 }}</ref><ref>{{cite journal|vauthors=Baianu IC|title=X-ray scattering by partially disordered membrane systems|journal=Acta Crystallogr A|volume=34|issue=5|pages=751–53|year=1978|doi=10.1107/S0567739478001540|bibcode=1978AcCrA..34..751B|url=http://journals.iucr.org/a/issues/1978/05/00/a15615/a15615.pdf|access-date=29 August 2019|archive-date=14 March 2020|archive-url=https://web.archive.org/web/20200314050140/http://journals.iucr.org/a/issues/1978/05/00/a15615/a15615.pdf|url-status=dead}}</ref> | |||
| Compared to B-DNA, the A-DNA form is a wider ] spiral, with a shallow, wide minor groove and a narrower, deeper major groove. The A form occurs under non-physiological conditions in partly dehydrated samples of DNA, while in the cell it may be produced in hybrid pairings of DNA and RNA strands, and in enzyme-DNA complexes.<ref>{{cite journal | vauthors = Wahl MC, Sundaralingam M | title = Crystal structures of A-DNA duplexes | journal = Biopolymers | volume = 44 | issue = 1 | pages = 45–63 | year = 1997 | pmid = 9097733 | doi = 10.1002/(SICI)1097-0282(1997)44:1<45::AID-BIP4>3.0.CO;2-# }}</ref><ref>{{cite journal | vauthors = Lu XJ, Shakked Z, Olson WK | title = A-form conformational motifs in ligand-bound DNA structures | journal = Journal of Molecular Biology | volume = 300 | issue = 4 | pages = 819–40 | date = July 2000 | pmid = 10891271 | doi = 10.1006/jmbi.2000.3690 }}</ref> Segments of DNA where the bases have been chemically modified by ] may undergo a larger change in conformation and adopt the ]. Here, the strands turn about the helical axis in a left-handed spiral, the opposite of the more common B form.<ref>{{cite journal | vauthors = Rothenburg S, Koch-Nolte F, Haag F | title = DNA methylation and Z-DNA formation as mediators of quantitative differences in the expression of alleles | journal = Immunological Reviews | volume = 184 | pages = 286–98 | date = December 2001 | pmid = 12086319 | doi = 10.1034/j.1600-065x.2001.1840125.x | s2cid = 20589136 }}</ref> These unusual structures can be recognized by specific Z-DNA binding proteins and may be involved in the regulation of transcription.<ref>{{cite journal | vauthors = Oh DB, Kim YG, Rich A | title = Z-DNA-binding proteins can act as potent effectors of gene expression in vivo | journal = Proceedings of the National Academy of Sciences of the United States of America | volume = 99 | issue = 26 | pages = 16666–71 | date = December 2002 | pmid = 12486233 | pmc = 139201 | doi = 10.1073/pnas.262672699 | bibcode = 2002PNAS...9916666O | doi-access = free }}</ref> | |||
| === Alternative DNA chemistry === | |||
| {{further|hypothetical types of biochemistry}} | |||
| For many years, ] have proposed the existence of a ], a postulated microbial ] of Earth that uses radically different biochemical and molecular processes than currently known life. One of the proposals was the existence of lifeforms that use ]. A report in 2010 of the possibility in the ] ] was announced,<ref name='arsenic extremophile'>{{cite news | vauthors = Palmer J |title=Arsenic-loving bacteria may help in hunt for alien life |date=2 December 2010 |url=https://www.bbc.co.uk/news/science-environment-11886943 |work=BBC News |access-date=2 December 2010 |url-status=live |archive-url=https://web.archive.org/web/20101203045804/http://www.bbc.co.uk/news/science-environment-11886943 |archive-date=3 December 2010 }}</ref><ref name="Space">{{cite news | vauthors = Bortman H |title=Arsenic-Eating Bacteria Opens New Possibilities for Alien Life |date=2 December 2010 |url=http://www.space.com/scienceastronomy/arsenic-bacteria-alien-life-101202.html |website=Space.com |access-date=2 December 2010 |url-status=live |archive-url=https://web.archive.org/web/20101204235915/http://www.space.com/scienceastronomy/arsenic-bacteria-alien-life-101202.html |archive-date=4 December 2010 }}</ref> though the research was disputed,<ref name="Space" /><ref>{{cite journal | vauthors = Katsnelson A |title=Arsenic-eating microbe may redefine chemistry of life |date=2 December 2010 |url=http://www.nature.com/news/2010/101202/full/news.2010.645.html |journal=Nature News |doi=10.1038/news.2010.645 |url-status=live |archive-url=https://web.archive.org/web/20120212155007/http://www.nature.com/news/2010/101202/full/news.2010.645.html |archive-date=12 February 2012 }}</ref> and evidence suggests the bacterium actively prevents the incorporation of arsenic into the DNA backbone and other biomolecules.<ref name="Nature">{{cite journal | vauthors = Cressey D |s2cid=87341731 |title='Arsenic-life' Bacterium Prefers Phosphorus after all |date=3 October 2012 |journal=Nature News |doi=10.1038/nature.2012.11520}}</ref> | |||
| === Quadruplex structures === | |||
| {{further|G-quadruplex}} | |||
| ] repeats. The looped conformation of the DNA backbone is very different from the typical DNA helix. The green spheres in the center represent potassium ions.<ref>{{Cite web|title=Structure and packing of human telomeric DNA|url=http://ndbserver.rutgers.edu/service/ndb/atlas/summary?searchTarget=UD0017|access-date=2023-05-18|website=ndbserver.rutgers.edu}}</ref>]] | |||
| At the ends of the linear chromosomes are specialized regions of DNA called ]s. The main function of these regions is to allow the cell to replicate chromosome ends using the enzyme ], as the enzymes that normally replicate DNA cannot copy the extreme 3′ ends of chromosomes.<ref name=Greider>{{cite journal | vauthors = Greider CW, Blackburn EH | title = Identification of a specific telomere terminal transferase activity in Tetrahymena extracts | journal = Cell | volume = 43 | issue = 2 Pt 1 | pages = 405–13 | date = December 1985 | pmid = 3907856 | doi = 10.1016/0092-8674(85)90170-9 | doi-access = free }}</ref> These specialized chromosome caps also help protect the DNA ends, and stop the ] systems in the cell from treating them as damage to be corrected.<ref name=Nugent>{{cite journal | vauthors = Nugent CI, Lundblad V | title = The telomerase reverse transcriptase: components and regulation | journal = Genes & Development | volume = 12 | issue = 8 | pages = 1073–85 | date = April 1998 | pmid = 9553037 | doi = 10.1101/gad.12.8.1073 | doi-access = free }}</ref> In ], telomeres are usually lengths of single-stranded DNA containing several thousand repeats of a simple TTAGGG sequence.<ref>{{cite journal | vauthors = Wright WE, Tesmer VM, Huffman KE, Levene SD, Shay JW | title = Normal human chromosomes have long G-rich telomeric overhangs at one end | journal = Genes & Development | volume = 11 | issue = 21 | pages = 2801–09 | date = November 1997 | pmid = 9353250 | pmc = 316649 | doi = 10.1101/gad.11.21.2801 }}</ref> | |||
| These guanine-rich sequences may stabilize chromosome ends by forming structures of stacked sets of four-base units, rather than the usual base pairs found in other DNA molecules. Here, four guanine bases, known as a ], form a flat plate. These flat four-base units then stack on top of each other to form a stable ] structure.<ref name=Burge>{{cite journal | vauthors = Burge S, Parkinson GN, Hazel P, Todd AK, Neidle S | title = Quadruplex DNA: sequence, topology and structure | journal = Nucleic Acids Research | volume = 34 | issue = 19 | pages = 5402–15 | year = 2006 | pmid = 17012276 | pmc = 1636468 | doi = 10.1093/nar/gkl655 }}</ref> These structures are stabilized by hydrogen bonding between the edges of the bases and ] of a metal ion in the centre of each four-base unit.<ref>{{cite journal | vauthors = Parkinson GN, Lee MP, Neidle S | title = Crystal structure of parallel quadruplexes from human telomeric DNA | journal = Nature | volume = 417 | issue = 6891 | pages = 876–80 | date = June 2002 | pmid = 12050675 | doi = 10.1038/nature755 | bibcode = 2002Natur.417..876P | s2cid = 4422211 }}</ref> Other structures can also be formed, with the central set of four bases coming from either a single strand folded around the bases, or several different parallel strands, each contributing one base to the central structure. | |||
| In addition to these stacked structures, telomeres also form large loop structures called telomere loops, or T-loops. Here, the single-stranded DNA curls around in a long circle stabilized by telomere-binding proteins.<ref>{{cite journal | vauthors = Griffith JD, Comeau L, Rosenfield S, Stansel RM, Bianchi A, Moss H, de Lange T | s2cid = 721901 | title = Mammalian telomeres end in a large duplex loop | journal = Cell | volume = 97 | issue = 4 | pages = 503–14 | date = May 1999 | pmid = 10338214 | doi = 10.1016/S0092-8674(00)80760-6 | citeseerx = 10.1.1.335.2649 }}</ref> At the very end of the T-loop, the single-stranded telomere DNA is held onto a region of double-stranded DNA by the telomere strand disrupting the double-helical DNA and base pairing to one of the two strands. This ] structure is called a displacement loop or ].<ref name=Burge /> | |||
| === Branched DNA === | |||
| {{further|Branched DNA|DNA nanotechnology}} | |||
| <div class="thumb tright" style="background:#f9f9f9; border:1px solid #ccc; margin:0.5em;"> | |||
| {| border="0" cellpadding="2" cellspacing="0" style="width:200px; font-size:85%; border:1px solid #ccc; margin:0.3em;" | |||
| |] | |||
| |] | |||
| |- | |- | ||
| |align=center|Single branch | |align=center|Single branch | ||
| Line 90: | Line 154: | ||
| |} | |} | ||
| <div style="border: none; width:200px;font-size: 90%;"><div class="thumbcaption">] can form networks containing multiple branches.</div></div></div> | <div style="border: none; width:200px;font-size: 90%;"><div class="thumbcaption">] can form networks containing multiple branches.</div></div></div> | ||
| In DNA, ] occurs when non-complementary regions exist at the end of an otherwise complementary double-strand of DNA. However, branched DNA can occur if a third strand of DNA is introduced and contains adjoining regions able to hybridize with the frayed regions of the pre-existing double-strand. Although the simplest example of branched DNA involves only three strands of DNA, complexes involving additional strands and multiple branches are also possible.<ref>{{cite journal | vauthors = Seeman NC | title = DNA enables nanoscale control of the structure of matter | journal = Quarterly Reviews of Biophysics | volume = 38 | issue = 4 | pages = 363–71 | date = November 2005 | pmid = 16515737 | pmc = 3478329 | doi = 10.1017/S0033583505004087 }}</ref> Branched DNA can be used in ] to construct geometric shapes, see the section on ] below. | |||
| === |
=== Artificial bases === | ||
| {{Main|Nucleic acid analogue}} | |||
| {{Further|] and ]}} | |||
| Several artificial nucleobases have been synthesized, and successfully incorporated in the eight-base DNA analogue named ]. Dubbed S, B, P, and Z, these artificial bases are capable of bonding with each other in a predictable way (S–B and P–Z), maintain the double helix structure of DNA, and be transcribed to RNA. Their existence could be seen as an indication that there is nothing special about the four natural nucleobases that evolved on Earth.<ref>{{cite journal | vauthors = Warren M |title=Four new DNA letters double life's alphabet | journal = Nature |date=21 February 2019 | doi = 10.1038/d41586-019-00650-8 | pmid = 30809059 | volume=566 | issue = 7745 | page=436| doi-access = free | bibcode = 2019Natur.566..436W }}</ref><ref>{{cite journal| vauthors = Hoshika S, Leal NA, Kim MJ, Kim MS, Karalkar NB, Kim HJ, Bates AM, Watkins NE, SantaLucia HA, Meyer AJ, DasGupta S, Piccirilli JA, Ellington AD, SantaLucia J, Georgiadis MM, Benner SA | display-authors = 6 |title=Hachimoji DNA and RNA: A genetic system with eight building blocks (paywall)|journal=] |volume=363 |issue=6429 |pages=884–887 |date=22 February 2019 | doi = 10.1126/science.aat0971 | pmid = 30792304 | pmc=6413494 | bibcode=2019Sci...363..884H}}</ref> On the other hand, DNA is tightly related to ] which does not only act as a transcript of DNA but also performs as molecular machines many tasks in cells. For this purpose it has to fold into a structure. It has been shown that to allow to create all possible structures at least four bases are required for the corresponding ],<ref>{{cite journal | vauthors = Burghardt B, Hartmann AK | title = RNA secondary structure design | journal = Physical Review E | volume = 75 | issue = 2 | pages = 021920 | date = February 2007 | doi = 10.1103/PhysRevE.75.021920 | pmid = 17358380 | url = https://link.aps.org/doi/10.1103/PhysRevE.75.021920| arxiv = physics/0609135 | bibcode = 2007PhRvE..75b1920B | s2cid = 17574854 }}</ref> while a higher number is also possible but this would be against the natural ]. | |||
| In DNA ] occurs when non-complementary regions exist at the end of an otherwise complementary double-strand of DNA. However, branched DNA can occur if a third strand of DNA is introduced and contains adjoining regions able to hybridize with the frayed regions of the pre-existing double-strand. Although the simplest example of branched DNA involves only three strands of DNA, complexes involving additional strands and multiple branches are also possible.<ref>{{cite journal |author=Seeman NC |title=DNA enables nanoscale control of the structure of matter |journal=Q. Rev. Biophys. |volume=38 |issue=4 |pages=363–71 |year=2005 |month=November |pmid=16515737 |doi=10.1017/S0033583505004087}}</ref> Branched DNA can be used in ] to construct geometric shapes, see the section on ] below. | |||
| === |
===Acidity=== | ||
| The phosphate groups of DNA give it similar ]ic properties to ] and it can be considered as a ]. It will be fully ionized at a normal cellular pH, releasing ]s which leave behind negative charges on the phosphate groups. These negative charges protect DNA from breakdown by ] by repelling ]s which could hydrolyze it.<ref name="Reusch">{{cite web | vauthors = Reusch W |title=Nucleic Acids |url=https://www2.chemistry.msu.edu/faculty/reusch/VirtTxtJml/nucacids.htm |publisher=Michigan State University |access-date=30 June 2022}}</ref> | |||
| {{Further|]}} | |||
| ===Macroscopic appearance=== | |||
| DNA may carry out low-frequency collective motion as observed by the ] | |||
| ] | |||
| <ref>Painter, P.C., Mosher, L.E. and Rhoads, C. (1981) Low-frequency modes in the Raman spectrum of DNA. ''Biopolymers'', 20, 243-247.</ref> | |||
| <ref>Urabe, H., Tominaga, Y. and Kubota, K. (1983) Experimental evidence of collective vibrations in DNA double helix Raman spectroscopy. ''Journal of Chemical Physics'', 78, 5937-5939.</ref> | |||
| and analyzed with the quasi-continuum model. | |||
| <ref>Kuo-Chen Chou (1984) Low-frequency vibration of DNA molecules. ''Biochemical Journal'', 221, 27-31.</ref> | |||
| <ref>Chou, K.C., Maggiora, G.M. and Mao, B. (1989) Quasi-continuum models of twist-like and accordion-like low-frequency motions in DNA. ''Biophysical Journal'', 56, 295-305.</ref> | |||
| Pure DNA extracted from cells forms white, stringy clumps.<ref>{{cite web |title=How To Extract DNA From Anything Living |url=https://learn.genetics.utah.edu/content/labs/extraction/howto/ |publisher=University of Utah |access-date=30 June 2022}}</ref> | |||
| ==Chemical modifications== | |||
| <div class="thumb tright" style="background-color: #f9f9f9; border: 1px solid #CCCCCC; margin:0.5em;"> | |||
| == Chemical modifications and altered DNA packaging == | |||
| {|border="0" width=300 border="0" cellpadding="2" cellspacing="0" style="font-size: 85%; border: 1px solid #CCCCCC; margin: 0.3em;" | |||
| |] | |||
| === Base modifications and DNA packaging === | |||
| |] | |||
| {{further|DNA methylation|Chromatin remodeling}} | |||
| |] | |||
| <div class="thumb tright" style="background:#f9f9f9; border:1px solid #ccc; margin:0.5em;"> | |||
| {| border="0" cellpadding="2" cellspacing="0" style="width:300px; font-size:85%; border:1px solid #ccc; margin:0.3em;" | |||
| |- | |||
| |] | |||
| |] | |||
| |] | |||
| |- | |- | ||
| |align=center|] | |align=center|] | ||
| Line 118: | Line 185: | ||
| |} | |} | ||
| <div style="border: none; width:300px;font-size: 90%;"><div class="thumbcaption">Structure of cytosine with and without the 5-methyl group. ] converts 5-methylcytosine into thymine.</div></div></div> | <div style="border: none; width:300px;font-size: 90%;"><div class="thumbcaption">Structure of cytosine with and without the 5-methyl group. ] converts 5-methylcytosine into thymine.</div></div></div> | ||
| The expression of genes is influenced by how the DNA is packaged in chromosomes, in a structure called ]. Base modifications can be involved in packaging, with regions that have low or no gene expression usually containing high levels of ] of ] bases. DNA packaging and its influence on gene expression can also occur by covalent modifications of the ] protein core around which DNA is wrapped in the chromatin structure or else by remodeling carried out by chromatin remodeling complexes (see ]). There is, further, ] between DNA methylation and histone modification, so they can coordinately affect chromatin and gene expression.<ref>{{cite journal | vauthors = Hu Q, Rosenfeld MG | title = Epigenetic regulation of human embryonic stem cells | journal = Frontiers in Genetics | volume = 3 | pages = 238 | year = 2012 | pmid = 23133442 | pmc = 3488762 | doi = 10.3389/fgene.2012.00238 | doi-access = free }}</ref> | |||
| For one example, cytosine methylation produces ], which is important for ] of chromosomes.<ref>{{cite journal | vauthors = Klose RJ, Bird AP | title = Genomic DNA methylation: the mark and its mediators | journal = Trends in Biochemical Sciences | volume = 31 | issue = 2 | pages = 89–97 | date = February 2006 | pmid = 16403636 | doi = 10.1016/j.tibs.2005.12.008 }}</ref> The average level of methylation varies between organisms—the worm '']'' lacks cytosine methylation, while ]s have higher levels, with up to 1% of their DNA containing 5-methylcytosine.<ref>{{cite journal | vauthors = Bird A | title = DNA methylation patterns and epigenetic memory | journal = Genes & Development | volume = 16 | issue = 1 | pages = 6–21 | date = January 2002 | pmid = 11782440 | doi = 10.1101/gad.947102 | doi-access = free }}</ref> Despite the importance of 5-methylcytosine, it can ] to leave a thymine base, so methylated cytosines are particularly prone to ]s.<ref>{{cite book | vauthors = Walsh CP, Xu GL | title = DNA Methylation: Basic Mechanisms | chapter = Cytosine methylation and DNA repair | volume = 301 | pages = 283–315 | year = 2006 | pmid = 16570853 | doi = 10.1007/3-540-31390-7_11 | isbn = 3-540-29114-8 | series = Current Topics in Microbiology and Immunology }}</ref> Other base modifications include adenine methylation in bacteria, the presence of ] in the ],<ref>{{cite journal | vauthors = Kriaucionis S, Heintz N | title = The nuclear DNA base 5-hydroxymethylcytosine is present in Purkinje neurons and the brain | journal = Science | volume = 324 | issue = 5929 | pages = 929–30 | date = May 2009 | pmid = 19372393 | pmc = 3263819 | doi = 10.1126/science.1169786 | bibcode = 2009Sci...324..929K }}</ref> and the ] of uracil to produce the "J-base" in ].<ref>{{cite journal | vauthors = Ratel D, Ravanat JL, Berger F, Wion D | title = N6-methyladenine: the other methylated base of DNA | journal = BioEssays | volume = 28 | issue = 3 | pages = 309–15 | date = March 2006 | pmid = 16479578 | pmc = 2754416 | doi = 10.1002/bies.20342 }}</ref><ref>{{cite journal | vauthors = Gommers-Ampt JH, Van Leeuwen F, de Beer AL, Vliegenthart JF, Dizdaroglu M, Kowalak JA, Crain PF, Borst P | s2cid = 24801094 | title = beta-D-glucosyl-hydroxymethyluracil: a novel modified base present in the DNA of the parasitic protozoan T. brucei | journal = Cell | volume = 75 | issue = 6 | pages = 1129–36 | date = December 1993 | pmid = 8261512 | doi = 10.1016/0092-8674(93)90322-H | hdl = 1874/5219 | hdl-access = free }}</ref> | |||
| ===Base modifications=== | |||
| {{Further|]}} | |||
| The expression of genes is influenced by how the DNA is packaged in chromosomes, in a structure called ]. Base modifications can be involved in packaging, with regions that have low or no gene expression usually containing high levels of ] of ] bases. For example, cytosine methylation, produces ], which is important for ].<ref>{{cite journal |author=Klose R, Bird A |title=Genomic DNA methylation: the mark and its mediators | journal=Trends Biochem Sci |volume=31 |issue=2 | pages=89–97 |year=2006 |pmid=16403636 |doi=10.1016/j.tibs.2005.12.008}}</ref> The average level of methylation varies between organisms - the worm '']'' lacks cytosine methylation, while ]s have higher levels, with up to 1% of their DNA containing 5-methylcytosine.<ref>{{cite journal |author=Bird A |title=DNA methylation patterns and epigenetic memory | journal=Genes Dev |volume=16 |issue=1 | pages=6–21 |year=2002 |pmid=11782440 |doi=10.1101/gad.947102}}</ref> Despite the importance of 5-methylcytosine, it can ] to leave a thymine base, methylated cytosines are therefore particularly prone to ]s.<ref>{{cite journal |author=Walsh C, Xu G |title=Cytosine methylation and DNA repair | journal=Curr Top Microbiol Immunol |volume=301 | pages=283–315 |year=2006|pmid=16570853 |doi=10.1007/3-540-31390-7_11}}</ref> Other base modifications include adenine methylation in bacteria, the presence of ] in the ],<ref>{{cite journal |author=Kriaucionis S, Heintz N |title=The nuclear DNA base 5-hydroxymethylcytosine is present in Purkinje neurons and the brain |journal=Science |volume=324 |issue=5929 |pages=929–30 |year=2009 |month=May |pmid=19372393 |doi=10.1126/science.1169786}}</ref> and the ] of uracil to produce the "J-base" in ]s.<ref>{{cite journal |author=Ratel D, Ravanat J, Berger F, Wion D |title=N6-methyladenine: the other methylated base of DNA | journal=Bioessays |volume=28 |issue=3 | pages=309–15 |year=2006 |pmid=16479578 | doi = 10.1002/bies.20342 |pmc=2754416}}</ref><ref>{{cite journal |author=Gommers-Ampt J, Van Leeuwen F, de Beer A, Vliegenthart J, Dizdaroglu M, Kowalak J, Crain P, Borst P |title=beta-D-glucosyl-hydroxymethyluracil: a novel modified base present in the DNA of the parasitic protozoan T. brucei | journal=Cell |volume=75 |issue=6 | pages=1129–36 |year=1993 |pmid=8261512 |doi=10.1016/0092-8674(93)90322-H}}</ref> | |||
| ===Damage=== | === Damage === | ||
| {{further|DNA damage (naturally occurring)|Mutation|DNA damage theory of aging}} | |||
| {{Further|]}} | |||
| ] ] between a ] form of pyrene]], the major ] in ], and DNA<ref>Created from </ref>]] | ] ] between a ] form of pyrene]], the major ] in ], and DNA<ref>Created from {{Webarchive|url=https://web.archive.org/web/20080922150848/http://www.rcsb.org/pdb/cgi/explore.cgi?pdbId=1JDG |date=22 September 2008 }}</ref>]] | ||
| DNA can be damaged by many sorts of ]s, which change the DNA sequence. Mutagens include ]s, ] and also high-energy ] such as ] light and ]s. The type of DNA damage produced depends on the type of mutagen. For example, UV light can damage DNA by producing ]s, which are cross-links between pyrimidine bases.<ref>{{cite journal | |
DNA can be damaged by many sorts of ]s, which change the ]. Mutagens include ]s, ] and also high-energy ] such as ] light and ]s. The type of DNA damage produced depends on the type of mutagen. For example, UV light can damage DNA by producing ]s, which are cross-links between pyrimidine bases.<ref>{{cite journal | vauthors = Douki T, Reynaud-Angelin A, Cadet J, Sage E | title = Bipyrimidine photoproducts rather than oxidative lesions are the main type of DNA damage involved in the genotoxic effect of solar UVA radiation | journal = Biochemistry | volume = 42 | issue = 30 | pages = 9221–26 | date = August 2003 | pmid = 12885257 | doi = 10.1021/bi034593c }}</ref> On the other hand, oxidants such as ] or ] produce multiple forms of damage, including base modifications, particularly of guanosine, and double-strand breaks.<ref>{{cite journal | vauthors = Cadet J, Delatour T, Douki T, Gasparutto D, Pouget JP, Ravanat JL, Sauvaigo S | title = Hydroxyl radicals and DNA base damage | journal = Mutation Research | volume = 424 | issue = 1–2 | pages = 9–21 | date = March 1999 | pmid = 10064846 | doi = 10.1016/S0027-5107(99)00004-4 | bibcode = 1999MRFMM.424....9C }}</ref> A typical human cell contains about 150,000 bases that have suffered oxidative damage.<ref>{{cite journal | vauthors = Beckman KB, Ames BN | title = Oxidative decay of DNA | journal = The Journal of Biological Chemistry | volume = 272 | issue = 32 | pages = 19633–36 | date = August 1997 | pmid = 9289489 | doi = 10.1074/jbc.272.32.19633 | doi-access = free }}</ref> Of these oxidative lesions, the most dangerous are double-strand breaks, as these are difficult to repair and can produce ]s, ], ] from the DNA sequence, and ]s.<ref>{{cite journal | vauthors = Valerie K, Povirk LF | title = Regulation and mechanisms of mammalian double-strand break repair | journal = Oncogene | volume = 22 | issue = 37 | pages = 5792–812 | date = September 2003 | pmid = 12947387 | doi = 10.1038/sj.onc.1206679 | doi-access = free }}</ref> These mutations can cause ]. Because of inherent limits in the DNA repair mechanisms, if humans lived long enough, they would all eventually develop cancer.<ref name=Weinberg>{{cite news | url = https://www.nytimes.com/2010/12/28/health/28cancer.html | title = Unearthing Prehistoric Tumors, and Debate | newspaper = ] | date = 28 December 2010 | vauthors = Johnson G | quote = If we lived long enough, sooner or later we all would get cancer. | url-status=live | archive-url = https://web.archive.org/web/20170624233156/http://www.nytimes.com/2010/12/28/health/28cancer.html | archive-date = 24 June 2017 | df = dmy-all }}</ref><ref>{{cite book |vauthors= Alberts B, Johnson A, Lewis J |title= Molecular biology of the cell |publisher= Garland Science |location= New York |year= 2002 |edition= 4th |chapter= The Preventable Causes of Cancer |isbn= 0-8153-4072-9 |chapter-url= https://www.ncbi.nlm.nih.gov/books/NBK26897/ |quote= A certain irreducible background incidence of cancer is to be expected regardless of circumstances: mutations can never be absolutely avoided, because they are an inescapable consequence of fundamental limitations on the accuracy of DNA replication, as discussed in Chapter 5. If a human could live long enough, it is inevitable that at least one of his or her cells would eventually accumulate a set of mutations sufficient for cancer to develop. |display-authors= etal |url-status=live |archive-url= https://web.archive.org/web/20160102193148/http://www.ncbi.nlm.nih.gov/books/NBK26897/ |archive-date= 2 January 2016 |df= dmy-all }}</ref> DNA damages that are ], due to normal cellular processes that produce reactive oxygen species, the hydrolytic activities of cellular water, etc., also occur frequently. Although most of these damages are repaired, in any cell some DNA damage may remain despite the action of repair processes. These remaining DNA damages accumulate with age in mammalian postmitotic tissues. This accumulation appears to be an important underlying cause of aging.<ref>{{cite book | veditors = Kimura H, Suzuki A | title = New Research on DNA Damage | date = 2008 | publisher = Nova Science Publishers | location = New York | isbn = 978-1-60456-581-2 | vauthors = Bernstein H, Payne CM, Bernstein C, Garewal H, Dvorak K | chapter = Cancer and aging as consequences of un-repaired DNA damage | chapter-url = https://www.novapublishers.com/catalog/product_info.php?products_id=43247 | pages = 1–47 | url-status=live | archive-url = https://web.archive.org/web/20141025091740/https://www.novapublishers.com/catalog/product_info.php?products_id=43247 | archive-date = 25 October 2014 | df = dmy-all }}</ref><ref>{{cite journal | vauthors = Hoeijmakers JH | title = DNA damage, aging, and cancer | journal = The New England Journal of Medicine | volume = 361 | issue = 15 | pages = 1475–85 | date = October 2009 | pmid = 19812404 | doi = 10.1056/NEJMra0804615 }}</ref><ref>{{cite journal | vauthors = Freitas AA, de Magalhães JP | title = A review and appraisal of the DNA damage theory of ageing | journal = Mutation Research | volume = 728 | issue = 1–2 | pages = 12–22 | year = 2011 | pmid = 21600302 | doi = 10.1016/j.mrrev.2011.05.001 | bibcode = 2011MRRMR.728...12F }}</ref> | ||
| Many mutagens fit into the space between two adjacent base pairs, this is called ]. Most intercalators are ] and planar molecules; examples include ], ], and ]. |
Many mutagens fit into the space between two adjacent base pairs, this is called '']''. Most intercalators are ] and planar molecules; examples include ], ]s, ], and ]. For an intercalator to fit between base pairs, the bases must separate, distorting the DNA strands by unwinding of the double helix. This inhibits both transcription and DNA replication, causing toxicity and mutations.<ref>{{cite journal | vauthors = Ferguson LR, Denny WA | title = The genetic toxicology of acridines | journal = Mutation Research | volume = 258 | issue = 2 | pages = 123–60 | date = September 1991 | pmid = 1881402 | doi = 10.1016/0165-1110(91)90006-H }}</ref> As a result, DNA intercalators may be ]s, and in the case of thalidomide, a ].<ref>{{cite journal | vauthors = Stephens TD, Bunde CJ, Fillmore BJ | title = Mechanism of action in thalidomide teratogenesis | journal = Biochemical Pharmacology | volume = 59 | issue = 12 | pages = 1489–99 | date = June 2000 | pmid = 10799645 | doi = 10.1016/S0006-2952(99)00388-3 }}</ref> Others such as pyrene diol epoxide]] and ] form DNA adducts that induce errors in replication.<ref>{{cite journal | vauthors = Jeffrey AM | title = DNA modification by chemical carcinogens | journal = Pharmacology & Therapeutics | volume = 28 | issue = 2 | pages = 237–72 | year = 1985 | pmid = 3936066 | doi = 10.1016/0163-7258(85)90013-0 }}</ref> Nevertheless, due to their ability to inhibit DNA transcription and replication, other similar toxins are also used in ] to inhibit rapidly growing ] cells.<ref>{{cite journal | vauthors = Braña MF, Cacho M, Gradillas A, de Pascual-Teresa B, Ramos A | title = Intercalators as anticancer drugs | journal = Current Pharmaceutical Design | volume = 7 | issue = 17 | pages = 1745–80 | date = November 2001 | pmid = 11562309 | doi = 10.2174/1381612013397113 }}</ref> | ||
| ==Biological functions== | == Biological functions == | ||
| ] within the chromosomes]] | |||
| DNA usually occurs as linear ]s in eukaryotes, and circular chromosomes in prokaryotes. The set of chromosomes in a cell makes up its ]; the ] has approximately 3 billion base pairs of DNA arranged into 46 chromosomes.<ref>{{cite journal |author=Venter J |title=The sequence of the human genome | journal=Science |volume=291 |issue=5507 | pages=1304–51 |year=2001 |pmid=11181995 |doi=10.1126/science.1058040 |last2=Adams |first2=MD |last3=Myers |first3=EW |last4=Li |first4=PW |last5=Mural |first5=RJ |last6=Sutton |first6=GG |last7=Smith |first7=HO |last8=Yandell |first8=M |last9=Evans |first9=CA}}</ref> The information carried by DNA is held in the ] of pieces of DNA called ]s. ] of genetic information in genes is achieved via complementary base pairing. For example, in transcription, when a cell uses the information in a gene, the DNA sequence is copied into a complementary RNA sequence through the attraction between the DNA and the correct RNA nucleotides. Usually, this RNA copy is then used to make a matching ] in a process called ] which depends on the same interaction between RNA nucleotides. Alternatively, a cell may simply copy its genetic information in a process called DNA replication. The details of these functions are covered in other articles; here we focus on the interactions between DNA and other molecules that mediate the function of the genome. | |||
| DNA usually occurs as linear ]s in ]s, and ] in ]s. The set of chromosomes in a cell makes up its ]; the ] has approximately 3 billion base pairs of DNA arranged into 46 chromosomes.<ref name="Venter_2001" /> The information carried by DNA is held in the ] of pieces of DNA called ]s. ] of genetic information in genes is achieved via complementary base pairing. For example, in transcription, when a cell uses the information in a gene, the DNA sequence is copied into a complementary RNA sequence through the attraction between the DNA and the correct RNA nucleotides. Usually, this RNA copy is then used to make a matching ] in a process called ], which depends on the same interaction between RNA nucleotides. In an alternative fashion, a cell may copy its genetic information in a process called ]. The details of these functions are covered in other articles; here the focus is on the interactions between DNA and other molecules that mediate the function of the genome. | |||
| ===Genes and genomes=== | === Genes and genomes === | ||
| {{ |
{{further|Cell nucleus|Chromatin|Chromosome|Gene|Noncoding DNA}} | ||
| Genomic DNA is |
Genomic DNA is tightly and orderly packed in the process called ], to fit the small available volumes of the cell. In eukaryotes, DNA is located in the ], with small amounts in ] and ]s. In prokaryotes, the DNA is held within an irregularly shaped body in the cytoplasm called the ].<ref>{{cite journal | vauthors = Thanbichler M, Wang SC, Shapiro L | title = The bacterial nucleoid: a highly organized and dynamic structure | journal = Journal of Cellular Biochemistry | volume = 96 | issue = 3 | pages = 506–21 | date = October 2005 | pmid = 15988757 | doi = 10.1002/jcb.20519 | doi-access = free }}</ref> The genetic information in a genome is held within genes, and the complete set of this information in an organism is called its ]. A gene is a unit of ] and is a region of DNA that influences a particular characteristic in an organism. Genes contain an ] that can be transcribed, and ]s such as ] and ], which control transcription of the open reading frame. | ||
| In many ], only a small fraction of the total sequence of the ] encodes protein. For example, only about 1.5% of the human genome consists of protein-coding ]s, with over 50% of human DNA consisting of non-coding ].<ref>{{cite journal | |
In many ], only a small fraction of the total sequence of the ] encodes protein. For example, only about 1.5% of the human genome consists of protein-coding ]s, with over 50% of human DNA consisting of non-coding ].<ref>{{cite journal | vauthors = Wolfsberg TG, McEntyre J, Schuler GD | title = Guide to the draft human genome | journal = Nature | volume = 409 | issue = 6822 | pages = 824–26 | date = February 2001 | pmid = 11236998 | doi = 10.1038/35057000 | bibcode = 2001Natur.409..824W | url = https://zenodo.org/record/1233093 | doi-access = free }}</ref> The reasons for the presence of so much ] in eukaryotic genomes and the extraordinary differences in ], or '']'', among species, represent a long-standing puzzle known as the "]".<ref>{{cite journal | vauthors = Gregory TR | title = The C-value enigma in plants and animals: a review of parallels and an appeal for partnership | journal = Annals of Botany | volume = 95 | issue = 1 | pages = 133–46 | date = January 2005 | pmid = 15596463 | doi = 10.1093/aob/mci009 | pmc = 4246714 }}</ref> However, some DNA sequences that do not code protein may still encode functional ] molecules, which are involved in the ].<ref name="Birney_2007" /> | ||
| ] (blue) producing |
] (blue) producing an ] (green) from a DNA template (orange)<ref>{{Cite web| vauthors = Yin YW, Steitz TA |title=RCSB PDB – 1MSW: Structural basis for the transition from initiation to elongation transcription in T7 RNA polymerase|url=https://www.rcsb.org/structure/1MSW|access-date=2023-03-27|website=www.rcsb.org|language=en-US}}</ref>]] | ||
| Some non-coding DNA sequences play structural roles in chromosomes. ]s and ]s typically contain few genes, but are important for the function and stability of chromosomes.<ref name=Nugent/><ref>{{cite journal |author=Pidoux A, Allshire R |title=The role of heterochromatin in centromere function | pmc=1569473 | journal=Philos Trans R Soc Lond B Biol Sci |volume=360 |issue=1455 | pages=569–79 |year=2005 |pmid=15905142 | doi = 10.1098/rstb.2004.1611}}</ref> An abundant form of non-coding DNA in humans are ]s, which are copies of genes that have been disabled by mutation.<ref>{{cite journal |author=Harrison P, Hegyi H, Balasubramanian S, Luscombe N, Bertone P, Echols N, Johnson T, Gerstein M |title=Molecular fossils in the human genome: identification and analysis of the pseudogenes in chromosomes 21 and 22 | url=http://www.genome.org/cgi/content/full/12/2/272 | journal=Genome Res |volume=12 |issue=2 | pages=272–80 |year=2002 |pmid=11827946 |doi=10.1101/gr.207102 |pmc=155275}}</ref> These sequences are usually just molecular ]s, although they can occasionally serve as raw genetic material for the creation of new genes through the process of ] and ].<ref>{{cite journal |author=Harrison P, Gerstein M |title=Studying genomes through the aeons: protein families, pseudogenes and proteome evolution | journal=J Mol Biol |volume=318 |issue=5 | pages=1155–74 |year=2002 |pmid=12083509 |doi=10.1016/S0022-2836(02)00109-2}}</ref> | |||
| Some noncoding DNA sequences play structural roles in chromosomes. ]s and ]s typically contain few genes but are important for the function and stability of chromosomes.<ref name=Nugent /><ref>{{cite journal | vauthors = Pidoux AL, Allshire RC | title = The role of heterochromatin in centromere function | journal = Philosophical Transactions of the Royal Society of London. Series B, Biological Sciences | volume = 360 | issue = 1455 | pages = 569–79 | date = March 2005 | pmid = 15905142 | pmc = 1569473 | doi = 10.1098/rstb.2004.1611 }}</ref> An abundant form of noncoding DNA in humans are ]s, which are copies of genes that have been disabled by mutation.<ref>{{cite journal | vauthors = Harrison PM, Hegyi H, Balasubramanian S, Luscombe NM, Bertone P, Echols N, Johnson T, Gerstein M | title = Molecular fossils in the human genome: identification and analysis of the pseudogenes in chromosomes 21 and 22 | journal = Genome Research | volume = 12 | issue = 2 | pages = 272–80 | date = February 2002 | pmid = 11827946 | pmc = 155275 | doi = 10.1101/gr.207102 }}</ref> These sequences are usually just molecular ]s, although they can occasionally serve as raw ] for the creation of new genes through the process of ] and ].<ref>{{cite journal | vauthors = Harrison PM, Gerstein M | title = Studying genomes through the aeons: protein families, pseudogenes and proteome evolution | journal = Journal of Molecular Biology | volume = 318 | issue = 5 | pages = 1155–74 | date = May 2002 | pmid = 12083509 | doi = 10.1016/S0022-2836(02)00109-2 }}</ref> | |||
| ===Transcription and translation=== | |||
| {{Further|], ], ]}} | |||
| === Transcription and translation === | |||
| {{further|Genetic code|Transcription (genetics)|Protein biosynthesis}} | |||
| A gene is a sequence of DNA that contains genetic information and can influence the ] of an organism. Within a gene, the sequence of bases along a DNA strand defines a ] sequence, which then defines one or more protein sequences. The relationship between the nucleotide sequences of genes and the ] sequences of proteins is determined by the rules of ], known collectively as the ]. The genetic code consists of three-letter 'words' called ''codons'' formed from a sequence of three nucleotides (e.g. ACT, CAG, TTT). | A gene is a sequence of DNA that contains genetic information and can influence the ] of an organism. Within a gene, the sequence of bases along a DNA strand defines a ] sequence, which then defines one or more protein sequences. The relationship between the nucleotide sequences of genes and the ] sequences of proteins is determined by the rules of ], known collectively as the ]. The genetic code consists of three-letter 'words' called ''codons'' formed from a sequence of three nucleotides (e.g. ACT, CAG, TTT). | ||
| In transcription, the codons of a gene are copied into messenger RNA by ]. This RNA copy is then decoded by a ] that reads the RNA sequence by base-pairing the messenger RNA to ], which carries amino acids. Since there are 4 bases in 3-letter combinations, there are 64 possible codons (< |
In transcription, the codons of a gene are copied into messenger RNA by ]. This RNA copy is then decoded by a ] that reads the RNA sequence by base-pairing the messenger RNA to ], which carries amino acids. Since there are 4 bases in 3-letter combinations, there are 64 possible codons (4<sup>3</sup> combinations). These encode the twenty ], giving most amino acids more than one possible codon. There are also three 'stop' or 'nonsense' codons signifying the end of the coding region; these are the TAG, TAA, and TGA codons, (UAG, UAA, and UGA on the mRNA). | ||
| === Replication === | |||
| ] and ]. Next, one ] produces the ] copy. Another DNA polymerase binds to the ]. This enzyme makes discontinuous segments (called ]s) before ] joins them together.]] | |||
| {{further|DNA replication}} | |||
| ] and ]. Next, one ] produces the ] copy. Another DNA polymerase binds to the ]. This enzyme makes discontinuous segments (called ]s) before ] joins them together.]] | |||
| ] is essential for an organism to grow, but, when a cell divides, it must replicate the DNA in its genome so that the two daughter cells have the same genetic information as their parent. The double-stranded structure of DNA provides a simple mechanism for ]. Here, the two strands are separated and then each strand's ] sequence is recreated by an ] called ]. This enzyme makes the complementary strand by finding the correct base through complementary base pairing and bonding it onto the original strand. As DNA polymerases can only extend a DNA strand in a 5′ to 3′ direction, different mechanisms are used to copy the antiparallel strands of the double helix.<ref>{{cite journal | vauthors = Albà M | title = Replicative DNA polymerases | journal = Genome Biology | volume = 2 | issue = 1 | pages = REVIEWS3002 | year = 2001 | pmid = 11178285 | pmc = 150442 | doi = 10.1186/gb-2001-2-1-reviews3002 | doi-access = free }}</ref> In this way, the base on the old strand dictates which base appears on the new strand, and the cell ends up with a perfect copy of its DNA. | |||
| === Extracellular nucleic acids === | |||
| ===Replication=== | |||
| Naked extracellular DNA (eDNA), most of it released by cell death, is nearly ubiquitous in the environment. Its concentration in soil may be as high as 2 μg/L, and its concentration in natural aquatic environments may be as high at 88 μg/L.<ref name=Tani_2010>{{cite book | vauthors = Tani K, Nasu M | veditors = Kikuchi Y, Rykova EY | title = Extracellular Nucleic Acids |url=https://archive.org/details/extracellularnuc00kiku |url-access=limited |publisher=Springer |date=2010 |pages=–38 |chapter=Roles of Extracellular DNA in Bacterial Ecosystems |isbn=978-3-642-12616-1}}</ref> Various possible functions have been proposed for eDNA: it may be involved in ];<ref name="Vlassov_2007">{{cite journal | vauthors = Vlassov VV, Laktionov PP, Rykova EY | title = Extracellular nucleic acids | journal = BioEssays | volume = 29 | issue = 7 | pages = 654–67 | date = July 2007 | pmid = 17563084 | doi = 10.1002/bies.20604 | s2cid = 32463239 }}</ref> it may provide nutrients;<ref name="pmid11591672">{{cite journal | vauthors = Finkel SE, Kolter R | title = DNA as a nutrient: novel role for bacterial competence gene homologs | journal = Journal of Bacteriology | volume = 183 | issue = 21 | pages = 6288–93 | date = November 2001 | pmid = 11591672 | pmc = 100116 | doi = 10.1128/JB.183.21.6288-6293.2001 }}</ref> and it may act as a buffer to recruit or titrate ions or antibiotics.<ref name=Mulcahy_2008>{{cite journal | vauthors = Mulcahy H, Charron-Mazenod L, Lewenza S | title = Extracellular DNA chelates cations and induces antibiotic resistance in Pseudomonas aeruginosa biofilms | journal = PLOS Pathogens | volume = 4 | issue = 11 | pages = e1000213 | date = November 2008 | pmid = 19023416 | pmc = 2581603 | doi = 10.1371/journal.ppat.1000213 | doi-access = free }}</ref> Extracellular DNA acts as a functional extracellular matrix component in the ]s of several bacterial species. It may act as a recognition factor to regulate the attachment and dispersal of specific cell types in the biofilm;<ref name=Berne_2010>{{cite journal | vauthors = Berne C, Kysela DT, Brun YV | title = A bacterial extracellular DNA inhibits settling of motile progeny cells within a biofilm | journal = Molecular Microbiology | volume = 77 | issue = 4 | pages = 815–29 | date = August 2010 | pmid = 20598083 | pmc = 2962764 | doi = 10.1111/j.1365-2958.2010.07267.x }}</ref> it may contribute to biofilm formation;<ref name=Whitchurch_2002>{{cite journal | vauthors = Whitchurch CB, Tolker-Nielsen T, Ragas PC, Mattick JS | title = Extracellular DNA required for bacterial biofilm formation | journal = Science | volume = 295 | issue = 5559 | pages = 1487 | date = February 2002 | pmid = 11859186 | doi = 10.1126/science.295.5559.1487 }}</ref> and it may contribute to the biofilm's physical strength and resistance to biological stress.<ref name=Hu_2012>{{cite journal | vauthors = Hu W, Li L, Sharma S, Wang J, McHardy I, Lux R, Yang Z, He X, Gimzewski JK, Li Y, Shi W | title = DNA builds and strengthens the extracellular matrix in Myxococcus xanthus biofilms by interacting with exopolysaccharides | journal = PLOS ONE | volume = 7 | issue = 12 | pages = e51905 | year = 2012 | pmid = 23300576 | pmc = 3530553 | doi = 10.1371/journal.pone.0051905 | bibcode = 2012PLoSO...751905H | doi-access = free }}</ref> | |||
| {{Further|]}} | |||
| ] is found in the blood of the mother, and can be sequenced to determine a great deal of information about the developing fetus.<ref name="Hui_2013">{{cite journal | vauthors = Hui L, Bianchi DW | title = Recent advances in the prenatal interrogation of the human fetal genome | journal = Trends in Genetics | volume = 29 | issue = 2 | pages = 84–91 | date = February 2013 | pmid = 23158400 | pmc = 4378900 | doi = 10.1016/j.tig.2012.10.013 }}</ref> | |||
| ] is essential for an organism to grow, but when a cell divides it must replicate the DNA in its genome so that the two daughter cells have the same genetic information as their parent. The double-stranded structure of DNA provides a simple mechanism for ]. Here, the two strands are separated and then each strand's ] sequence is recreated by an ] called ]. This enzyme makes the complementary strand by finding the correct base through complementary base pairing, and bonding it onto the original strand. As DNA polymerases can only extend a DNA strand in a 5′ to 3′ direction, different mechanisms are used to copy the antiparallel strands of the double helix.<ref>{{cite journal |author=Albà M |title=Replicative DNA polymerases | journal=Genome Biol |volume=2 |issue=1 | pages=REVIEWS3002 |year=2001 |pmid=11178285 |pmc=150442 |url=http://genomebiology.com/1465-6906/2/REVIEWS3002 |doi=10.1186/gb-2001-2-1-reviews3002 |nopp=true}}</ref> In this way, the base on the old strand dictates which base appears on the new strand, and the cell ends up with a perfect copy of its DNA. | |||
| Under the name of ] eDNA has seen increased use in the natural sciences as a survey tool for ], monitoring the movements and presence of species in water, air, or on land, and assessing an area's biodiversity.<ref>{{cite journal | vauthors = Foote AD, Thomsen PF, Sveegaard S, Wahlberg M, Kielgast J, Kyhn LA, Salling AB, Galatius A, Orlando L, Gilbert MT | display-authors = 6 | title = Investigating the potential use of environmental DNA (eDNA) for genetic monitoring of marine mammals | journal = PLOS ONE | volume = 7 | issue = 8 | pages = e41781 | year = 2012 | pmid = 22952587 | pmc = 3430683 | doi = 10.1371/journal.pone.0041781 | bibcode = 2012PLoSO...741781F | doi-access = free }}</ref><ref>{{Cite web | url=https://www.the-scientist.com/news-opinion/researchers-detect-land-animals-using-dna-in-nearby-water-bodies-67481 | title=Researchers Detect Land Animals Using DNA in Nearby Water Bodies}}</ref> | |||
| ==Interactions with proteins== | |||
| === Neutrophil extracellular traps === | |||
| {{Main|Neutrophil extracellular traps}} | |||
| Neutrophil extracellular traps (NETs) are networks of extracellular fibers, primarily composed of DNA, which allow ], a type of white blood cell, to kill extracellular pathogens while minimizing damage to the host cells. | |||
| == Interactions with proteins == | |||
| All the functions of DNA depend on interactions with proteins. These ] can be non-specific, or the protein can bind specifically to a single DNA sequence. Enzymes can also bind to DNA and of these, the polymerases that copy the DNA base sequence in transcription and DNA replication are particularly important. | All the functions of DNA depend on interactions with proteins. These ] can be non-specific, or the protein can bind specifically to a single DNA sequence. Enzymes can also bind to DNA and of these, the polymerases that copy the DNA base sequence in transcription and DNA replication are particularly important. | ||
| ===DNA-binding proteins=== | === DNA-binding proteins === | ||
| {{ |
{{further|DNA-binding protein}} | ||
| ]s (in blue). These proteins' basic amino acids bind to the acidic phosphate groups on DNA.]] | |||
| <div class="thumb tleft" style="background-color: #f9f9f9; border: 1px solid #CCCCCC; margin:0.5em;"> | |||
| {|border="0" width=260 border="0" cellpadding="0" cellspacing="0" style="font-size: 85%; border: 1px solid #CCCCCC; margin: 0.3em;" | |||
| |] | |||
| |- | |||
| |] | |||
| |} | |||
| <div style="border: none; width:260px;"><div class="thumbcaption">Interaction of DNA with ]s (shown in white, top). These proteins' basic amino acids (below left, blue) bind to the acidic phosphate groups on DNA (below right, red).</div></div></div> | |||
| Structural proteins that bind DNA are well-understood examples of non-specific DNA-protein interactions. Within chromosomes, DNA is held in complexes with structural proteins. These proteins organize the DNA into a compact structure called ]. In eukaryotes this structure involves DNA binding to a complex of small basic proteins called ]s, while in prokaryotes multiple types of proteins are involved.<ref>{{cite journal | |
Structural proteins that bind DNA are well-understood examples of non-specific DNA-protein interactions. Within chromosomes, DNA is held in complexes with structural proteins. These proteins organize the DNA into a compact structure called ]. In eukaryotes, this structure involves DNA binding to a complex of small basic proteins called ]s, while in prokaryotes multiple types of proteins are involved.<ref>{{cite journal | vauthors = Sandman K, Pereira SL, Reeve JN | s2cid = 21101836 | title = Diversity of prokaryotic chromosomal proteins and the origin of the nucleosome | journal = Cellular and Molecular Life Sciences | volume = 54 | issue = 12 | pages = 1350–64 | date = December 1998 | pmid = 9893710 | doi = 10.1007/s000180050259 | pmc = 11147202 }}</ref><ref>{{cite journal | vauthors = Dame RT | title = The role of nucleoid-associated proteins in the organization and compaction of bacterial chromatin | journal = Molecular Microbiology | volume = 56 | issue = 4 | pages = 858–70 | date = May 2005 | pmid = 15853876 | doi = 10.1111/j.1365-2958.2005.04598.x | s2cid = 26965112 | doi-access = free }}</ref> The histones form a disk-shaped complex called a ], which contains two complete turns of double-stranded DNA wrapped around its surface. These non-specific interactions are formed through basic residues in the histones, making ]s to the acidic sugar-phosphate backbone of the DNA, and are thus largely independent of the base sequence.<ref>{{cite journal | vauthors = Luger K, Mäder AW, Richmond RK, Sargent DF, Richmond TJ | title = Crystal structure of the nucleosome core particle at 2.8 A resolution | journal = Nature | volume = 389 | issue = 6648 | pages = 251–60 | date = September 1997 | pmid = 9305837 | doi = 10.1038/38444 | bibcode = 1997Natur.389..251L | s2cid = 4328827 }}</ref> Chemical modifications of these basic amino acid residues include ], ], and ].<ref>{{cite journal | vauthors = Jenuwein T, Allis CD | title = Translating the histone code | journal = Science | volume = 293 | issue = 5532 | pages = 1074–80 | date = August 2001 | pmid = 11498575 | doi = 10.1126/science.1063127 | s2cid = 1883924 | url = http://www.gs.washington.edu/academics/courses/braun/55104/readings/jenuwein.pdf | url-status=live | archive-url = https://web.archive.org/web/20170808142426/http://www.gs.washington.edu/academics/courses/braun/55104/readings/jenuwein.pdf | archive-date = 8 August 2017 | df = dmy-all }}</ref> These chemical changes alter the strength of the interaction between the DNA and the histones, making the DNA more or less accessible to ]s and changing the rate of transcription.<ref>{{cite book | vauthors = Ito T | title = Protein Complexes that Modify Chromatin | chapter = Nucleosome Assembly and Remodeling | series = Current Topics in Microbiology and Immunology | volume = 274 | pages = 1–22 | year = 2003 | pmid = 12596902 | doi = 10.1007/978-3-642-55747-7_1 | isbn = 978-3-540-44208-0 }}</ref> Other non-specific DNA-binding proteins in chromatin include the high-mobility group proteins, which bind to bent or distorted DNA.<ref>{{cite journal | vauthors = Thomas JO | title = HMG1 and 2: architectural DNA-binding proteins | journal = Biochemical Society Transactions | volume = 29 | issue = Pt 4 | pages = 395–401 | date = August 2001 | pmid = 11497996 | doi = 10.1042/BST0290395 }}</ref> These proteins are important in bending arrays of nucleosomes and arranging them into the larger structures that make up chromosomes.<ref>{{cite journal | vauthors = Grosschedl R, Giese K, Pagel J | title = HMG domain proteins: architectural elements in the assembly of nucleoprotein structures | journal = Trends in Genetics | volume = 10 | issue = 3 | pages = 94–100 | date = March 1994 | pmid = 8178371 | doi = 10.1016/0168-9525(94)90232-1 }}</ref> | ||
| A distinct group of DNA-binding proteins |
A distinct group of DNA-binding proteins is the DNA-binding proteins that specifically bind single-stranded DNA. In humans, replication ] is the best-understood member of this family and is used in processes where the double helix is separated, including DNA replication, recombination, and DNA repair.<ref>{{cite journal | vauthors = Iftode C, Daniely Y, Borowiec JA | title = Replication protein A (RPA): the eukaryotic SSB | journal = Critical Reviews in Biochemistry and Molecular Biology | volume = 34 | issue = 3 | pages = 141–80 | year = 1999 | pmid = 10473346 | doi = 10.1080/10409239991209255 }}</ref> These binding proteins seem to stabilize single-stranded DNA and protect it from forming ]s or being degraded by ]s. | ||
| ] transcription factor bound to its DNA target<ref> |
] transcription factor bound to its DNA target<ref>{{Cite web| vauthors = Beamer LJ, Pabo CO |title=RCSB PDB – 1LMB: Refined 1.8 Å crystal structure of the lambda repressor-operator complex |url=https://www.rcsb.org/structure/1LMB|access-date=2023-03-27|website=www.rcsb.org|language=en-US}}</ref>]] | ||
| In contrast, other proteins have evolved to bind to particular DNA sequences. The most intensively studied of these are the various ]s, which are proteins that regulate transcription. Each transcription factor binds to one particular set of DNA sequences and activates or inhibits the transcription of genes that have these sequences close to their promoters. The transcription factors do this in two ways. Firstly, they can bind the RNA polymerase responsible for transcription, either directly or through other mediator proteins; this locates the polymerase at the promoter and allows it to begin transcription.<ref>{{cite journal | |
In contrast, other proteins have evolved to bind to particular DNA sequences. The most intensively studied of these are the various ]s, which are proteins that regulate transcription. Each transcription factor binds to one particular set of DNA sequences and activates or inhibits the transcription of genes that have these sequences close to their promoters. The transcription factors do this in two ways. Firstly, they can bind the RNA polymerase responsible for transcription, either directly or through other mediator proteins; this locates the polymerase at the promoter and allows it to begin transcription.<ref>{{cite journal | vauthors = Myers LC, Kornberg RD | title = Mediator of transcriptional regulation | journal = Annual Review of Biochemistry | volume = 69 | pages = 729–49 | year = 2000 | pmid = 10966474 | doi = 10.1146/annurev.biochem.69.1.729 }}</ref> Alternatively, transcription factors can bind ]s that modify the histones at the promoter. This changes the accessibility of the DNA template to the polymerase.<ref>{{cite journal | vauthors = Spiegelman BM, Heinrich R | title = Biological control through regulated transcriptional coactivators | journal = Cell | volume = 119 | issue = 2 | pages = 157–67 | date = October 2004 | pmid = 15479634 | doi = 10.1016/j.cell.2004.09.037 | doi-access = free }}</ref> | ||
| As these DNA targets can occur throughout an organism's genome, changes in the activity of one type of transcription factor can affect thousands of genes.<ref>{{cite journal | |
As these DNA targets can occur throughout an organism's genome, changes in the activity of one type of transcription factor can affect thousands of genes.<ref>{{cite journal | vauthors = Li Z, Van Calcar S, Qu C, Cavenee WK, Zhang MQ, Ren B | title = A global transcriptional regulatory role for c-Myc in Burkitt's lymphoma cells | journal = Proceedings of the National Academy of Sciences of the United States of America | volume = 100 | issue = 14 | pages = 8164–69 | date = July 2003 | pmid = 12808131 | pmc = 166200 | doi = 10.1073/pnas.1332764100 | bibcode = 2003PNAS..100.8164L | doi-access = free }}</ref> Consequently, these proteins are often the targets of the ] processes that control responses to environmental changes or ] and development. The specificity of these transcription factors' interactions with DNA come from the proteins making multiple contacts to the edges of the DNA bases, allowing them to "read" the DNA sequence. Most of these base-interactions are made in the major groove, where the bases are most accessible.<ref name="Pabo1984" /> | ||
| === DNA-modifying enzymes === | |||
| ] ] (green) in a complex with its substrate DNA<ref>Created from </ref>]] | |||
| ==== Nucleases and ligases ==== | |||
| ===DNA-modifying enzymes=== | |||
| ] ] (green) in a complex with its substrate DNA<ref>{{Cite web| vauthors = Kostrewa D, Winkler FK |title=RCSB PDB – 1RVA: Mg2+ binding to the active site of EcoRV endonuclease: a crystallographic study of complexes with substrate and product DNA at 2 Å resolution |url=https://www.rcsb.org/structure/1RVA|access-date=2023-03-27|website=www.rcsb.org|language=en-US}}</ref>]] | |||
| ====Nucleases and ligases==== | |||
| ]s are ]s that cut DNA strands by catalyzing the ] of the ]s. Nucleases that hydrolyse nucleotides from the ends of DNA strands are called ]s, while ]s cut within strands. The most frequently used nucleases in ] are the ], which cut DNA at specific sequences. For instance, the EcoRV enzyme shown to the left recognizes the 6-base sequence 5′- |
]s are ]s that cut DNA strands by catalyzing the ] of the ]s. Nucleases that hydrolyse nucleotides from the ends of DNA strands are called ]s, while ]s cut within strands. The most frequently used nucleases in ] are the ], which cut DNA at specific sequences. For instance, the EcoRV enzyme shown to the left recognizes the 6-base sequence 5′-GATATC-3′ and makes a cut at the horizontal line. In nature, these enzymes protect ] against ] infection by digesting the phage DNA when it enters the bacterial cell, acting as part of the ].<ref>{{cite journal | vauthors = Bickle TA, Krüger DH | title = Biology of DNA restriction | journal = Microbiological Reviews | volume = 57 | issue = 2 | pages = 434–50 | date = June 1993 | pmid = 8336674 | pmc = 372918 | doi = 10.1128/MMBR.57.2.434-450.1993 }}</ref> In technology, these sequence-specific nucleases are used in ] and ]. | ||
| Enzymes called ]s can rejoin cut or broken DNA strands.<ref name=Doherty>{{cite journal | |
Enzymes called ]s can rejoin cut or broken DNA strands.<ref name=Doherty>{{cite journal | vauthors = Doherty AJ, Suh SW | title = Structural and mechanistic conservation in DNA ligases | journal = Nucleic Acids Research | volume = 28 | issue = 21 | pages = 4051–58 | date = November 2000 | pmid = 11058099 | pmc = 113121 | doi = 10.1093/nar/28.21.4051 }}</ref> Ligases are particularly important in ] DNA replication, as they join the short segments of DNA produced at the ] into a complete copy of the DNA template. They are also used in ] and ].<ref name=Doherty /> | ||
| ====Topoisomerases and helicases==== | ==== Topoisomerases and helicases ==== | ||
| ]s are enzymes with both nuclease and ligase activity. These proteins change the amount of ] in DNA. Some of these enzymes work by cutting the DNA helix and allowing one section to rotate, thereby reducing its level of supercoiling; the enzyme then seals the DNA break.<ref name=Champoux/> Other types of these enzymes are capable of cutting one DNA helix and then passing a second strand of DNA through this break, before rejoining the helix.<ref>{{cite journal | |
]s are enzymes with both nuclease and ligase activity. These proteins change the amount of ] in DNA. Some of these enzymes work by cutting the DNA helix and allowing one section to rotate, thereby reducing its level of supercoiling; the enzyme then seals the DNA break.<ref name=Champoux /> Other types of these enzymes are capable of cutting one DNA helix and then passing a second strand of DNA through this break, before rejoining the helix.<ref>{{cite journal | vauthors = Schoeffler AJ, Berger JM | title = Recent advances in understanding structure-function relationships in the type II topoisomerase mechanism | journal = Biochemical Society Transactions | volume = 33 | issue = Pt 6 | pages = 1465–70 | date = December 2005 | pmid = 16246147 | doi = 10.1042/BST20051465 }}</ref> Topoisomerases are required for many processes involving DNA, such as DNA replication and transcription.<ref name=Wang /> | ||
| ]s are proteins that are a type of ]. They use the chemical energy in ]s, predominantly ], to break hydrogen bonds between bases and unwind the DNA double helix into single strands.<ref>{{cite journal | |
]s are proteins that are a type of ]. They use the chemical energy in ]s, predominantly ] (ATP), to break hydrogen bonds between bases and unwind the DNA double helix into single strands.<ref>{{cite journal | vauthors = Tuteja N, Tuteja R | title = Unraveling DNA helicases. Motif, structure, mechanism and function | journal = European Journal of Biochemistry | volume = 271 | issue = 10 | pages = 1849–63 | date = May 2004 | pmid = 15128295 | doi = 10.1111/j.1432-1033.2004.04094.x | url = http://repository.ias.ac.in/52775/1/40-pub.pdf | doi-access = free }}</ref> These enzymes are essential for most processes where enzymes need to access the DNA bases. | ||
| ====Polymerases==== | ==== Polymerases ==== | ||
| ]s are ]s that synthesize polynucleotide chains from ]s. The sequence of their products |
]s are ]s that synthesize polynucleotide chains from ]s. The sequence of their products is created based on existing polynucleotide chains—which are called ''templates''. These enzymes function by repeatedly adding a nucleotide to the 3′ ] group at the end of the growing polynucleotide chain. As a consequence, all polymerases work in a 5′ to 3′ direction.<ref name=Joyce>{{cite journal | vauthors = Joyce CM, Steitz TA | title = Polymerase structures and function: variations on a theme? | journal = Journal of Bacteriology | volume = 177 | issue = 22 | pages = 6321–29 | date = November 1995 | pmid = 7592405 | pmc = 177480 | doi=10.1128/jb.177.22.6321-6329.1995}}</ref> In the ] of these enzymes, the incoming nucleoside triphosphate base-pairs to the template: this allows polymerases to accurately synthesize the complementary strand of their template. Polymerases are classified according to the type of template that they use. | ||
| In DNA replication, |
In DNA replication, DNA-dependent ]s make copies of DNA polynucleotide chains. To preserve biological information, it is essential that the sequence of bases in each copy are precisely complementary to the sequence of bases in the template strand. Many DNA polymerases have a ] activity. Here, the polymerase recognizes the occasional mistakes in the synthesis reaction by the lack of base pairing between the mismatched nucleotides. If a mismatch is detected, a 3′ to 5′ ] activity is activated and the incorrect base removed.<ref>{{cite journal | vauthors = Hubscher U, Maga G, Spadari S | s2cid = 26171993 | title = Eukaryotic DNA polymerases | journal = Annual Review of Biochemistry | volume = 71 | pages = 133–63 | year = 2002 | pmid = 12045093 | doi = 10.1146/annurev.biochem.71.090501.150041 | url = http://pdfs.semanticscholar.org/e941/98efed7eb8fa606b87d9a44c118c235a62e9.pdf | archive-url = https://web.archive.org/web/20210126170051/http://pdfs.semanticscholar.org/e941/98efed7eb8fa606b87d9a44c118c235a62e9.pdf | url-status = dead | archive-date = 26 January 2021 }}</ref> In most organisms, DNA polymerases function in a large complex called the ] that contains multiple accessory subunits, such as the ] or ]s.<ref>{{cite journal | vauthors = Johnson A, O'Donnell M | title = Cellular DNA replicases: components and dynamics at the replication fork | journal = Annual Review of Biochemistry | volume = 74 | pages = 283–315 | year = 2005 | pmid = 15952889 | doi = 10.1146/annurev.biochem.73.011303.073859 }}</ref> | ||
| RNA-dependent DNA polymerases are a specialized class of polymerases that copy the sequence of an RNA strand into DNA. They include ], which is a ] enzyme involved in the infection of cells by ]es, and ], which is required for the replication of telomeres.<ref name=Greider/><ref>{{cite journal | |
RNA-dependent DNA polymerases are a specialized class of polymerases that copy the sequence of an RNA strand into DNA. They include ], which is a ] enzyme involved in the infection of cells by ]es, and ], which is required for the replication of telomeres.<ref name=Greider /><ref name=Tarrago-Litvak1994>{{cite journal | vauthors = Tarrago-Litvak L, Andréola ML, Nevinsky GA, Sarih-Cottin L, Litvak S | title = The reverse transcriptase of HIV-1: from enzymology to therapeutic intervention | journal = FASEB Journal | volume = 8 | issue = 8 | pages = 497–503 | date = May 1994 | pmid = 7514143 | doi = 10.1096/fasebj.8.8.7514143 | doi-access = free | s2cid = 39614573 }}</ref> For example, HIV reverse transcriptase is an enzyme for AIDS virus replication.<ref name=Tarrago-Litvak1994 /> Telomerase is an unusual polymerase because it contains its own RNA template as part of its structure. It synthesizes ] at the ends of chromosomes. Telomeres prevent fusion of the ends of neighboring chromosomes and protect chromosome ends from damage.<ref name=Nugent /> | ||
| Transcription is carried out by a DNA-dependent ] that copies the sequence of a DNA strand into RNA. To begin transcribing a gene, the RNA polymerase binds to a sequence of DNA called a promoter and separates the DNA strands. It then copies the gene sequence into a ] transcript until it reaches a region of DNA called the ], where it halts and detaches from the DNA. As with human DNA-dependent DNA polymerases, ], the enzyme that transcribes most of the genes in the human genome, operates as part of a large ] with multiple regulatory and accessory subunits.<ref>{{cite journal | |
Transcription is carried out by a DNA-dependent ] that copies the sequence of a DNA strand into RNA. To begin transcribing a gene, the RNA polymerase binds to a sequence of DNA called a promoter and separates the DNA strands. It then copies the gene sequence into a ] transcript until it reaches a region of DNA called the ], where it halts and detaches from the DNA. As with human DNA-dependent DNA polymerases, ], the enzyme that transcribes most of the genes in the human genome, operates as part of a large ] with multiple regulatory and accessory subunits.<ref>{{cite journal | vauthors = Martinez E | s2cid = 24946189 | title = Multi-protein complexes in eukaryotic gene transcription | journal = Plant Molecular Biology | volume = 50 | issue = 6 | pages = 925–47 | date = December 2002 | pmid = 12516863 | doi = 10.1023/A:1021258713850 }}</ref> | ||
| ==Genetic recombination== | == Genetic recombination == | ||
| {{further|Genetic recombination}} | |||
| <div class="thumb tright" style="background-color: #f9f9f9; border: 1px solid #CCCCCC; margin:0.5em;"> | |||
| <div class="thumb tright" style="background:#f9f9f9; border:1px solid #ccc; margin:0.5em;"> | |||
| {| border="0" cellpadding="0" cellspacing="0" style="width:250px; font-size:85%; border:1px solid #ccc; margin:0.3em;" | |||
| |] | |||
| |- | |||
| |] | |||
| |- | |- | ||
| |] | |] | ||
| |} | |} | ||
| <div style="border: none; width:250px;"><div class="thumbcaption">Structure of the ] intermediate in ]. The four separate DNA strands are coloured red, blue, green and yellow.<ref> |
<div style="border: none; width:250px;"><div class="thumbcaption">Structure of the ] intermediate in ]. The four separate DNA strands are coloured red, blue, green and yellow.<ref>{{Cite web| vauthors = Thorpe JH, Gale BC, Teixeira SC, Cardin CJ |title=RCSB PDB – 1M6G: Structural Characterisation of the Holliday Junction TCGGTACCGA|url=https://www.rcsb.org/structure/1M6G|access-date=2023-03-27|website=www.rcsb.org|language=en-US}}</ref></div></div></div> | ||
| ] | |||
| {{Further|]}} | |||
| ] | |||
| A DNA helix usually does not interact with other segments of DNA, and in human cells the different chromosomes even occupy separate areas in the nucleus called "chromosome territories".<ref>{{cite journal | |
A DNA helix usually does not interact with other segments of DNA, and in human cells, the different chromosomes even occupy separate areas in the nucleus called "]".<ref>{{cite journal | vauthors = Cremer T, Cremer C | title = Chromosome territories, nuclear architecture and gene regulation in mammalian cells | journal = Nature Reviews Genetics | volume = 2 | issue = 4 | pages = 292–301 | date = April 2001 | pmid = 11283701 | doi = 10.1038/35066075 | s2cid = 8547149 }}</ref> This physical separation of different chromosomes is important for the ability of DNA to function as a stable repository for information, as one of the few times chromosomes interact is in ] which occurs during ], when ] occurs. Chromosomal crossover is when two DNA helices break, swap a section and then rejoin. | ||
| Recombination allows chromosomes to exchange genetic information and produces new combinations of genes, which increases the efficiency of ] and can be important in the rapid evolution of new proteins.<ref>{{cite journal | |
Recombination allows chromosomes to exchange genetic information and produces new combinations of genes, which increases the efficiency of ] and can be important in the rapid evolution of new proteins.<ref>{{cite journal | vauthors = Pál C, Papp B, Lercher MJ | title = An integrated view of protein evolution | journal = Nature Reviews Genetics | volume = 7 | issue = 5 | pages = 337–48 | date = May 2006 | pmid = 16619049 | doi = 10.1038/nrg1838 | s2cid = 23225873 }}</ref> Genetic recombination can also be involved in DNA repair, particularly in the cell's response to double-strand breaks.<ref>{{cite journal | vauthors = O'Driscoll M, Jeggo PA | title = The role of double-strand break repair – insights from human genetics | journal = Nature Reviews Genetics | volume = 7 | issue = 1 | pages = 45–54 | date = January 2006 | pmid = 16369571 | doi = 10.1038/nrg1746 | s2cid = 7779574 }}</ref> | ||
| The most common form of chromosomal crossover is ], where the two chromosomes involved share very similar sequences. Non-homologous recombination can be damaging to cells, as it can produce ]s and genetic abnormalities. The recombination reaction is catalyzed by enzymes known as ]s, such as ].<ref>{{cite journal | |
The most common form of chromosomal crossover is ], where the two chromosomes involved share very similar sequences. ] can be damaging to cells, as it can produce ]s and genetic abnormalities. The recombination reaction is catalyzed by enzymes known as ]s, such as ].<ref>{{cite journal | vauthors = Vispé S, Defais M | title = Mammalian Rad51 protein: a RecA homologue with pleiotropic functions | journal = Biochimie | volume = 79 | issue = 9–10 | pages = 587–92 | date = October 1997 | pmid = 9466696 | doi = 10.1016/S0300-9084(97)82007-X }}</ref> The first step in recombination is a double-stranded break caused by either an ] or damage to the DNA.<ref>{{cite journal | vauthors = Neale MJ, Keeney S | title = Clarifying the mechanics of DNA strand exchange in meiotic recombination | journal = Nature | volume = 442 | issue = 7099 | pages = 153–58 | date = July 2006 | pmid = 16838012 | doi = 10.1038/nature04885 | bibcode = 2006Natur.442..153N | pmc = 5607947 }}</ref> A series of steps catalyzed in part by the recombinase then leads to joining of the two helices by at least one ], in which a segment of a single strand in each helix is annealed to the complementary strand in the other helix. The Holliday junction is a tetrahedral junction structure that can be moved along the pair of chromosomes, swapping one strand for another. The recombination reaction is then halted by cleavage of the junction and re-ligation of the released DNA.<ref>{{cite journal | vauthors = Dickman MJ, Ingleston SM, Sedelnikova SE, Rafferty JB, Lloyd RG, Grasby JA, Hornby DP | s2cid = 39505263 | title = The RuvABC resolvasome | journal = European Journal of Biochemistry | volume = 269 | issue = 22 | pages = 5492–501 | date = November 2002 | pmid = 12423347 | doi = 10.1046/j.1432-1033.2002.03250.x | doi-access = free }}</ref> Only strands of like polarity exchange DNA during recombination. There are two types of cleavage: east-west cleavage and north–south cleavage. The north–south cleavage nicks both strands of DNA, while the east–west cleavage has one strand of DNA intact. The formation of a Holliday junction during recombination makes it possible for genetic diversity, genes to exchange on chromosomes, and expression of wild-type viral genomes. | ||
| ==Evolution== | == Evolution == | ||
| {{ |
{{further|RNA world hypothesis}} | ||
| DNA contains the genetic information that allows all |
DNA contains the genetic information that allows all forms of life to function, grow and reproduce. However, it is unclear how long in the 4-billion-year ] DNA has performed this function, as it has been proposed that the earliest forms of life may have used RNA as their genetic material.<ref name="Joyce-2002">{{cite journal | vauthors = Joyce GF | title = The antiquity of RNA-based evolution | journal = Nature | volume = 418 | issue = 6894 | pages = 214–21 | date = July 2002 | pmid = 12110897 | doi = 10.1038/418214a | bibcode = 2002Natur.418..214J | s2cid = 4331004 }}</ref><ref>{{cite journal | vauthors = Orgel LE | title = Prebiotic chemistry and the origin of the RNA world | journal = Critical Reviews in Biochemistry and Molecular Biology | volume = 39 | issue = 2 | pages = 99–123 | year = 2004 | pmid = 15217990 | doi = 10.1080/10409230490460765 | citeseerx = 10.1.1.537.7679 | s2cid = 4939632 }}</ref> RNA may have acted as the central part of early ] as it can both transmit genetic information and carry out ] as part of ]s.<ref>{{cite journal | vauthors = Davenport RJ | s2cid = 85976762 | title = Ribozymes. Making copies in the RNA world | journal = Science | volume = 292 | issue = 5520 | pages = 1278a–1278 | date = May 2001 | pmid = 11360970 | doi = 10.1126/science.292.5520.1278a }}</ref> This ancient ] where nucleic acid would have been used for both catalysis and genetics may have influenced the ] of the current genetic code based on four nucleotide bases. This would occur, since the number of different bases in such an organism is a trade-off between a small number of bases increasing replication accuracy and a large number of bases increasing the catalytic efficiency of ribozymes.<ref>{{cite journal | vauthors = Szathmáry E | title = What is the optimum size for the genetic alphabet? | journal = Proceedings of the National Academy of Sciences of the United States of America | volume = 89 | issue = 7 | pages = 2614–18 | date = April 1992 | pmid = 1372984 | pmc = 48712 | doi = 10.1073/pnas.89.7.2614 | bibcode = 1992PNAS...89.2614S | doi-access = free }}</ref> However, there is no direct evidence of ancient genetic systems, as recovery of DNA from most fossils is impossible because DNA survives in the environment for less than one million years, and slowly degrades into short fragments in solution.<ref>{{cite journal | vauthors = Lindahl T | title = Instability and decay of the primary structure of DNA | journal = Nature | volume = 362 | issue = 6422 | pages = 709–15 | date = April 1993 | pmid = 8469282 | doi = 10.1038/362709a0 | bibcode = 1993Natur.362..709L | s2cid = 4283694 }}</ref> Claims for older DNA have been made, most notably a report of the isolation of a viable bacterium from a salt crystal 250 million years old,<ref>{{cite journal | vauthors = Vreeland RH, Rosenzweig WD, Powers DW | title = Isolation of a 250 million-year-old halotolerant bacterium from a primary salt crystal | journal = Nature | volume = 407 | issue = 6806 | pages = 897–900 | date = October 2000 | pmid = 11057666 | doi = 10.1038/35038060 | bibcode = 2000Natur.407..897V | s2cid = 9879073 }}</ref> but these claims are controversial.<ref>{{cite journal | vauthors = Hebsgaard MB, Phillips MJ, Willerslev E | title = Geologically ancient DNA: fact or artefact? | journal = Trends in Microbiology | volume = 13 | issue = 5 | pages = 212–20 | date = May 2005 | pmid = 15866038 | doi = 10.1016/j.tim.2005.03.010 }}</ref><ref>{{cite journal | vauthors = Nickle DC, Learn GH, Rain MW, Mullins JI, Mittler JE | title = Curiously modern DNA for a "250 million-year-old" bacterium | journal = Journal of Molecular Evolution | volume = 54 | issue = 1 | pages = 134–37 | date = January 2002 | pmid = 11734907 | doi = 10.1007/s00239-001-0025-x | bibcode = 2002JMolE..54..134N | s2cid = 24740859 }}</ref> | ||
| Building blocks of DNA (], ], and related ]) may have been formed extraterrestrially in ].<ref name="Callahan">{{cite journal | vauthors = Callahan MP, Smith KE, Cleaves HJ, Ruzicka J, Stern JC, Glavin DP, House CH, Dworkin JP | title = Carbonaceous meteorites contain a wide range of extraterrestrial nucleobases | journal = Proceedings of the National Academy of Sciences of the United States of America | volume = 108 | issue = 34 | pages = 13995–98 | date = August 2011 | pmid = 21836052 | pmc = 3161613 | doi = 10.1073/pnas.1106493108 | bibcode = 2011PNAS..10813995C | doi-access = free }}</ref><ref name="Steigerwald">{{cite web | vauthors = Steigerwald J |title=NASA Researchers: DNA Building Blocks Can Be Made in Space |url=http://www.nasa.gov/topics/solarsystem/features/dna-meteorites.html |publisher=] |date=8 August 2011 |access-date=10 August 2011 |url-status=live |archive-url=https://web.archive.org/web/20150623004556/http://www.nasa.gov/topics/solarsystem/features/dna-meteorites.html |archive-date=23 June 2015 }}</ref><ref name="DNA">{{cite web |author=ScienceDaily Staff |title=DNA Building Blocks Can Be Made in Space, NASA Evidence Suggests |url=https://www.sciencedaily.com/releases/2011/08/110808220659.htm |date=9 August 2011 |website=] |access-date=9 August 2011 |url-status=live |archive-url=https://web.archive.org/web/20110905105043/https://www.sciencedaily.com/releases/2011/08/110808220659.htm |archive-date=5 September 2011 }}</ref> Complex DNA and ] ]s of ], including ], ], and ], have also been formed in the laboratory under conditions mimicking those found in ], using starting chemicals, such as ], found in ]s. Pyrimidine, like ] (PAHs), the most carbon-rich chemical found in the ], may have been formed in ]s or in interstellar ] and gas clouds.<ref name="NASA-20150303">{{cite web | vauthors = Marlaire R |title=NASA Ames Reproduces the Building Blocks of Life in Laboratory |url=http://www.nasa.gov/content/nasa-ames-reproduces-the-building-blocks-of-life-in-laboratory |date=3 March 2015 |work=] |access-date=5 March 2015 |url-status=live |archive-url=https://web.archive.org/web/20150305083306/http://www.nasa.gov/content/nasa-ames-reproduces-the-building-blocks-of-life-in-laboratory/ |archive-date=5 March 2015 }}</ref> | |||
| Unfortunately, there is no direct evidence of ancient genetic systems, as recovery of DNA from most fossils is impossible. This is because DNA will survive in the environment for less than one million years and slowly degrades into short fragments in solution.<ref>{{cite journal |author=Lindahl T |title=Instability and decay of the primary structure of DNA |journal=Nature |volume=362 |issue=6422 |pages=709–15 |year=1993 |pmid=8469282 | doi = 10.1038/362709a0}}</ref> Claims for older DNA have been made, most notably a report of the isolation of a viable bacterium from a salt crystal 250 million years old,<ref>{{cite journal |author=Vreeland R, Rosenzweig W, Powers D |title=Isolation of a 250 million-year-old halotolerant bacterium from a primary salt crystal |journal=Nature |volume=407 |issue=6806 |pages=897–900 |year=2000 |pmid=11057666 | doi = 10.1038/35038060}}</ref> but these claims are controversial.<ref>{{cite journal |author=Hebsgaard M, Phillips M, Willerslev E |title=Geologically ancient DNA: fact or artefact? |journal=Trends Microbiol |volume=13 |issue=5 |pages=212–20 |year=2005 |pmid=15866038 |doi=10.1016/j.tim.2005.03.010}}</ref><ref>{{cite journal |author=Nickle D, Learn G, Rain M, Mullins J, Mittler J |title=Curiously modern DNA for a "250 million-year-old" bacterium |journal=J Mol Evol |volume=54 |issue=1 |pages=134–7 |year=2002 |pmid=11734907 | doi = 10.1007/s00239-001-0025-x}}</ref> | |||
| ] has been recovered from ancient organisms at a timescale where genome evolution can be directly observed, including from extinct organisms up to millions of years old, such as the ].<ref name="CNN-20210217">{{cite news | vauthors = Hunt K |title=World's oldest DNA sequenced from a mammoth that lived more than a million years ago |url=https://www.cnn.com/2021/02/17/world/mammoth-oldest-dna-million-years-ago-scn/index.html |date=17 February 2021 |work=] |access-date=17 February 2021 }}</ref><ref name="NAT-20210217">{{cite journal | vauthors = Callaway E | title = Million-year-old mammoth genomes shatter record for oldest ancient DNA – Permafrost-preserved teeth, up to 1.6 million years old, identify a new kind of mammoth in Siberia. |date=17 February 2021 |journal=] |volume=590 |issue=7847 |pages=537–538 |doi=10.1038/d41586-021-00436-x |issn=0028-0836 |pmid=33597786 | bibcode = 2021Natur.590..537C |doi-access=free }}</ref> | |||
| ==Uses in technology== | |||
| ===Genetic engineering=== | |||
| {{Further|], ] and ]}} | |||
| == Uses in technology == | |||
| Methods have been developed to purify DNA from organisms, such as ] and manipulate it in the laboratory, such as ]s and the ]. Modern ] and ] make intensive use of these techniques in recombinant DNA technology. ] is a man-made DNA sequence that has been assembled from other DNA sequences. They can be ] into organisms in the form of ]s or in the appropriate format, by using a ].<ref>{{cite journal |author=Goff SP, Berg P |title=Construction of hybrid viruses containing SV40 and lambda phage DNA segments and their propagation in cultured monkey cells |journal=Cell |volume=9 |issue=4 PT 2 |pages=695–705 |year=1976 |pmid=189942 |doi=10.1016/0092-8674(76)90133-1}}</ref> The ] organisms produced can be used to produce products such as recombinant ]s, used in ],<ref>{{cite journal |author=Houdebine L |title=Transgenic animal models in biomedical research |journal=Methods Mol Biol |volume=360 |pages=163–202 |pmid=17172731 |year=2007 |doi=10.1385/1-59745-165-7:163}}</ref> or be grown in ].<ref>{{cite journal |author=Daniell H, Dhingra A |title=Multigene engineering: dawn of an exciting new era in biotechnology |journal=Curr Opin Biotechnol |volume=13 |issue=2 |pages=136–41 |year=2002 |pmid=11950565 |doi=10.1016/S0958-1669(02)00297-5}}</ref><ref>{{cite journal |author=Job D |title=Plant biotechnology in agriculture |journal=Biochimie |volume=84 |issue=11 |pages=1105–10 |year=2002 |pmid=12595138 |doi=10.1016/S0300-9084(02)00013-5}}</ref> | |||
| === |
=== Genetic engineering === | ||
| {{further|Molecular biology|Nucleic acid methods|Genetic engineering}} | |||
| {{Further|]}} | |||
| Methods have been developed to purify DNA from organisms, such as ], and to manipulate it in the laboratory, such as ]s and the ]. Modern ] and ] make intensive use of these techniques in recombinant DNA technology. ] is a man-made DNA sequence that has been assembled from other DNA sequences. They can be ] into organisms in the form of ]s or in the appropriate format, by using a ].<ref>{{cite journal | vauthors = Goff SP, Berg P | s2cid = 41788896 | title = Construction of hybrid viruses containing SV40 and lambda phage DNA segments and their propagation in cultured monkey cells | journal = Cell | volume = 9 | issue = 4 PT 2 | pages = 695–705 | date = December 1976 | pmid = 189942 | doi = 10.1016/0092-8674(76)90133-1 }}</ref> The ] organisms produced can be used to produce products such as recombinant ]s, used in ],<ref>{{cite book | vauthors = Houdebine LM | title = Target Discovery and Validation Reviews and Protocols | chapter = Transgenic animal models in biomedical research | series = Methods in Molecular Biology | volume = 360 | pages = 163–202 | year = 2007 | pmid = 17172731 | doi = 10.1385/1-59745-165-7:163 | isbn = 978-1-59745-165-9 }}</ref> or be grown in ].<ref>{{cite journal | vauthors = Daniell H, Dhingra A | title = Multigene engineering: dawn of an exciting new era in biotechnology | journal = Current Opinion in Biotechnology | volume = 13 | issue = 2 | pages = 136–41 | date = April 2002 | pmid = 11950565 | pmc = 3481857 | doi = 10.1016/S0958-1669(02)00297-5 }}</ref><ref>{{cite journal | vauthors = Job D | title = Plant biotechnology in agriculture | journal = Biochimie | volume = 84 | issue = 11 | pages = 1105–10 | date = November 2002 | pmid = 12595138 | doi = 10.1016/S0300-9084(02)00013-5 }}</ref> | |||
| === DNA profiling === | |||
| ] can use DNA in ], ], ], ] or ] found at a ] to identify a matching DNA of an individual, such as a perpetrator. This process is called genetic fingerprinting, or more accurately, ]. In DNA profiling, the lengths of variable sections of repetitive DNA, such as ]s and ]s, are compared between people. This method is usually an extremely reliable technique for identifying a matching DNA.<ref>{{cite journal |author=Collins A, Morton N |title=Likelihood ratios for DNA identification | url=http://www.pnas.org/cgi/reprint/91/13/6007.pdf | journal=Proc Natl Acad Sci USA |volume=91 |issue=13 | pages=6007–11 |year=1994 |pmid=8016106 |doi=10.1073/pnas.91.13.6007|format=PDF |pmc=44126}}</ref> However, identification can be complicated if the scene is contaminated with DNA from several people.<ref>{{cite journal |author=Weir B, Triggs C, Starling L, Stowell L, Walsh K, Buckleton J |title=Interpreting DNA mixtures | journal=J Forensic Sci |volume=42 |issue=2 | pages=213–22 |year=1997 |pmid=9068179}}</ref> DNA profiling was developed in 1984 by British geneticist Sir ],<ref>{{cite journal |author=Jeffreys A, Wilson V, Thein S |title=Individual-specific 'fingerprints' of human DNA | journal=Nature |volume=316 |issue=6023 | pages=76–9 |year=1985|pmid=2989708 |doi=10.1038/316076a0}}</ref> and first used in forensic science to convict Colin Pitchfork in the 1988 ] case.<ref> Forensic Science Service Accessed 23 December 2006</ref> | |||
| {{further|DNA profiling}} | |||
| ] can use DNA in ], ], ], ] or ] found at a ] to identify a matching DNA of an individual, such as a perpetrator.<ref>{{Cite news|url=https://theconversation.com/from-the-crime-scene-to-the-courtroom-the-journey-of-a-dna-sample-82250|title=From the crime scene to the courtroom: the journey of a DNA sample| vauthors = Curtis C, Hereward J |date=29 August 2017 |work=The Conversation |access-date=22 October 2017 |archive-url=https://web.archive.org/web/20171022033110/http://theconversation.com/from-the-crime-scene-to-the-courtroom-the-journey-of-a-dna-sample-82250 |archive-date=22 October 2017 |url-status=live }}</ref> This process is formally termed ], also called ''DNA fingerprinting''. In DNA profiling, the lengths of variable sections of repetitive DNA, such as ]s and ]s, are compared between people. This method is usually an extremely reliable technique for identifying a matching DNA.<ref>{{cite journal | vauthors = Collins A, Morton NE | title = Likelihood ratios for DNA identification | journal = Proceedings of the National Academy of Sciences of the United States of America | volume = 91 | issue = 13 | pages = 6007–11 | date = June 1994 | pmid = 8016106 | pmc = 44126 | doi = 10.1073/pnas.91.13.6007 | bibcode = 1994PNAS...91.6007C | doi-access = free }}</ref> However, identification can be complicated if the scene is contaminated with DNA from several people.<ref>{{cite journal | vauthors = Weir BS, Triggs CM, Starling L, Stowell LI, Walsh KA, Buckleton J | title = Interpreting DNA mixtures | journal = Journal of Forensic Sciences | volume = 42 | issue = 2 | pages = 213–22 | date = March 1997 | doi = 10.1520/JFS14100J | pmid = 9068179 | s2cid = 14511630 }}</ref> DNA profiling was developed in 1984 by British geneticist Sir ],<ref>{{cite journal | vauthors = Jeffreys AJ, Wilson V, Thein SL | title = Individual-specific 'fingerprints' of human DNA | journal = Nature | volume = 316 | issue = 6023 | pages = 76–79 | year = 1985 | pmid = 2989708 | doi = 10.1038/316076a0 | bibcode = 1985Natur.316...76J | s2cid = 4229883 | doi-access = free }}</ref> and first used in forensic science to convict Colin Pitchfork in the 1988 ] case.<ref>{{Cite web|date=2006-12-14|title=Colin Pitchfork|url=http://www.forensic.gov.uk/forensic_t/inside/news/list_casefiles.php?case=1|access-date=2023-03-27|archive-url=https://web.archive.org/web/20061214004903/http://www.forensic.gov.uk/forensic_t/inside/news/list_casefiles.php?case=1 |archive-date=14 December 2006 }}</ref> | |||
| People convicted of certain types of crimes may be required to provide a sample of DNA for a database. This has helped investigators solve old cases where only a DNA sample was obtained from the scene. DNA profiling can also be used to identify victims of mass casualty incidents.<ref>{{cite web |url=http://massfatality.dna.gov/Introduction/ |title=DNA Identification in Mass Fatality Incidents |month=September | year=2006 |publisher=National Institute of Justice}}</ref> On the other hand, many convicted people have been released from prison on the basis of DNA techniques, which were not available when a crime had originally been committed. | |||
| The development of forensic science and the ability to now obtain genetic matching on minute samples of blood, skin, saliva, or hair has led to re-examining many cases. Evidence can now be uncovered that was scientifically impossible at the time of the original examination. Combined with the removal of the ] law in some places, this can allow cases to be reopened where prior trials have failed to produce sufficient evidence to convince a jury. People charged with serious crimes may be required to provide a sample of DNA for matching purposes. The most obvious defense to DNA matches obtained forensically is to claim that cross-contamination of evidence has occurred. This has resulted in meticulous strict handling procedures with new cases of serious crime. | |||
| ===Bioinformatics=== | |||
| {{Further|]}} | |||
| ] involves the manipulation, searching, and ] of DNA sequence data. The development of techniques to store and search DNA sequences have led to widely applied advances in ], especially ]s, ] and ].<ref>{{Cite book | last1=Baldi|first1= Pierre|author1-link=Pierre Baldi|last2= Brunak|first2= Soren | title=Bioinformatics: The Machine Learning Approach | publisher= MIT Press | year=2001| isbn=978-0-262-02506-5 | oclc=45951728 | author=Pierre Baldi; Søren Brunak.}}.</ref> String searching or matching algorithms, which find an occurrence of a sequence of letters inside a larger sequence of letters, were developed to search for specific sequences of nucleotides.<ref>Gusfield, Dan. ''Algorithms on Strings, Trees, and Sequences: Computer Science and Computational Biology''. ], 15 January 1997. ISBN 978-0-521-58519-4.</ref> In other applications such as ]s, even simple algorithms for this problem usually suffice, but DNA sequences cause these algorithms to exhibit near-worst-case behaviour due to their small number of distinct characters. The related problem of ] aims to identify ] sequences and locate the specific ]s that make them distinct. These techniques, especially ], are used in studying ] relationships and protein function.<ref>{{cite journal |author=Sjölander K |title=Phylogenomic inference of protein molecular function: advances and challenges | url=http://bioinformatics.oxfordjournals.org/cgi/reprint/20/2/170 | journal=Bioinformatics |volume=20 |issue=2 | pages=170–9 |year=2004 |pmid=14734307 | doi = 10.1093/bioinformatics/bth021}}</ref> Data sets representing entire genomes' worth of DNA sequences, such as those produced by the ], are difficult to use without annotations, which label the locations of genes and regulatory elements on each chromosome. Regions of DNA sequence that have the characteristic patterns associated with protein- or RNA-coding genes can be identified by ] algorithms, which allow researchers to predict the presence of particular ]s in an organism even before they have been isolated experimentally.<ref name="Mount">{{cite book|author = Mount DM |title=Bioinformatics: Sequence and Genome Analysis | edition = 2 | publisher = Cold Spring Harbor Laboratory Press | year = 2004 | isbn = 0879697121|oclc = 55106399|location=Cold Spring Harbor, NY}}</ref> | |||
| DNA profiling is also used successfully to positively identify victims of mass casualty incidents,<ref>{{cite web|url=http://massfatality.dna.gov/Introduction/ |title=DNA Identification in Mass Fatality Incidents |date=September 2006 |publisher=National Institute of Justice |url-status=dead |archive-url=https://web.archive.org/web/20061112000837/http://massfatality.dna.gov/Introduction/ |archive-date=12 November 2006 }}</ref> bodies or body parts in serious accidents, and individual victims in mass war graves, via matching to family members. | |||
| ===DNA nanotechnology=== | |||
| ] at right. ] is the field which seeks to design nanoscale structures using the ] properties of DNA molecules. Image from {{doi-inline|10.1371/journal.pbio.0020073|Strong, 2004}}.]] | |||
| DNA profiling is also used in ] to determine if someone is the biological parent or grandparent of a child with the probability of parentage is typically 99.99% when the alleged parent is biologically related to the child. Normal ] methods happen after birth, but there are new methods to test paternity while a mother is still pregnant.<ref>{{Cite news| vauthors = Pollack A |date=2012-06-19|title=Before Birth, Dad's ID|language=en-US|work=The New York Times|url=https://www.nytimes.com/2012/06/20/health/paternity-blood-tests-that-work-early-in-a-pregnancy.html|access-date=2023-03-27|issn=0362-4331|archive-url=https://web.archive.org/web/20170624231639/http://www.nytimes.com/2012/06/20/health/paternity-blood-tests-that-work-early-in-a-pregnancy.html|archive-date=2017-06-24|url-status=live}}</ref> | |||
| {{Further|]}} | |||
| DNA nanotechnology uses the unique ] properties of DNA and other nucleic acids to create self-assembling branched DNA complexes with useful properties.<ref>{{cite journal |author=Rothemund PW |title=Folding DNA to create nanoscale shapes and patterns |journal=Nature |volume=440 |issue=7082 |pages=297–302 |year=2006 |month=March |pmid=16541064 |doi=10.1038/nature04586}}</ref> DNA is thus used as a structural material rather than as a carrier of biological information. This has led to the creation of two-dimensional periodic lattices (both tile-based as well as using the "]" method) as well as three-dimensional structures in the shapes of ].<ref>{{cite journal |author=Andersen ES, Dong M, Nielsen MM |title=Self-assembly of a nanoscale DNA box with a controllable lid |journal=Nature |volume=459 |issue=7243 |pages=73–6 |year=2009 |month=May |pmid=19424153 |doi=10.1038/nature07971}}</ref> ] and ] have also been demonstrated,<ref>{{cite journal |author=Ishitsuka Y, Ha T |title=DNA nanotechnology: a nanomachine goes live |journal=Nat Nanotechnol |volume=4 |issue=5 |pages=281–2 |year=2009 |month=May |pmid=19421208 |doi=10.1038/nnano.2009.101}}</ref> and these DNA structures have been used to template the arrangement of other molecules such as ] and ] proteins.<ref>{{cite journal |author=Aldaye FA, Palmer AL, Sleiman HF |title=Assembling materials with DNA as the guide |journal=Science |volume=321 |issue=5897 |pages=1795–9 |year=2008 |month=September |pmid=18818351 |doi=10.1126/science.1154533}}</ref> | |||
| === DNA enzymes or catalytic DNA === | |||
| ===History and anthropology=== | |||
| {{further|Deoxyribozyme}} | |||
| {{Further|] and ]}} | |||
| ]s, also called DNAzymes or catalytic DNA, were first discovered in 1994.<ref name="Breaker 223–229">{{cite journal | vauthors = Breaker RR, Joyce GF | title = A DNA enzyme that cleaves RNA | journal = Chemistry & Biology | volume = 1 | issue = 4 | pages = 223–29 | date = December 1994 | pmid = 9383394 | doi = 10.1016/1074-5521(94)90014-0 }}</ref> They are mostly single stranded DNA sequences isolated from a large pool of random DNA sequences through a combinatorial approach called ] selection or ] (SELEX). DNAzymes catalyze variety of chemical reactions including RNA-DNA cleavage, RNA-DNA ligation, amino acids phosphorylation-dephosphorylation, carbon-carbon bond formation, etc. DNAzymes can enhance catalytic rate of chemical reactions up to 100,000,000,000-fold over the uncatalyzed reaction.<ref>{{cite journal | vauthors = Chandra M, Sachdeva A, Silverman SK | title = DNA-catalyzed sequence-specific hydrolysis of DNA | journal = Nature Chemical Biology | volume = 5 | issue = 10 | pages = 718–20 | date = October 2009 | pmid = 19684594 | pmc = 2746877 | doi = 10.1038/nchembio.201 }}</ref> The most extensively studied class of DNAzymes is RNA-cleaving types which have been used to detect different metal ions and designing therapeutic agents. Several metal-specific DNAzymes have been reported including the GR-5 DNAzyme (lead-specific),<ref name="Breaker 223–229" /> the CA1-3 DNAzymes (copper-specific),<ref>{{cite journal | vauthors = Carmi N, Shultz LA, Breaker RR | title = In vitro selection of self-cleaving DNAs | journal = Chemistry & Biology | volume = 3 | issue = 12 | pages = 1039–46 | date = December 1996 | pmid = 9000012 | doi = 10.1016/S1074-5521(96)90170-2 | doi-access = free }}</ref> the 39E DNAzyme (uranyl-specific) and the NaA43 DNAzyme (sodium-specific).<ref>{{cite journal | vauthors = Torabi SF, Wu P, McGhee CE, Chen L, Hwang K, Zheng N, Cheng J, Lu Y | title = In vitro selection of a sodium-specific DNAzyme and its application in intracellular sensing | journal = Proceedings of the National Academy of Sciences of the United States of America | volume = 112 | issue = 19 | pages = 5903–08 | date = May 2015 | pmid = 25918425 | pmc = 4434688 | doi = 10.1073/pnas.1420361112 | bibcode = 2015PNAS..112.5903T | doi-access = free }}</ref> The NaA43 DNAzyme, which is reported to be more than 10,000-fold selective for sodium over other metal ions, was used to make a real-time sodium sensor in cells. | |||
| Because DNA collects mutations over time, which are then inherited, it contains historical information and by comparing DNA sequences, geneticists can infer the evolutionary history of organisms, their ].<ref>{{cite journal |author=Wray G |title=Dating branches on the tree of life using DNA | journal=Genome Biol |volume=3 |issue=1 | pages=REVIEWS0001 |year=2002 |pmid=11806830 |pmc=150454 |doi=10.1046/j.1525-142X.1999.99010.x |url=http://genomebiology.com/1465-6906/3/REVIEWS0001 |nopp=true |last2=Martindale |first2=Mark Q.}}</ref> This field of phylogenetics is a powerful tool in ]. If DNA sequences within a species are compared, ] can learn the history of particular populations. This can be used in studies ranging from ] to ]; for example, DNA evidence is being used to try to identify the ].<ref>''Lost Tribes of Israel'', ], PBS airdate: 22 February 2000. Transcript available from (last accessed on 4 March 2006)</ref><ref>Kleiman, Yaakov. ''aish.com'' (January 13, 2000). Accessed 4 March 2006.</ref> | |||
| === Bioinformatics === | |||
| DNA has also been used to look at modern family relationships, such as establishing family relationships between the descendants of ] and ]. This usage is closely related to the use of DNA in criminal investigations detailed above. Indeed, some criminal investigations have been solved when DNA from crime scenes has matched relatives of the guilty individual.<ref>Bhattacharya, Shaoni. ''newscientist.com'' (20 April 2004). Accessed 22 December 06</ref> | |||
| {{further|Bioinformatics}} | |||
| ] involves the development of techniques to store, ], search and manipulate biological data, including DNA ] data. These have led to widely applied advances in ], especially ]s, ], and ].<ref>{{cite book | vauthors = Baldi P, Brunak S |author1-link=Pierre Baldi |title=Bioinformatics: The Machine Learning Approach |publisher= MIT Press |year=2001| isbn=978-0-262-02506-5 |oclc=45951728}}</ref> String searching or matching algorithms, which find an occurrence of a sequence of letters inside a larger sequence of letters, were developed to search for specific sequences of nucleotides.<ref>{{cite book | vauthors = Gusfield D | title = Algorithms on Strings, Trees, and Sequences: Computer Science and Computational Biology | publisher = ] | date = 15 January 1997 | isbn = 978-0-521-58519-4 }}</ref> The DNA sequence may be ] with other DNA sequences to identify ] and locate the specific ]s that make them distinct. These techniques, especially ], are used in studying ] relationships and protein function.<ref>{{cite journal | vauthors = Sjölander K | title = Phylogenomic inference of protein molecular function: advances and challenges | journal = Bioinformatics | volume = 20 | issue = 2 | pages = 170–79 | date = January 2004 | pmid = 14734307 | doi = 10.1093/bioinformatics/bth021 | citeseerx = 10.1.1.412.943 }}</ref> Data sets representing entire genomes' worth of DNA sequences, such as those produced by the ], are difficult to use without the annotations that identify the locations of genes and regulatory elements on each chromosome. Regions of DNA sequence that have the characteristic patterns associated with protein- or RNA-coding genes can be identified by ] algorithms, which allow researchers to predict the presence of particular ]s and their possible functions in an organism even before they have been isolated experimentally.<ref name="Mount">{{cite book | vauthors = Mount DM |title=Bioinformatics: Sequence and Genome Analysis |edition= 2nd |publisher= Cold Spring Harbor Laboratory Press |year= 2004 |isbn= 0-87969-712-1|oclc= 55106399|location=Cold Spring Harbor, NY}}</ref> Entire genomes may also be compared, which can shed light on the evolutionary history of particular organism and permit the examination of complex evolutionary events. | |||
| === DNA nanotechnology === | |||
| ], co-creator of the single X-ray diffraction image]] | |||
| {{further|DNA nanotechnology}} | |||
| ], co-creator of the single X-ray diffraction image]] | |||
| ] at right. ] is the field that seeks to design nanoscale structures using the ] properties of DNA molecules.<ref>{{cite journal | vauthors = Strong M | title = Protein nanomachines | journal = PLOS Biology | volume = 2 | issue = 3 | pages = E73 | date = March 2004 | pmid = 15024422 | pmc = 368168 | doi = 10.1371/journal.pbio.0020073 | s2cid = 13222080 | doi-access = free }}</ref>]] | |||
| ], co-originator of the double-helix model]] | |||
| DNA nanotechnology uses the unique ] properties of DNA and other nucleic acids to create self-assembling branched DNA complexes with useful properties.<ref>{{cite journal | vauthors = Rothemund PW | s2cid = 4316391 | title = Folding DNA to create nanoscale shapes and patterns | journal = Nature | volume = 440 | issue = 7082 | pages = 297–302 | date = March 2006 | pmid = 16541064 | doi = 10.1038/nature04586 | bibcode = 2006Natur.440..297R | url = https://authors.library.caltech.edu/22244/3/nature04586-s2.pdf }}</ref> DNA is thus used as a structural material rather than as a carrier of biological information. This has led to the creation of two-dimensional periodic lattices (both tile-based and using the '']'' method) and three-dimensional structures in the shapes of ].<ref>{{cite journal | vauthors = Andersen ES, Dong M, Nielsen MM, Jahn K, Subramani R, Mamdouh W, Golas MM, Sander B, Stark H, Oliveira CL, Pedersen JS, Birkedal V, Besenbacher F, Gothelf KV, Kjems J | s2cid = 4430815 | title = Self-assembly of a nanoscale DNA box with a controllable lid | journal = Nature | volume = 459 | issue = 7243 | pages = 73–76 | date = May 2009 | pmid = 19424153 | doi = 10.1038/nature07971 | bibcode = 2009Natur.459...73A | hdl = 11858/00-001M-0000-0010-9362-B | hdl-access = free }}</ref> ] and ] have also been demonstrated,<ref>{{cite journal | vauthors = Ishitsuka Y, Ha T | title = DNA nanotechnology: a nanomachine goes live | journal = Nature Nanotechnology | volume = 4 | issue = 5 | pages = 281–82 | date = May 2009 | pmid = 19421208 | doi = 10.1038/nnano.2009.101 | bibcode = 2009NatNa...4..281I }}</ref> and these DNA structures have been used to template the arrangement of other molecules such as ] and ] proteins.<ref>{{cite journal | vauthors = Aldaye FA, Palmer AL, Sleiman HF | title = Assembling materials with DNA as the guide | journal = Science | volume = 321 | issue = 5897 | pages = 1795–99 | date = September 2008 | pmid = 18818351 | doi = 10.1126/science.1154533 | bibcode = 2008Sci...321.1795A | s2cid = 2755388 }}</ref> DNA and other nucleic acids are the basis of ], synthetic oligonucleotide ligands for specific target molecules used in a range of biotechnology and biomedical applications.<ref>{{cite journal | vauthors = Dunn MR, Jimenez RM, Chaput JC |title=Analysis of aptamer discovery and technology |journal=Nature Reviews Chemistry |date=2017 |volume=1 |issue=10 |doi=10.1038/s41570-017-0076 |url=https://www.nature.com/articles/s41570-017-0076 |access-date=30 June 2022}}</ref> | |||
| ==History |
=== History and anthropology === | ||
| {{further|Phylogenetics|Genetic genealogy}} | |||
| {{Further|]}} | |||
| Because DNA collects mutations over time, which are then inherited, it contains historical information, and, by comparing DNA sequences, geneticists can infer the evolutionary history of organisms, their ].<ref>{{cite journal | vauthors = Wray GA | title = Dating branches on the tree of life using DNA | journal = Genome Biology | volume = 3 | issue = 1 | pages = REVIEWS0001 | year = 2002 | pmid = 11806830 | pmc = 150454 | doi = 10.1186/gb-2001-3-1-reviews0001 | doi-access = free }}</ref> This field of phylogenetics is a powerful tool in ]. If DNA sequences within a species are compared, ] can learn the history of particular populations. This can be used in studies ranging from ] to ]. | |||
| DNA was first isolated by the ] physician ] who, in 1869, discovered a microscopic substance in the ] of discarded surgical bandages. As it resided in the nuclei of cells, he called it "nuclein".<ref>{{cite journal |author=Dahm R |title=Discovering DNA: Friedrich Miescher and the early years of nucleic acid research |journal=Hum. Genet. |volume=122 |issue=6 |pages=565–81 |year=2008 |month=January |pmid=17901982 |doi=10.1007/s00439-007-0433-0}}</ref> In 1919, ] identified the base, sugar and phosphate nucleotide unit.<ref>{{cite journal |author=Levene P, |title=The structure of yeast nucleic acid | url=http://www.jbc.org/cgi/reprint/40/2/415 | journal=J Biol Chem |volume=40 |issue=2 | pages=415–24 |date=1 December 1919}}</ref> Levene suggested that DNA consisted of a string of nucleotide units linked together through the phosphate groups. However, Levene thought the chain was short and the bases repeated in a fixed order. In 1937 ] produced the first ] patterns that showed that DNA had a regular structure.<ref>{{cite journal | author =Astbury W, |title=Nucleic acid | journal=Symp. SOC. Exp. Bbl |volume=1 |issue=66 |year=1947}}</ref> | |||
| === Information storage === | |||
| In 1928, ] discovered that ] of the "smooth" form of the ''Pneumococcus'' could be transferred to the "rough" form of the same bacteria by mixing killed "smooth" bacteria with the live "rough" form.<ref>{{cite journal |author=Lorenz MG, Wackernagel W |title=Bacterial gene transfer by natural genetic transformation in the environment |journal=Microbiol. Rev. |volume=58 |issue=3 |pages=563–602 |date=1 September 1994|pmid=7968924 |pmc=372978 |url=http://mmbr.asm.org/cgi/pmidlookup?view=long&pmid=7968924 }}</ref> This system provided the first clear suggestion that DNA carried genetic information—the ]—when ], along with coworkers ] and ], identified DNA as the ] in 1943.<ref>{{cite journal |author=Avery O, MacLeod C, McCarty M |title=Studies on the chemical nature of the substance inducing transformation of pneumococcal types. Inductions of transformation by a desoxyribonucleic acid fraction isolated from pneumococcus type III | url=http://www.jem.org/cgi/reprint/149/2/297 | journal=J Exp Med |volume=79 |issue=2 | pages=137–158 |year=1944 |doi=10.1084/jem.79.2.137 |pmid=19871359 |pmc=2135445}}</ref> DNA's role in ] was confirmed in 1952, when ] and ] in the ] showed that DNA is the ] of the ].<ref>{{cite journal |author=Hershey A, Chase M |title=Independent functions of viral protein and nucleic acid in growth of bacteriophage | url=http://www.jgp.org/cgi/reprint/36/1/39.pdf | journal=J Gen Physiol |volume=36 |issue=1 | pages=39–56 |year=1952 |pmid=12981234 |doi=10.1085/jgp.36.1.39|format=PDF |pmc=2147348}}</ref> | |||
| {{Main|DNA digital data storage}} | |||
| DNA as a ] for information has enormous potential since it has much higher ] compared to electronic devices. However, high costs, slow read and write times (]), and insufficient ] has prevented its practical use.<ref name="pmid29744271">{{cite journal | vauthors = Panda D, Molla KA, Baig MJ, Swain A, Behera D, Dash M | title = DNA as a digital information storage device: hope or hype? | journal = 3 Biotech | volume = 8 | issue = 5 | pages = 239 | date = May 2018 | pmid = 29744271 | doi = 10.1007/s13205-018-1246-7 | pmc=5935598}}</ref><ref name="pmid30073589">{{cite journal | vauthors = Akram F, Haq IU, Ali H, Laghari AT | s2cid = 51905843 | title = Trends to store digital data in DNA: an overview | journal = Molecular Biology Reports | volume = 45 | issue = 5 | pages = 1479–1490 | date = October 2018 | pmid = 30073589 | doi = 10.1007/s11033-018-4280-y }}</ref> | |||
| In 1953 James D. Watson and Francis Crick suggested what is now accepted as the first correct double-helix model of ] in the journal ].<ref name=FWPUB/> Their double-helix, molecular model of DNA was then based on a single ] image (labeled as "]")<ref>The B-DNA X-ray pattern was obtained by ] and ] in May 1952 at high hydration levels of DNA and it has been labeled as "Photo 51"</ref> taken by Rosalind Franklin and Raymond Gosling in May 1952, as well as the information that the DNA bases were paired—also obtained through private communications from Erwin Chargaff in the previous years. ] played a very important role in establishing double-helix configurations for B-DNA as well as A-DNA. | |||
| == History == | |||
| Experimental evidence supporting the Watson and Crick model were published in a series of five articles in the same issue of ''Nature''.<ref name=NatureDNA50>Nature Archives </ref> Of these, Franklin and Gosling's paper was the first publication of their own X-ray diffraction data and original analysis method that partially supported the ] model<ref name=NatFranGos/><ref></ref>; this issue also contained an article on DNA structure by ] and two of his colleagues, whose analysis and ''in vivo'' B-DNA X-ray patterns also supported the presence ''in vivo'' of the double-helical DNA configurations as proposed by Crick and Watson for their double-helix molecular model of DNA in the previous two pages of ''Nature''.<ref name=NatWilk>{{cite journal| title=Molecular Structure of Deoxypentose Nucleic Acids | author= Wilkins M.H.F., A.R. Stokes A.R. & Wilson, H.R. | journal=Nature | volume= 171 | pages= 738–740 | year=1953 | url=http://www.nature.com/nature/dna50/wilkins.pdf| pmid=13054693 | doi=10.1038/171738a0|format=PDF| issue=4356}}</ref> In 1962, after Franklin's death, Watson, Crick, and Wilkins jointly received the ] in ].<ref> Nobelprize .org Accessed 22 December 06</ref> Unfortunately, Nobel rules of the time allowed only living recipients, but a vigorous debate continues on who should receive credit for the discovery.<ref>{{cite journal | title=The double helix and the 'wronged heroine' | author= Brenda Maddox| journal= Nature | volume= 421 | pages= 407–408 | date=23 January 2003 | url=http://www.biomath.nyu.edu/index/course/hw_articles/nature4.pdf | pmid=12540909 | doi = 10.1038/nature01399 |format=PDF | issue=6921}}</ref> | |||
| {{anchor|History of DNA research}} | |||
| {{further|History of molecular biology}} | |||
| ] (left) shakes hands with ] and ], co-originators of the double-helix model based on the X-ray diffraction data and insights of ] and ].]] | |||
| DNA was first isolated by the Swiss physician ] who, in 1869, discovered a microscopic substance in the ] of discarded surgical bandages. As it resided in the nuclei of cells, he called it "nuclein".<ref>{{cite journal | vauthors = Miescher F | year = 1871 | url = https://books.google.com/books?id=YJRTAAAAcAAJ&pg=PA441 | title = Ueber die chemische Zusammensetzung der Eiterzellen | language = de | trans-title = On the chemical composition of pus cells | journal = Medicinisch-chemische Untersuchungen | volume = 4 | pages = 441–60 | quote = ''Ich habe mich daher später mit meinen Versuchen an die ganzen Kerne gehalten, die Trennung der Körper, die ich einstweilen ohne weiteres Präjudiz als lösliches und unlösliches Nuclein bezeichnen will, einem günstigeren Material überlassend.'' (Therefore, in my experiments I subsequently limited myself to the whole nucleus, leaving to a more favorable material the separation of the substances, that for the present, without further prejudice, I will designate as soluble and insoluble nuclear material ("Nuclein"))}}</ref><ref>{{cite journal | vauthors = Dahm R | s2cid = 915930 | title = Discovering DNA: Friedrich Miescher and the early years of nucleic acid research | journal = Human Genetics | volume = 122 | issue = 6 | pages = 565–81 | date = January 2008 | pmid = 17901982 | doi = 10.1007/s00439-007-0433-0 }}</ref> In 1878, ] isolated the non-protein component of "nuclein", nucleic acid, and later isolated its five primary ]s.<ref>See: | |||
| In an influential presentation in 1957, Crick laid out the ], which foretold the relationship between DNA, RNA, and proteins, and articulated the "adaptor hypothesis".<ref>Crick, F.H.C. genome.wellcome.ac.uk (Lecture, 1955). Accessed 22 December 2006</ref> Final confirmation of the replication mechanism that was implied by the double-helical structure followed in 1958 through the ].<ref>{{cite journal |author=Meselson M, Stahl F |title=The replication of DNA in ''Escherichia coli'' | journal=Proc Natl Acad Sci USA |volume=44 |issue=7 | pages=671–82 |year=1958 |pmid=16590258 |doi=10.1073/pnas.44.7.671 |pmc=528642}}</ref> Further work by Crick and coworkers showed that the genetic code was based on non-overlapping triplets of bases, called codons, allowing ], ] and ] to decipher the genetic code.<ref> Nobelprize.org Accessed 22 December 06</ref> These findings represent the birth of ]. | |||
| * {{cite journal | vauthors = Kossel A | year = 1879 | url = https://books.google.com/books?id=4H5NAAAAYAAJ&pg=PA284 | title = Ueber Nucleïn der Hefe | language = de| trans-title = On nuclein in yeast | journal = Zeitschrift für physiologische Chemie | volume = 3 | pages = 284–91 }} | |||
| * {{cite journal | vauthors = Kossel A | year = 1880 | url = https://books.google.com/books?id=u4s1AQAAMAAJ&pg=PA290 | title = Ueber Nucleïn der Hefe II | language = de | trans-title = On nuclein in yeast, Part 2 | journal = Zeitschrift für physiologische Chemie | volume = 4 | pages = 290–95 }} | |||
| * {{cite journal | vauthors = Kossel A | year = 1881 | url = https://books.google.com/books?id=xYs1AQAAMAAJ&pg=PA267 | title = Ueber die Verbreitung des Hypoxanthins im Thier- und Pflanzenreich | language = de | trans-title = On the distribution of hypoxanthins in the animal and plant kingdoms | journal = Zeitschrift für physiologische Chemie | volume = 5 | pages = 267–71 }} | |||
| * {{cite book | vauthors = Kossel A | title = Untersuchungen über die Nucleine und ihre Spaltungsprodukte | language = de | trans-title = Investigations into nuclein and its cleavage products | location = Strassburg, Germany | publisher = K.J. Trübner | year = 1881 | pages = 19 }} | |||
| * {{cite journal | vauthors = Kossel A | year = 1882 | url = https://books.google.com/books?id=z4s1AQAAMAAJ&pg=PA422 | title = Ueber Xanthin und Hypoxanthin |trans-title= On xanthin and hypoxanthin | journal = Zeitschrift für physiologische Chemie | volume = 6 | pages = 422–31 }} | |||
| * Albrect Kossel (1883) {{webarchive|url=https://web.archive.org/web/20171117235430/https://books.google.com/books?id=2os1AQAAMAAJ&pg=PA7 |date=17 November 2017 }} (On the chemistry of the cell nucleus), ''Zeitschrift für physiologische Chemie'', '''7''': 7–22. | |||
| * {{cite journal | vauthors = Kossel A | year = 1886 | title = Weitere Beiträge zur Chemie des Zellkerns | language = de | trans-title = Further contributions to the chemistry of the cell nucleus | journal = Zeitschrift für Physiologische Chemie | volume = 10 | pages = 248–64 | url = http://vlp.mpiwg-berlin.mpg.de/library/data/lit16615/index_html?pn=1&ws=1.5 | quote = On p. 264, Kossel remarked presciently: Der Erforschung der quantitativen Verhältnisse der vier stickstoffreichen Basen, der Abhängigkeit ihrer Menge von den physiologischen Zuständen der Zelle, verspricht wichtige Aufschlüsse über die elementaren physiologisch-chemischen Vorgänge. (The study of the quantitative relations of the four nitrogenous bases— of the dependence of their quantity on the physiological states of the cell—promises important insights into the fundamental physiological-chemical processes.) }}</ref><ref name="Yale_Jones_1953">{{cite journal | vauthors = Jones ME | title = Albrecht Kossel, a biographical sketch | journal = The Yale Journal of Biology and Medicine | volume = 26 | issue = 1 | pages = 80–97 | date = September 1953 | pmid = 13103145 | pmc = 2599350 }}</ref> | |||
| In 1909, ] identified the base, sugar, and phosphate nucleotide unit of RNA (then named "yeast nucleic acid").<ref>{{cite journal | vauthors = Levene PA, Jacobs WA | year = 1909 | title = Über Inosinsäure | language = de | journal = Berichte der Deutschen Chemischen Gesellschaft | volume = 42 | pages = 1198–203 |url=https://babel.hathitrust.org/cgi/pt?id=iau.31858002459620&view=1up&seq=1054 | doi=10.1002/cber.190904201196}}</ref><ref>{{cite journal | vauthors = Levene PA, Jacobs WA | year = 1909 | title = Über die Hefe-Nucleinsäure | language = de | journal = Berichte der Deutschen Chemischen Gesellschaft | volume = 42 | issue = 2 | pages = 2474–78 | doi=10.1002/cber.190904202148| url = https://zenodo.org/record/2175598 }}</ref><ref>{{cite journal | vauthors = Levene P |title=The structure of yeast nucleic acid| journal=J Biol Chem |volume=40 |issue=2 |pages=415–24 |year=1919|doi=10.1016/S0021-9258(18)87254-4|doi-access=free }}</ref> In 1929, Levene identified deoxyribose sugar in "thymus nucleic acid" (DNA).<ref>{{cite journal | vauthors = Cohen JS, Portugal FH | year = 1974 | title = The search for the chemical structure of DNA | journal = Connecticut Medicine | volume = 38 | issue = 10 | pages = 551–52, 554–57 | pmid = 4609088 | url = https://profiles.nlm.nih.gov/ps/access/CCAAHW.pdf }}</ref> Levene suggested that DNA consisted of a string of four nucleotide units linked together through the phosphate groups ("tetranucleotide hypothesis"). Levene thought the chain was short and the bases repeated in a fixed order. In 1927, ] proposed that inherited traits would be inherited via a "giant hereditary molecule" made up of "two mirror strands that would replicate in a semi-conservative fashion using each strand as a template".<ref>Koltsov proposed that a cell's genetic information was encoded in a long chain of amino acids. See: | |||
| ==See also== | |||
| * {{cite speech | vauthors = Koltsov HK | title = Физико-химические основы морфологии | trans-title = The physical-chemical basis of morphology | language = ru | event = 3rd All-Union Meeting of Zoologist, Anatomists, and Histologists | location = Leningrad, U.S.S.R. | date = 12 December 1927 }} | |||
| {{Portal|Molecular and Cellular Biology}} | |||
| * Reprinted in: {{cite journal | vauthors = Koltsov HK | title = Физико-химические основы морфологии | trans-title = The physical-chemical basis of morphology | language = ru| journal = Успехи экспериментальной биологии (Advances in Experimental Biology) series B | volume = 7 | issue = 1 | pages = ? | date = 1928 }} | |||
| <div style="-moz-column-count:2; column-count:2;"> | |||
| * Reprinted in German as: {{cite journal | vauthors = Koltzoff NK | date = 1928 | title = Physikalisch-chemische Grundlagen der Morphologie | trans-title = The physical-chemical basis of morphology | language = de | journal = Biologisches Zentralblatt | volume = 48 | issue = 6 | pages = 345–69 }} | |||
| * ] | |||
| * In 1934, Koltsov contended that the proteins that contain a cell's genetic information replicate. See: {{cite journal | vauthors = Koltzoff N | title = The structure of the chromosomes in the salivary glands of Drosophila | journal = Science | volume = 80 | issue = 2075 | pages = 312–13 | date = October 1934 | pmid = 17769043 | doi = 10.1126/science.80.2075.312 | quote = From page 313: "I think that the size of the chromosomes in the salivary glands is determined through the multiplication of ''genonemes''. By this term I designate the axial thread of the chromosome, in which the geneticists locate the linear combination of genes; … In the normal chromosome there is usually only one genoneme; before cell-division this genoneme has become divided into two strands."| bibcode = 1934Sci....80..312K }}</ref><ref name="Soyfer">{{cite journal | vauthors = Soyfer VN | s2cid = 46277758 | title = The consequences of political dictatorship for Russian science | journal = Nature Reviews Genetics | volume = 2 | issue = 9 | pages = 723–29 | date = September 2001 | pmid = 11533721 | doi = 10.1038/35088598 }}</ref> In 1928, ] in his ] discovered that ] of the "smooth" form of ''Pneumococcus'' could be transferred to the "rough" form of the same bacteria by mixing killed "smooth" bacteria with the live "rough" form.<ref>{{cite journal | vauthors = Griffith F | title = The Significance of Pneumococcal Types | journal = The Journal of Hygiene | volume = 27 | issue = 2 | pages = 113–59 | date = January 1928 | pmid = 20474956 | pmc = 2167760 | doi = 10.1017/S0022172400031879 }}</ref><ref>{{cite journal | vauthors = Lorenz MG, Wackernagel W | title = Bacterial gene transfer by natural genetic transformation in the environment | journal = Microbiological Reviews | volume = 58 | issue = 3 | pages = 563–602 | date = September 1994 | pmid = 7968924 | pmc = 372978 | doi = 10.1128/MMBR.58.3.563-602.1994 }}</ref> This system provided the first clear suggestion that DNA carries genetic information. | |||
| * ] | |||
| * ] | |||
| * ] | |||
| * ] | |||
| * ] | |||
| * ] | |||
| * ] | |||
| * ] | |||
| * ] | |||
| * ] | |||
| * ] | |||
| * ] | |||
| * ] | |||
| * ] | |||
| * ] | |||
| * ] | |||
| * {{Proteopedia|DNA}} | |||
| * ] | |||
| * ] | |||
| </div> | |||
| In 1933, while studying virgin ] eggs, ] suggested that DNA is found in the ] and that ] is present exclusively in the ]. At the time, "yeast nucleic acid" (RNA) was thought to occur only in plants, while "thymus nucleic acid" (DNA) only in animals. The latter was thought to be a tetramer, with the function of buffering cellular pH.<ref>{{cite journal | vauthors = Brachet J | year = 1933 | title = Recherches sur la synthese de l'acide thymonucleique pendant le developpement de l'oeuf d'Oursin | language = it | journal = Archives de Biologie | volume = 44 | pages = 519–76 }}</ref><ref>{{cite book | vauthors = Burian R | year = 1994 | chapter = Jean Brachet's Cytochemical Embryology: Connections with the Renovation of Biology in France? | veditors = Debru C, Gayon J, Picard JF | title = Les sciences biologiques et médicales en France 1920–1950 | volume = 2 | series = Cahiers pour I'histoire de la recherche | location = Paris | publisher = CNRS Editions | pages = 207–20 | chapter-url = http://www.histcnrs.fr/ColloqDijon/Burian-Brachet.pdf }}</ref> | |||
| ==References== | |||
| {{Reflist|colwidth=30em}} | |||
| In 1937, ] produced the first X-ray diffraction patterns that showed that DNA had a regular structure.<ref>See: | |||
| ==Further reading== | |||
| * {{cite journal | vauthors = Astbury WT, Bell FO | year = 1938 | title = Some recent developments in the X-ray study of proteins and related structures | journal = Cold Spring Harbor Symposia on Quantitative Biology | volume = 6 | pages = 109–21 | url = http://www.leeds.ac.uk/heritage/Astbury/bibliography/CSHSQB_Astbury_and_Bell_1938.pdf | archive-url = https://web.archive.org/web/20140714204539/http://www.leeds.ac.uk/heritage/Astbury/bibliography/CSHSQB_Astbury_and_Bell_1938.pdf | archive-date = 14 July 2014 | doi=10.1101/sqb.1938.006.01.013}} | |||
| *{{cite book |title=Understanding DNA: the molecule & how it works |author=Calladine, Chris R.; Drew, Horace R.; Luisi, Ben F. and Travers, Andrew A. |year=2003 |publisher=Elsevier Academic Press |location=Amsterdam |isbn=0-12-155089-3 }} | |||
| * {{cite journal | vauthors = Astbury WT | title = X-ray studies of nucleic acids | journal = Symposia of the Society for Experimental Biology | issue = 1 | pages = 66–76 | year = 1947 | pmid = 20257017 | url = http://scarc.library.oregonstate.edu/coll/pauling/dna/papers/astbury-xray.html | archive-url = https://web.archive.org/web/20140705132403/http://scarc.library.oregonstate.edu/coll/pauling/dna/papers/astbury-xray.html | archive-date=5 July 2014 }}</ref> | |||
| *{{cite book |author=Dennis, Carina; Julie Clayton |title=50 years of DNA |publisher=Palgrave Macmillan |location=Basingstoke |year=2003 |isbn=1-4039-1479-6 }} | |||
| *{{cite book |author=Judson, Horace Freeland |title=The eighth day of creation: makers of the revolution in biology |publisher=CSHL Press |location=Plainview, N.Y |year=1996 |isbn=0-87969-478-5 }} | |||
| *{{cite book |author=Olby, Robert C. |authorlink=Robert Olby |title=The path to the double helix: the discovery of DNA |publisher=Dover Publications |location=New York |year=1994 |isbn=0-486-68117-3 }}, first published in October 1974 by MacMillan, with foreword by Francis Crick;the definitive DNA textbook,revised in 1994 with a 9 page postscript. | |||
| *{{cite book |author=Olby, Robert C. |title=Francis Crick: A Biography |publisher=Cold Spring Harbor Laboratory Press |location=Plainview, N.Y |year=2009 |isbn=0-87969-798-9 }} | |||
| *{{cite book |author=Ridley, Matt |authorlink=Matt Ridley |title=Francis Crick: discoverer of the genetic code |publisher=Eminent Lives, Atlas Books |location=[Ashland, OH |year=2006 |isbn=0-06-082333-X }} | |||
| *{{cite book |author=Berry, Andrew; Watson, James D. |title=DNA: the secret of life |publisher=Alfred A. Knopf |location=New York |year=2003 |isbn=0-375-41546-7 }} | |||
| *{{cite book |author=Stent, Gunther Siegmund; Watson, James D. |title=] |publisher=Norton |location=New York |year=1980 |isbn=0-393-95075-1 }} | |||
| *{{cite book |author=Wilkins, Maurice |title=The third man of the double helix the autobiography of Maurice Wilkins |publisher=University Press |location=Cambridge, Eng |year=2003 |isbn=0-19-860665-6 }} | |||
| In 1943, ], along with co-workers ] and ], identified DNA as the ], supporting Griffith's suggestion (]).<ref>{{cite journal | vauthors = Avery OT, Macleod CM, McCarty M | title = Studies on the Chemical Nature of the Substance Inducing Transformation of Pneumococcal Types: Induction of Transformation by a Desoxyribonucleic Acid Fraction Isolated from Pneumococcus Type III | journal = The Journal of Experimental Medicine | volume = 79 | issue = 2 | pages = 137–158 | date = February 1944 | pmid = 19871359 | pmc = 2135445 | doi = 10.1084/jem.79.2.137 }}</ref> ] developed and published observations now known as ], stating that in DNA from any species of any organism, the amount of ] should be equal to ] and the amount of ] should be equal to ].<ref name="chargaff_1950">{{cite journal | vauthors = Chargaff E | title = Chemical specificity of nucleic acids and mechanism of their enzymatic degradation | journal = Experientia | volume = 6 | issue = 6 | pages = 201–209 | date = June 1950 | pmid = 15421335 | doi = 10.1007/BF02173653 | s2cid = 2522535 }}</ref><ref name="kresge_2005">{{cite journal | vauthors = Kresge N, Simoni RD, Hill RL |title=Chargaff's Rules: the Work of Erwin Chargaff |journal=Journal of Biological Chemistry |date=June 2005 |volume=280 |issue=24 |pages=172–174 |doi=10.1016/S0021-9258(20)61522-8|doi-access=free }}</ref> | |||
| ==External links== | |||
| {{Spoken Misplaced Pages|dna.ogg|2007-02-12}} | |||
| ] outside ] ] in Cambridge, England commemorating Crick and Watson]] | |||
| {{Commons category|DNA}} | |||
| * {{dmoz|Science/Biology/Biochemistry_and_Molecular_Biology/Biomolecules/Nucleic_Acids/DNA/|DNA}} | |||
| Late in 1951, ] started working with ] at the ] within the ]. DNA's role in ] was confirmed in 1952 when ] and ] in the ] showed that DNA is the ] of the ].<ref>{{cite journal | vauthors = Hershey AD, Chase M | title = Independent functions of viral protein and nucleic acid in growth of bacteriophage | journal = The Journal of General Physiology | volume = 36 | issue = 1 | pages = 39–56 | date = May 1952 | pmid = 12981234 | pmc = 2147348 | doi = 10.1085/jgp.36.1.39 }}</ref> | |||
| * | |||
| ] | |||
| * | |||
| In May 1952, ], a graduate student working under the supervision of ], took an ] image, labeled as "]",<ref>{{Cite web|title=Pictures and Illustrations: Crystallographic photo of Sodium Thymonucleate, Type B. "Photo 51." May 1952|url=http://scarc.library.oregonstate.edu/coll/pauling/dna/pictures/sci9.001.5.html|access-date=2023-05-18|website=scarc.library.oregonstate.edu}}</ref> at high hydration levels of DNA. This photo was given to Watson and Crick by ] and was critical to their obtaining the correct structure of DNA. Franklin told Crick and Watson that the backbones had to be on the outside. Before then, Linus Pauling, and Watson and Crick, had erroneous models with the chains inside and the bases pointing outwards. Franklin's identification of the ] for DNA crystals revealed to Crick that the two DNA strands were ].<ref name="ReferenceA">{{cite book| vauthors = Schwartz J |title=In pursuit of the gene: from Darwin to DNA |url= https://archive.org/details/inpursuitofgenef00schw |url-access=registration |year=2008|publisher=Harvard University Press|location=Cambridge, Mass.|isbn=978-0-674-02670-4 }}</ref> In February 1953, ] and ] proposed a model for nucleic acids containing three intertwined chains, with the phosphates near the axis, and the bases on the outside.<ref name="pmid16578429">{{cite journal | vauthors = Pauling L, Corey RB | title = A Proposed Structure For The Nucleic Acids | journal = Proceedings of the National Academy of Sciences of the United States of America | volume = 39 | issue = 2 | pages = 84–97 | date = February 1953 | pmid = 16578429 | pmc = 1063734 | doi = 10.1073/pnas.39.2.84| url = http://scarc.library.oregonstate.edu/coll/pauling/dna/papers/1953p.9-084.html | bibcode = 1953PNAS...39...84P | doi-access = free }}</ref> Watson and Crick completed their model, which is now accepted as the first correct model of the double helix of ]. On 28 February 1953 Crick interrupted patrons' lunchtime at ] ] in Cambridge, England to announce that he and Watson had "discovered the secret of life".<ref>{{cite book | vauthors = Regis E | date = 2009 | title = What Is Life?: investigating the nature of life in the age of synthetic biology | location = Oxford | publisher = ] | isbn = 978-0-19-538341-6 | page = 52 }}</ref> | |||
| * Another DNA Learning Center site on DNA, genes, and heredity from Mendel to the human genome project. | |||
| * | |||
| ] | |||
| * From the official Nobel Prize web site | |||
| The 25 April 1953 issue of the journal '']'' published a series of five articles giving the Watson and Crick double-helix structure DNA and evidence supporting it.<ref name=NatureDNA50>{{cite web | work = Nature Archives | url = http://www.nature.com/nature/dna50/archive.html | title = Double Helix of DNA: 50 Years | archive-url = https://web.archive.org/web/20150405140401/http://www.nature.com/nature/dna50/archive.html | archive-date=5 April 2015 | url-status=dead }}</ref> The structure was reported in a letter titled "''MOLECULAR STRUCTURE OF NUCLEIC ACIDS A Structure for Deoxyribose Nucleic Acid''{{-"}}, in which they said, "It has not escaped our notice that the specific pairing we have postulated immediately suggests a possible copying mechanism for the genetic material."<ref name="Watson-1953" /> This letter was followed by a letter from Franklin and Gosling, which was the first publication of their own X-ray diffraction data and of their original analysis method.<ref name=NatFranGos /><ref>{{cite web | url = http://osulibrary.oregonstate.edu/specialcollections/coll/pauling/dna/pictures/franklin-typeBphoto.html | title = Original X-ray diffraction image | publisher = Oregon State Library | access-date = 6 February 2011 | url-status=live | archive-url = https://web.archive.org/web/20090130111849/http://osulibrary.oregonstate.edu/specialcollections/coll/pauling/dna/pictures/franklin-typeBphoto.html | archive-date=30 January 2009 }}</ref> Then followed a letter by Wilkins and two of his colleagues, which contained an analysis of ''in vivo'' B-DNA X-ray patterns, and which supported the presence ''in vivo'' of the Watson and Crick structure.<ref name="NatWilk" /> | |||
| In April 2023, scientists, based on new evidence, concluded that Rosalind Franklin was a contributor and "equal player" in the discovery process of DNA, rather than otherwise, as may have been presented subsequently after the time of the discovery.<ref name="AP-20230425">{{cite news | vauthors = Burakoff M |title=Rosalind Franklin's role in DNA discovery gets a new twist |url=https://apnews.com/article/dna-double-helix-rosalind-franklin-watson-crick-69ec8164c720e0b23374da69a1d3708d |date=25 April 2023 |work=] |accessdate=25 April 2023 }}</ref><ref name="NYT-20230425">{{cite news | vauthors = Anthes E |title=Untangling Rosalind Franklin's Role in DNA Discovery, 70 Years On – Historians have long debated the role that Dr. Franklin played in identifying the double helix. A new opinion essay argues that she was an "equal contributor." |url=https://www.nytimes.com/2023/04/25/science/rosalind-franklin-dna.html |date=25 April 2023 |work=] |url-status=live |archiveurl=https://archive.today/20230425182515/https://www.nytimes.com/2023/04/25/science/rosalind-franklin-dna.html |archivedate=25 April 2023 |accessdate=26 April 2023 }}</ref><ref name="NAT-202304254">{{cite journal | vauthors = Cobb M, Comfort N |title=What Rosalind Franklin truly contributed to the discovery of DNA's structure – Franklin was no victim in how the DNA double helix was solved. An overlooked letter and an unpublished news article, both written in 1953, reveal that she was an equal player. |date=25 April 2023 |journal=] |volume=616 |issue=7958 |pages=657–660 |doi=10.1038/d41586-023-01313-5 |pmid=37100935 |s2cid=258314143 |doi-access=free |bibcode=2023Natur.616..657C }}</ref> | |||
| In 1962, after Franklin's death, Watson, Crick, and Wilkins jointly received the ].<ref>{{cite web | url = http://nobelprize.org/nobel_prizes/medicine/laureates/1962/ | title = The Nobel Prize in Physiology or Medicine 1962 | work = Nobelprize.org }}</ref> Nobel Prizes are awarded only to living recipients. A debate continues about who should receive credit for the discovery.<ref>{{cite journal | vauthors = Maddox B | s2cid = 4428347 | title = The double helix and the 'wronged heroine' | journal = Nature | volume = 421 | issue = 6921 | pages = 407–08 | date = January 2003 | pmid = 12540909 | doi = 10.1038/nature01399 | url = http://www.biomath.nyu.edu/index/course/hw_articles/nature4.pdf | bibcode = 2003Natur.421..407M | url-status=live | archive-url = https://web.archive.org/web/20161017011403/http://www.biomath.nyu.edu/index/course/hw_articles/nature4.pdf | archive-date = 17 October 2016 | df = dmy-all | doi-access = free }}</ref> | |||
| In an influential presentation in 1957, Crick laid out the ], which foretold the relationship between DNA, RNA, and proteins, and articulated the "adaptor hypothesis".<ref>{{cite speech | vauthors = Crick FH |title=A Note for the RNA Tie Club | date= 1955 | location = Cambridge, England |url= http://genome.wellcome.ac.uk/assets/wtx030893.pdf | archive-url = https://web.archive.org/web/20081001223217/http://genome.wellcome.ac.uk/assets/wtx030893.pdf | url-status=dead | archive-date = 1 October 2008 }}</ref> Final confirmation of the replication mechanism that was implied by the double-helical structure followed in 1958 through the ].<ref>{{cite journal | vauthors = Meselson M, Stahl FW | title = The Replication of DNA in Escherichia Coli | journal = Proceedings of the National Academy of Sciences of the United States of America | volume = 44 | issue = 7 | pages = 671–82 | date = July 1958 | pmid = 16590258 | pmc = 528642 | doi = 10.1073/pnas.44.7.671 | bibcode = 1958PNAS...44..671M | doi-access = free }}</ref> Further work by Crick and co-workers showed that the genetic code was based on non-overlapping triplets of bases, called ], allowing ], ], and ] to decipher the genetic code.<ref>{{cite web | url = http://nobelprize.org/nobel_prizes/medicine/laureates/1968/ | title = The Nobel Prize in Physiology or Medicine 1968 | work = Nobelprize.org }}</ref> These findings represent the birth of ].<ref>{{cite journal | vauthors = Pray L | year = 2008 | title = Discovery of DNA structure and function: Watson and Crick. | journal = Nature Education | volume = 1 | issue = 1 | pages = 100 }}</ref> | |||
| {{clear}} | |||
| In 1986, DNA analysis was first used for criminal investigative purposes when police in the UK requested ] of the University of Leicester to verify or disprove a suspect's rape-murder "confession". In this particular case, the suspect had confessed to two rape-murders, but had later retracted his confession. DNA testing at the university labs soon disproved the veracity of the suspect's original "confession", and the suspect was exonerated from the murder-rape charges.<ref>{{cite journal | pmc=3561883 | year=2003 | vauthors=Panneerchelvam S, Norazmi MN | title=Forensic DNA Profiling and Database | journal=The Malaysian Journal of Medical Sciences | volume=10 | issue=2 | pages=20–26 | pmid=23386793 }}</ref> | |||
| == See also == | |||
| {{Div col}} | |||
| * {{annotated link|Autosome}} | |||
| * {{annotated link|Crystallography}} | |||
| * {{annotated link|DNA Day}} | |||
| * {{annotated link|DNA microarray}} | |||
| * {{annotated link|DNA sequencing}} | |||
| * {{annotated link|Genetic disorder}} | |||
| * {{annotated link|Genetic genealogy}} | |||
| * {{annotated link|Haplotype}} | |||
| * {{annotated link|Meiosis}} | |||
| * {{annotated link|Nucleic acid notation}} | |||
| * {{annotated link|Nucleic acid sequence}} | |||
| * {{annotated link|Ribosomal DNA}} | |||
| * {{annotated link|Southern blot}} | |||
| * {{annotated link|X-ray scattering techniques}} | |||
| * {{annotated link|Xeno nucleic acid}} | |||
| {{Div col end}} | |||
| == References == | |||
| {{Reflist|refs = | |||
| <ref name="Birney_2007">{{cite journal |author-link2=John Stamatoyannopoulos | vauthors = Birney E, Stamatoyannopoulos JA, Dutta A, Guigó R, Gingeras TR, Margulies EH, Weng Z, Snyder M, Dermitzakis ET, Thurman RE, Kuehn MS, Taylor CM, Neph S, Koch CM, Asthana S, Malhotra A, Adzhubei I, Greenbaum JA, Andrews RM, Flicek P, Boyle PJ, Cao H, Carter NP, Clelland GK, Davis S, Day N, Dhami P, Dillon SC, Dorschner MO, Fiegler H, Giresi PG, Goldy J, Hawrylycz M, Haydock A, Humbert R, James KD, Johnson BE, Johnson EM, Frum TT, Rosenzweig ER, Karnani N, Lee K, Lefebvre GC, Navas PA, Neri F, Parker SC, Sabo PJ, Sandstrom R, Shafer A, Vetrie D, Weaver M, Wilcox S, Yu M, Collins FS, Dekker J, Lieb JD, Tullius TD, Crawford GE, Sunyaev S, Noble WS, Dunham I, Denoeud F, Reymond A, Kapranov P, Rozowsky J, Zheng D, Castelo R, Frankish A, Harrow J, Ghosh S, Sandelin A, Hofacker IL, Baertsch R, Keefe D, Dike S, Cheng J, Hirsch HA, Sekinger EA, Lagarde J, Abril JF, Shahab A, Flamm C, Fried C, Hackermüller J, Hertel J, Lindemeyer M, Missal K, Tanzer A, Washietl S, Korbel J, Emanuelsson O, Pedersen JS, Holroyd N, Taylor R, Swarbreck D, Matthews N, Dickson MC, Thomas DJ, Weirauch MT, Gilbert J, Drenkow J, Bell I, Zhao X, Srinivasan KG, Sung WK, Ooi HS, Chiu KP, Foissac S, Alioto T, Brent M, Pachter L, Tress ML, Valencia A, Choo SW, Choo CY, Ucla C, Manzano C, Wyss C, Cheung E, Clark TG, Brown JB, Ganesh M, Patel S, Tammana H, Chrast J, Henrichsen CN, Kai C, Kawai J, Nagalakshmi U, Wu J, Lian Z, Lian J, Newburger P, Zhang X, Bickel P, Mattick JS, Carninci P, Hayashizaki Y, Weissman S, Hubbard T, Myers RM, Rogers J, Stadler PF, Lowe TM, Wei CL, Ruan Y, Struhl K, Gerstein M, Antonarakis SE, Fu Y, Green ED, Karaöz U, Siepel A, Taylor J, Liefer LA, Wetterstrand KA, Good PJ, Feingold EA, Guyer MS, Cooper GM, Asimenos G, Dewey CN, Hou M, Nikolaev S, Montoya-Burgos JI, Löytynoja A, Whelan S, Pardi F, Massingham T, Huang H, Zhang NR, Holmes I, Mullikin JC, Ureta-Vidal A, Paten B, Seringhaus M, Church D, Rosenbloom K, Kent WJ, Stone EA, Batzoglou S, Goldman N, Hardison RC, Haussler D, Miller W, Sidow A, Trinklein ND, Zhang ZD, Barrera L, Stuart R, King DC, Ameur A, Enroth S, Bieda MC, Kim J, Bhinge AA, Jiang N, Liu J, Yao F, Vega VB, Lee CW, Ng P, Shahab A, Yang A, Moqtaderi Z, Zhu Z, Xu X, Squazzo S, Oberley MJ, Inman D, Singer MA, Richmond TA, Munn KJ, Rada-Iglesias A, Wallerman O, Komorowski J, Fowler JC, Couttet P, Bruce AW, Dovey OM, Ellis PD, Langford CF, Nix DA, Euskirchen G, Hartman S, Urban AE, Kraus P, Van Calcar S, Heintzman N, Kim TH, Wang K, Qu C, Hon G, Luna R, Glass CK, Rosenfeld MG, Aldred SF, Cooper SJ, Halees A, Lin JM, Shulha HP, Zhang X, Xu M, Haidar JN, Yu Y, Ruan Y, Iyer VR, Green RD, Wadelius C, Farnham PJ, Ren B, Harte RA, Hinrichs AS, Trumbower H, Clawson H, Hillman-Jackson J, Zweig AS, Smith K, Thakkapallayil A, Barber G, Kuhn RM, Karolchik D, Armengol L, Bird CP, de Bakker PI, Kern AD, Lopez-Bigas N, Martin JD, Stranger BE, Woodroffe A, Davydov E, Dimas A, Eyras E, Hallgrímsdóttir IB, Huppert J, Zody MC, Abecasis GR, Estivill X, Bouffard GG, Guan X, Hansen NF, Idol JR, Maduro VV, Maskeri B, McDowell JC, Park M, Thomas PJ, Young AC, Blakesley RW, Muzny DM, Sodergren E, Wheeler DA, Worley KC, Jiang H, Weinstock GM, Gibbs RA, Graves T, Fulton R, Mardis ER, Wilson RK, Clamp M, Cuff J, Gnerre S, Jaffe DB, Chang JL, Lindblad-Toh K, Lander ES, Koriabine M, Nefedov M, Osoegawa K, Yoshinaga Y, Zhu B, de Jong PJ | display-authors = 6 | title = Identification and analysis of functional elements in 1% of the human genome by the ENCODE pilot project | journal = Nature | volume = 447 | issue = 7146 | pages = 799–816 | date = June 2007 | pmid = 17571346 | pmc = 2212820 | doi = 10.1038/nature05874 | bibcode = 2007Natur.447..799B }}</ref> | |||
| <ref name="Gregory_2006">{{cite journal | vauthors = Gregory SG, Barlow KF, McLay KE, Kaul R, Swarbreck D, Dunham A, Scott CE, Howe KL, Woodfine K, Spencer CC, Jones MC, Gillson C, Searle S, Zhou Y, Kokocinski F, McDonald L, Evans R, Phillips K, Atkinson A, Cooper R, Jones C, Hall RE, Andrews TD, Lloyd C, Ainscough R, Almeida JP, Ambrose KD, Anderson F, Andrew RW, Ashwell RI, Aubin K, Babbage AK, Bagguley CL, Bailey J, Beasley H, Bethel G, Bird CP, Bray-Allen S, Brown JY, Brown AJ, Buckley D, Burton J, Bye J, Carder C, Chapman JC, Clark SY, Clarke G, Clee C, Cobley V, Collier RE, Corby N, Coville GJ, Davies J, Deadman R, Dunn M, Earthrowl M, Ellington AG, Errington H, Frankish A, Frankland J, French L, Garner P, Garnett J, Gay L, Ghori MR, Gibson R, Gilby LM, Gillett W, Glithero RJ, Grafham DV, Griffiths C, Griffiths-Jones S, Grocock R, Hammond S, Harrison ES, Hart E, Haugen E, Heath PD, Holmes S, Holt K, Howden PJ, Hunt AR, Hunt SE, Hunter G, Isherwood J, James R, Johnson C, Johnson D, Joy A, Kay M, Kershaw JK, Kibukawa M, Kimberley AM, King A, Knights AJ, Lad H, Laird G, Lawlor S, Leongamornlert DA, Lloyd DM, Loveland J, Lovell J, Lush MJ, Lyne R, Martin S, Mashreghi-Mohammadi M, Matthews L, Matthews NS, McLaren S, Milne S, Mistry S, Moore MJ, Nickerson T, O'Dell CN, Oliver K, Palmeiri A, Palmer SA, Parker A, Patel D, Pearce AV, Peck AI, Pelan S, Phelps K, Phillimore BJ, Plumb R, Rajan J, Raymond C, Rouse G, Saenphimmachak C, Sehra HK, Sheridan E, Shownkeen R, Sims S, Skuce CD, Smith M, Steward C, Subramanian S, Sycamore N, Tracey A, Tromans A, Van Helmond Z, Wall M, Wallis JM, White S, Whitehead SL, Wilkinson JE, Willey DL, Williams H, Wilming L, Wray PW, Wu Z, Coulson A, Vaudin M, Sulston JE, Durbin R, Hubbard T, Wooster R, Dunham I, Carter NP, McVean G, Ross MT, Harrow J, Olson MV, Beck S, Rogers J, Bentley DR, Banerjee R, Bryant SP, Burford DC, Burrill WD, Clegg SM, Dhami P, Dovey O, Faulkner LM, Gribble SM, Langford CF, Pandian RD, Porter KM, Prigmore E | display-authors = 6 | title = The DNA sequence and biological annotation of human chromosome 1 | journal = Nature | volume = 441 | issue = 7091 | pages = 315–21 | date = May 2006 | pmid = 16710414 | doi = 10.1038/nature04727 | bibcode = 2006Natur.441..315G | doi-access = free }}</ref> | |||
| <ref name="Venter_2001">{{cite journal | vauthors = Venter JC, Adams MD, Myers EW, Li PW, Mural RJ, Sutton GG, Smith HO, Yandell M, Evans CA, Holt RA, Gocayne JD, Amanatides P, Ballew RM, Huson DH, Wortman JR, Zhang Q, Kodira CD, Zheng XH, Chen L, Skupski M, Subramanian G, Thomas PD, Zhang J, Gabor Miklos GL, Nelson C, Broder S, Clark AG, Nadeau J, McKusick VA, Zinder N, Levine AJ, Roberts RJ, Simon M, Slayman C, Hunkapiller M, Bolanos R, Delcher A, Dew I, Fasulo D, Flanigan M, Florea L, Halpern A, Hannenhalli S, Kravitz S, Levy S, Mobarry C, Reinert K, Remington K, Abu-Threideh J, Beasley E, Biddick K, Bonazzi V, Brandon R, Cargill M, Chandramouliswaran I, Charlab R, Chaturvedi K, Deng Z, Di Francesco V, Dunn P, Eilbeck K, Evangelista C, Gabrielian AE, Gan W, Ge W, Gong F, Gu Z, Guan P, Heiman TJ, Higgins ME, Ji RR, Ke Z, Ketchum KA, Lai Z, Lei Y, Li Z, Li J, Liang Y, Lin X, Lu F, Merkulov GV, Milshina N, Moore HM, Naik AK, Narayan VA, Neelam B, Nusskern D, Rusch DB, Salzberg S, Shao W, Shue B, Sun J, Wang Z, Wang A, Wang X, Wang J, Wei M, Wides R, Xiao C, Yan C, Yao A, Ye J, Zhan M, Zhang W, Zhang H, Zhao Q, Zheng L, Zhong F, Zhong W, Zhu S, Zhao S, Gilbert D, Baumhueter S, Spier G, Carter C, Cravchik A, Woodage T, Ali F, An H, Awe A, Baldwin D, Baden H, Barnstead M, Barrow I, Beeson K, Busam D, Carver A, Center A, Cheng ML, Curry L, Danaher S, Davenport L, Desilets R, Dietz S, Dodson K, Doup L, Ferriera S, Garg N, Gluecksmann A, Hart B, Haynes J, Haynes C, Heiner C, Hladun S, Hostin D, Houck J, Howland T, Ibegwam C, Johnson J, Kalush F, Kline L, Koduru S, Love A, Mann F, May D, McCawley S, McIntosh T, McMullen I, Moy M, Moy L, Murphy B, Nelson K, Pfannkoch C, Pratts E, Puri V, Qureshi H, Reardon M, Rodriguez R, Rogers YH, Romblad D, Ruhfel B, Scott R, Sitter C, Smallwood M, Stewart E, Strong R, Suh E, Thomas R, Tint NN, Tse S, Vech C, Wang G, Wetter J, Williams S, Williams M, Windsor S, Winn-Deen E, Wolfe K, Zaveri J, Zaveri K, Abril JF, Guigó R, Campbell MJ, Sjolander KV, Karlak B, Kejariwal A, Mi H, Lazareva B, Hatton T, Narechania A, Diemer K, Muruganujan A, Guo N, Sato S, Bafna V, Istrail S, Lippert R, Schwartz R, Walenz B, Yooseph S, Allen D, Basu A, Baxendale J, Blick L, Caminha M, Carnes-Stine J, Caulk P, Chiang YH, Coyne M, Dahlke C, Mays A, Dombroski M, Donnelly M, Ely D, Esparham S, Fosler C, Gire H, Glanowski S, Glasser K, Glodek A, Gorokhov M, Graham K, Gropman B, Harris M, Heil J, Henderson S, Hoover J, Jennings D, Jordan C, Jordan J, Kasha J, Kagan L, Kraft C, Levitsky A, Lewis M, Liu X, Lopez J, Ma D, Majoros W, McDaniel J, Murphy S, Newman M, Nguyen T, Nguyen N, Nodell M, Pan S, Peck J, Peterson M, Rowe W, Sanders R, Scott J, Simpson M, Smith T, Sprague A, Stockwell T, Turner R, Venter E, Wang M, Wen M, Wu D, Wu M, Xia A, Zandieh A, Zhu X | display-authors = 6 | title = The sequence of the human genome | journal = Science | volume = 291 | issue = 5507 | pages = 1304–51 | date = February 2001 | pmid = 11181995 | doi = 10.1126/science.1058040 | bibcode = 2001Sci...291.1304V | doi-access = free }}</ref> | |||
| }} | |||
| == Further reading == | |||
| {{refbegin}} | |||
| * {{cite book | vauthors = Berry A, Watson J | author-link2 = James Watson | name-list-style = vanc | title = DNA: the secret of life | publisher = Alfred A. Knopf | location = New York | year = 2003 | isbn = 0-375-41546-7 | url = https://archive.org/details/dnasecretoflife00wats }} | |||
| * {{cite book |title=Understanding DNA: the molecule & how it works | vauthors = Calladine CR, Drew HR, Luisi BF, Travers AA | year = 2003 |publisher=Elsevier Academic Press |location=Amsterdam |isbn=0-12-155089-3}} | |||
| * {{cite book | vauthors = Carina D, Clayton J | title = 50 years of DNA | publisher = Palgrave Macmillan | location = Basingstoke | year = 2003 | isbn = 1-4039-1479-6 | url = https://archive.org/details/50yearsofdna00clay }} | |||
| * {{cite book | author-link = Horace Freeland Judson | vauthors = Judson HF | date = 1979 | title = The Eighth Day of Creation: Makers of the Revolution in Biology | isbn = 0-671-22540-5 | edition = 2nd | publisher = Cold Spring Harbor Laboratory Press }} | |||
| * {{cite book | vauthors = Olby RC | author-link = Robert Olby | title = The path to the double helix: the discovery of DNA |publisher=Dover Publications |location=New York |year=1994 |isbn=0-486-68117-3}} First published in October 1974 by MacMillan, with foreword by Francis Crick; the definitive DNA textbook, revised in 1994 with a nine-page postscript. | |||
| * {{cite journal | vauthors = Olby R | title = Quiet debut for the double helix | journal = Nature | volume = 421 | issue = 6921 | pages = 402–05 | date = January 2003 | pmid = 12540907 | doi = 10.1038/nature01397 | author-link = Robert Olby | bibcode = 2003Natur.421..402O | doi-access = free }} | |||
| * {{cite book | vauthors = Olby RC | title=Francis Crick: A Biography |publisher=Cold Spring Harbor Laboratory Press |location=Plainview, N.Y |year=2009 |isbn=978-0-87969-798-3}} | |||
| * {{cite book | vauthors = Micklas D | date = 2003 | title = DNA Science: A First Course | publisher = Cold Spring Harbor Press | isbn = 978-0-87969-636-8 }} | |||
| * {{cite book | vauthors = Ridley M | author-link = Matt Ridley |title=Francis Crick: discoverer of the genetic code |publisher=Eminent Lives, Atlas Books |location=Ashland, OH |year=2006 |isbn=0-06-082333-X}} | |||
| * {{cite book | vauthors = Rosenfeld I | date = 2010 | title = DNA: A Graphic Guide to the Molecule that Shook the World | publisher = Columbia University Press | isbn = 978-0-231-14271-7 }} | |||
| * {{cite book | vauthors = Schultz M, Cannon Z | date = 2009 | title = The Stuff of Life: A Graphic Guide to Genetics and DNA | url = https://archive.org/details/stuffoflifegraph00schu | url-access = registration | publisher = Hill and Wang | isbn = 978-0-8090-8947-5 }} | |||
| * {{cite book | author-link1 = Gunther Stent | vauthors = Stent GS, Watson J |title=The Double Helix: A Personal Account of the Discovery of the Structure of DNA | url = https://archive.org/details/doublehelixpers00wats_0 | url-access = registration |publisher=Norton |location=New York |year=1980 |isbn=0-393-95075-1}} | |||
| * {{cite book | vauthors = Watson J | date = 2004 | title = DNA: The Secret of Life | publisher = Random House | isbn = 978-0-09-945184-6 }} | |||
| * {{cite book | author-link = Maurice Wilkins | vauthors = Wilkins M |title=The third man of the double helix the autobiography of Maurice Wilkins |publisher=University Press |location=Cambridge, England |year=2003 |isbn=0-19-860665-6}} | |||
| {{refend}} | |||
| == External links == | |||
| {{Library resources box | |||
| |onlinebooks=yes | |||
| |by=no | |||
| |lcheading= DNA | |||
| |label=DNA | |||
| }} | |||
| {{Spoken Misplaced Pages|dna.ogg|date=12 February 2007}} | |||
| * | |||
| * From the official Nobel Prize web site | |||
| * | * | ||
| * | * | ||
| * , '']'' | * , '']'' | ||
| * {{Proteopedia|DNA}} | |||
| * National Centre for Biotechnology Education | |||
| * {{Proteopedia|Forms_of_DNA}} | |||
| * | |||
| * ENCODE home page at ] | |||
| * —''DNA from the Beginning'' Study Guide | |||
| * National Centre for Biotechnology Education | |||
| * | |||
| * – ''DNA from the Beginning'' Study Guide | |||
| * {{cite journal |author=Olby R |authorlink=Robert Olby |title=Quiet debut for the double helix |journal=Nature |volume=421 |issue=6921 |pages=402–5 |year=2003 |month=January |pmid=12540907 |doi=10.1038/nature01397 |url=http://chem-faculty.ucsd.edu/joseph/CHEM13/DNA1.pdf}} | |||
| * {{PDB Molecule of the Month| |
* {{PDB Molecule of the Month|23|DNA}} | ||
| * . '']'', June 1953. First American newspaper coverage of the discovery of the DNA structure | |||
| * | |||
| * Another DNA Learning Center site on DNA, genes, and heredity from Mendel to the human genome project. | |||
| * at Mandeville Special Collections Library, Geisel Library, ] | |||
| * at Mandeville Special Collections Library, ] | |||
| * —watch videos and participate in real-time chat with top scientists | |||
| * See , Nature, 5 April 2013. | |||
| * {{cite news |title=Clue to chemistry of heredity found |work=] |date=Saturday, June 13, 1953 |url=http://www.nytimes.com/packages/pdf/science/dna-article.pdf}} The first American newspaper coverage of the discovery of the DNA structure. | |||
| * from ]: Huntington's Disease Outreach Project for Education at Stanford | |||
| {{Nucleic acids}} | |||
| {{Genetics}} | {{Genetics}} | ||
| {{ |
{{Gene expression}} | ||
| {{Nucleic acids}} | |||
| {{Subject bar | |||
| |commons = yes | |||
| |d = yes | |||
| |v = yes | |||
| |portal1 = Biology | |||
| }} | |||
| {{Authority control}} | |||
| {{DEFAULTSORT:Dna}} | |||
| <!--Categories--> | |||
| ] | ] | ||
| ] | ] | ||
| ] | |||
| ] | ] | ||
| ] | ] | ||
| {{Link FA|ca}} | |||
| {{Link FA|de}} | |||
| {{Link FA|es}} | |||
| {{Link FA|eu}} | |||
| {{Link FA|pt}} | |||
| {{Link FA|ru}} | |||
| {{Link FA|uk}} | |||
| {{Link FA|zh}} | |||
| <!--Interwiki--> | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
Latest revision as of 18:18, 22 December 2024
Molecule that carries genetic informationFor a non-technical introduction to the topic, see Introduction to genetics. For other uses, see DNA (disambiguation).
 Chromosome
(10 - 10 bp)
DNA
Gene
(10 - 10 bp )
Function
Chromosome
(10 - 10 bp)
DNA
Gene
(10 - 10 bp )
Function
|

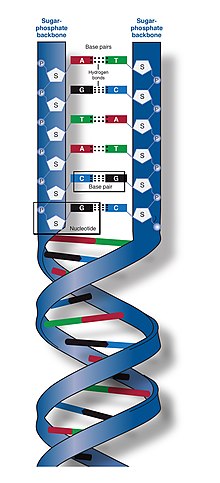
| Part of a series on |
| Genetics |
|---|
 |
|
Key components
|
| History and topics |
|
Research
|
| Fields |
| Personalized medicine |
Deoxyribonucleic acid (/diːˈɒksɪˌraɪboʊnjuːˌkliːɪk, -ˌkleɪ-/ ; DNA) is a polymer composed of two polynucleotide chains that coil around each other to form a double helix. The polymer carries genetic instructions for the development, functioning, growth and reproduction of all known organisms and many viruses. DNA and ribonucleic acid (RNA) are nucleic acids. Alongside proteins, lipids and complex carbohydrates (polysaccharides), nucleic acids are one of the four major types of macromolecules that are essential for all known forms of life.
The two DNA strands are known as polynucleotides as they are composed of simpler monomeric units called nucleotides. Each nucleotide is composed of one of four nitrogen-containing nucleobases (cytosine , guanine , adenine or thymine ), a sugar called deoxyribose, and a phosphate group. The nucleotides are joined to one another in a chain by covalent bonds (known as the phosphodiester linkage) between the sugar of one nucleotide and the phosphate of the next, resulting in an alternating sugar-phosphate backbone. The nitrogenous bases of the two separate polynucleotide strands are bound together, according to base pairing rules (A with T and C with G), with hydrogen bonds to make double-stranded DNA. The complementary nitrogenous bases are divided into two groups, the single-ringed pyrimidines and the double-ringed purines. In DNA, the pyrimidines are thymine and cytosine; the purines are adenine and guanine.
Both strands of double-stranded DNA store the same biological information. This information is replicated when the two strands separate. A large part of DNA (more than 98% for humans) is non-coding, meaning that these sections do not serve as patterns for protein sequences. The two strands of DNA run in opposite directions to each other and are thus antiparallel. Attached to each sugar is one of four types of nucleobases (or bases). It is the sequence of these four nucleobases along the backbone that encodes genetic information. RNA strands are created using DNA strands as a template in a process called transcription, where DNA bases are exchanged for their corresponding bases except in the case of thymine (T), for which RNA substitutes uracil (U). Under the genetic code, these RNA strands specify the sequence of amino acids within proteins in a process called translation.
Within eukaryotic cells, DNA is organized into long structures called chromosomes. Before typical cell division, these chromosomes are duplicated in the process of DNA replication, providing a complete set of chromosomes for each daughter cell. Eukaryotic organisms (animals, plants, fungi and protists) store most of their DNA inside the cell nucleus as nuclear DNA, and some in the mitochondria as mitochondrial DNA or in chloroplasts as chloroplast DNA. In contrast, prokaryotes (bacteria and archaea) store their DNA only in the cytoplasm, in circular chromosomes. Within eukaryotic chromosomes, chromatin proteins, such as histones, compact and organize DNA. These compacting structures guide the interactions between DNA and other proteins, helping control which parts of the DNA are transcribed.
Properties
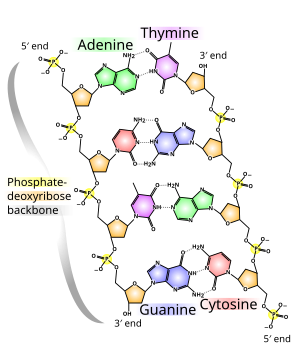
DNA is a long polymer made from repeating units called nucleotides. The structure of DNA is dynamic along its length, being capable of coiling into tight loops and other shapes. In all species it is composed of two helical chains, bound to each other by hydrogen bonds. Both chains are coiled around the same axis, and have the same pitch of 34 ångströms (3.4 nm). The pair of chains have a radius of 10 Å (1.0 nm). According to another study, when measured in a different solution, the DNA chain measured 22–26 Å (2.2–2.6 nm) wide, and one nucleotide unit measured 3.3 Å (0.33 nm) long. The buoyant density of most DNA is 1.7g/cm.
DNA does not usually exist as a single strand, but instead as a pair of strands that are held tightly together. These two long strands coil around each other, in the shape of a double helix. The nucleotide contains both a segment of the backbone of the molecule (which holds the chain together) and a nucleobase (which interacts with the other DNA strand in the helix). A nucleobase linked to a sugar is called a nucleoside, and a base linked to a sugar and to one or more phosphate groups is called a nucleotide. A biopolymer comprising multiple linked nucleotides (as in DNA) is called a polynucleotide.
The backbone of the DNA strand is made from alternating phosphate and sugar groups. The sugar in DNA is 2-deoxyribose, which is a pentose (five-carbon) sugar. The sugars are joined by phosphate groups that form phosphodiester bonds between the third and fifth carbon atoms of adjacent sugar rings. These are known as the 3′-end (three prime end), and 5′-end (five prime end) carbons, the prime symbol being used to distinguish these carbon atoms from those of the base to which the deoxyribose forms a glycosidic bond.
Therefore, any DNA strand normally has one end at which there is a phosphate group attached to the 5′ carbon of a ribose (the 5′ phosphoryl) and another end at which there is a free hydroxyl group attached to the 3′ carbon of a ribose (the 3′ hydroxyl). The orientation of the 3′ and 5′ carbons along the sugar-phosphate backbone confers directionality (sometimes called polarity) to each DNA strand. In a nucleic acid double helix, the direction of the nucleotides in one strand is opposite to their direction in the other strand: the strands are antiparallel. The asymmetric ends of DNA strands are said to have a directionality of five prime end (5′ ), and three prime end (3′), with the 5′ end having a terminal phosphate group and the 3′ end a terminal hydroxyl group. One major difference between DNA and RNA is the sugar, with the 2-deoxyribose in DNA being replaced by the related pentose sugar ribose in RNA.
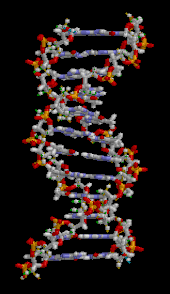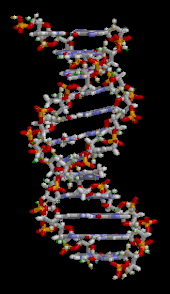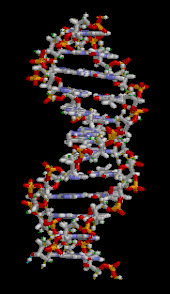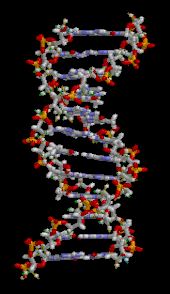
The DNA double helix is stabilized primarily by two forces: hydrogen bonds between nucleotides and base-stacking interactions among aromatic nucleobases. The four bases found in DNA are adenine (A), cytosine (C), guanine (G) and thymine (T). These four bases are attached to the sugar-phosphate to form the complete nucleotide, as shown for adenosine monophosphate. Adenine pairs with thymine and guanine pairs with cytosine, forming A-T and G-C base pairs.
Nucleobase classification
The nucleobases are classified into two types: the purines, A and G, which are fused five- and six-membered heterocyclic compounds, and the pyrimidines, the six-membered rings C and T. A fifth pyrimidine nucleobase, uracil (U), usually takes the place of thymine in RNA and differs from thymine by lacking a methyl group on its ring. In addition to RNA and DNA, many artificial nucleic acid analogues have been created to study the properties of nucleic acids, or for use in biotechnology.
Non-canonical bases
Modified bases occur in DNA. The first of these recognized was 5-methylcytosine, which was found in the genome of Mycobacterium tuberculosis in 1925. The reason for the presence of these noncanonical bases in bacterial viruses (bacteriophages) is to avoid the restriction enzymes present in bacteria. This enzyme system acts at least in part as a molecular immune system protecting bacteria from infection by viruses. Modifications of the bases cytosine and adenine, the more common and modified DNA bases, play vital roles in the epigenetic control of gene expression in plants and animals.
A number of noncanonical bases are known to occur in DNA. Most of these are modifications of the canonical bases plus uracil.
- Modified Adenine
- N6-carbamoyl-methyladenine
- N6-methyadenine
- Modified Guanine
- 7-Deazaguanine
- 7-Methylguanine
- Modified Cytosine
- N4-Methylcytosine
- 5-Carboxylcytosine
- 5-Formylcytosine
- 5-Glycosylhydroxymethylcytosine
- 5-Hydroxycytosine
- 5-Methylcytosine
- Modified Thymidine
- α-Glutamythymidine
- α-Putrescinylthymine
- Uracil and modifications
- Base J
- Uracil
- 5-Dihydroxypentauracil
- 5-Hydroxymethyldeoxyuracil
- Others
- Deoxyarchaeosine
- 2,6-Diaminopurine (2-Aminoadenine)
Grooves
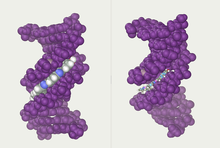
Twin helical strands form the DNA backbone. Another double helix may be found tracing the spaces, or grooves, between the strands. These voids are adjacent to the base pairs and may provide a binding site. As the strands are not symmetrically located with respect to each other, the grooves are unequally sized. The major groove is 22 ångströms (2.2 nm) wide, while the minor groove is 12 Å (1.2 nm) in width. Due to the larger width of the major groove, the edges of the bases are more accessible in the major groove than in the minor groove. As a result, proteins such as transcription factors that can bind to specific sequences in double-stranded DNA usually make contact with the sides of the bases exposed in the major groove. This situation varies in unusual conformations of DNA within the cell (see below), but the major and minor grooves are always named to reflect the differences in width that would be seen if the DNA was twisted back into the ordinary B form.
Base pairing
Further information: Base pair
|
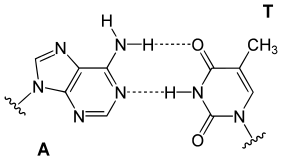
|
In a DNA double helix, each type of nucleobase on one strand bonds with just one type of nucleobase on the other strand. This is called complementary base pairing. Purines form hydrogen bonds to pyrimidines, with adenine bonding only to thymine in two hydrogen bonds, and cytosine bonding only to guanine in three hydrogen bonds. This arrangement of two nucleotides binding together across the double helix (from six-carbon ring to six-carbon ring) is called a Watson-Crick base pair. DNA with high GC-content is more stable than DNA with low GC-content. A Hoogsteen base pair (hydrogen bonding the 6-carbon ring to the 5-carbon ring) is a rare variation of base-pairing. As hydrogen bonds are not covalent, they can be broken and rejoined relatively easily. The two strands of DNA in a double helix can thus be pulled apart like a zipper, either by a mechanical force or high temperature. As a result of this base pair complementarity, all the information in the double-stranded sequence of a DNA helix is duplicated on each strand, which is vital in DNA replication. This reversible and specific interaction between complementary base pairs is critical for all the functions of DNA in organisms.
ssDNA vs. dsDNA
Most DNA molecules are actually two polymer strands, bound together in a helical fashion by noncovalent bonds; this double-stranded (dsDNA) structure is maintained largely by the intrastrand base stacking interactions, which are strongest for G,C stacks. The two strands can come apart—a process known as melting—to form two single-stranded DNA (ssDNA) molecules. Melting occurs at high temperatures, low salt and high pH (low pH also melts DNA, but since DNA is unstable due to acid depurination, low pH is rarely used).
The stability of the dsDNA form depends not only on the GC-content (% G,C basepairs) but also on sequence (since stacking is sequence specific) and also length (longer molecules are more stable). The stability can be measured in various ways; a common way is the melting temperature (also called Tm value), which is the temperature at which 50% of the double-strand molecules are converted to single-strand molecules; melting temperature is dependent on ionic strength and the concentration of DNA. As a result, it is both the percentage of GC base pairs and the overall length of a DNA double helix that determines the strength of the association between the two strands of DNA. Long DNA helices with a high GC-content have more strongly interacting strands, while short helices with high AT content have more weakly interacting strands. In biology, parts of the DNA double helix that need to separate easily, such as the TATAAT Pribnow box in some promoters, tend to have a high AT content, making the strands easier to pull apart.
In the laboratory, the strength of this interaction can be measured by finding the melting temperature Tm necessary to break half of the hydrogen bonds. When all the base pairs in a DNA double helix melt, the strands separate and exist in solution as two entirely independent molecules. These single-stranded DNA molecules have no single common shape, but some conformations are more stable than others.
Amount

In humans, the total female diploid nuclear genome per cell extends for 6.37 Gigabase pairs (Gbp), is 208.23 cm long and weighs 6.51 picograms (pg). Male values are 6.27 Gbp, 205.00 cm, 6.41 pg. Each DNA polymer can contain hundreds of millions of nucleotides, such as in chromosome 1. Chromosome 1 is the largest human chromosome with approximately 220 million base pairs, and would be 85 mm long if straightened.
In eukaryotes, in addition to nuclear DNA, there is also mitochondrial DNA (mtDNA) which encodes certain proteins used by the mitochondria. The mtDNA is usually relatively small in comparison to the nuclear DNA. For example, the human mitochondrial DNA forms closed circular molecules, each of which contains 16,569 DNA base pairs, with each such molecule normally containing a full set of the mitochondrial genes. Each human mitochondrion contains, on average, approximately 5 such mtDNA molecules. Each human cell contains approximately 100 mitochondria, giving a total number of mtDNA molecules per human cell of approximately 500. However, the amount of mitochondria per cell also varies by cell type, and an egg cell can contain 100,000 mitochondria, corresponding to up to 1,500,000 copies of the mitochondrial genome (constituting up to 90% of the DNA of the cell).
Sense and antisense
Further information: Sense (molecular biology)A DNA sequence is called a "sense" sequence if it is the same as that of a messenger RNA copy that is translated into protein. The sequence on the opposite strand is called the "antisense" sequence. Both sense and antisense sequences can exist on different parts of the same strand of DNA (i.e. both strands can contain both sense and antisense sequences). In both prokaryotes and eukaryotes, antisense RNA sequences are produced, but the functions of these RNAs are not entirely clear. One proposal is that antisense RNAs are involved in regulating gene expression through RNA-RNA base pairing.
A few DNA sequences in prokaryotes and eukaryotes, and more in plasmids and viruses, blur the distinction between sense and antisense strands by having overlapping genes. In these cases, some DNA sequences do double duty, encoding one protein when read along one strand, and a second protein when read in the opposite direction along the other strand. In bacteria, this overlap may be involved in the regulation of gene transcription, while in viruses, overlapping genes increase the amount of information that can be encoded within the small viral genome.
Supercoiling
Further information: DNA supercoilDNA can be twisted like a rope in a process called DNA supercoiling. With DNA in its "relaxed" state, a strand usually circles the axis of the double helix once every 10.4 base pairs, but if the DNA is twisted the strands become more tightly or more loosely wound. If the DNA is twisted in the direction of the helix, this is positive supercoiling, and the bases are held more tightly together. If they are twisted in the opposite direction, this is negative supercoiling, and the bases come apart more easily. In nature, most DNA has slight negative supercoiling that is introduced by enzymes called topoisomerases. These enzymes are also needed to relieve the twisting stresses introduced into DNA strands during processes such as transcription and DNA replication.
Alternative DNA structures
Further information: Molecular Structure of Nucleic Acids: A Structure for Deoxyribose Nucleic Acid, Molecular models of DNA, and DNA structure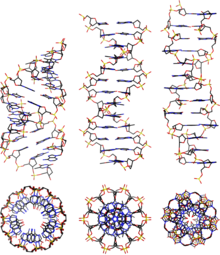
DNA exists in many possible conformations that include A-DNA, B-DNA, and Z-DNA forms, although only B-DNA and Z-DNA have been directly observed in functional organisms. The conformation that DNA adopts depends on the hydration level, DNA sequence, the amount and direction of supercoiling, chemical modifications of the bases, the type and concentration of metal ions, and the presence of polyamines in solution.
The first published reports of A-DNA X-ray diffraction patterns—and also B-DNA—used analyses based on Patterson functions that provided only a limited amount of structural information for oriented fibers of DNA. An alternative analysis was proposed by Wilkins et al. in 1953 for the in vivo B-DNA X-ray diffraction-scattering patterns of highly hydrated DNA fibers in terms of squares of Bessel functions. In the same journal, James Watson and Francis Crick presented their molecular modeling analysis of the DNA X-ray diffraction patterns to suggest that the structure was a double helix.
Although the B-DNA form is most common under the conditions found in cells, it is not a well-defined conformation but a family of related DNA conformations that occur at the high hydration levels present in cells. Their corresponding X-ray diffraction and scattering patterns are characteristic of molecular paracrystals with a significant degree of disorder.
Compared to B-DNA, the A-DNA form is a wider right-handed spiral, with a shallow, wide minor groove and a narrower, deeper major groove. The A form occurs under non-physiological conditions in partly dehydrated samples of DNA, while in the cell it may be produced in hybrid pairings of DNA and RNA strands, and in enzyme-DNA complexes. Segments of DNA where the bases have been chemically modified by methylation may undergo a larger change in conformation and adopt the Z form. Here, the strands turn about the helical axis in a left-handed spiral, the opposite of the more common B form. These unusual structures can be recognized by specific Z-DNA binding proteins and may be involved in the regulation of transcription.
Alternative DNA chemistry
Further information: hypothetical types of biochemistryFor many years, exobiologists have proposed the existence of a shadow biosphere, a postulated microbial biosphere of Earth that uses radically different biochemical and molecular processes than currently known life. One of the proposals was the existence of lifeforms that use arsenic instead of phosphorus in DNA. A report in 2010 of the possibility in the bacterium GFAJ-1 was announced, though the research was disputed, and evidence suggests the bacterium actively prevents the incorporation of arsenic into the DNA backbone and other biomolecules.
Quadruplex structures
Further information: G-quadruplex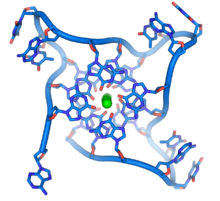
At the ends of the linear chromosomes are specialized regions of DNA called telomeres. The main function of these regions is to allow the cell to replicate chromosome ends using the enzyme telomerase, as the enzymes that normally replicate DNA cannot copy the extreme 3′ ends of chromosomes. These specialized chromosome caps also help protect the DNA ends, and stop the DNA repair systems in the cell from treating them as damage to be corrected. In human cells, telomeres are usually lengths of single-stranded DNA containing several thousand repeats of a simple TTAGGG sequence.
These guanine-rich sequences may stabilize chromosome ends by forming structures of stacked sets of four-base units, rather than the usual base pairs found in other DNA molecules. Here, four guanine bases, known as a guanine tetrad, form a flat plate. These flat four-base units then stack on top of each other to form a stable G-quadruplex structure. These structures are stabilized by hydrogen bonding between the edges of the bases and chelation of a metal ion in the centre of each four-base unit. Other structures can also be formed, with the central set of four bases coming from either a single strand folded around the bases, or several different parallel strands, each contributing one base to the central structure.
In addition to these stacked structures, telomeres also form large loop structures called telomere loops, or T-loops. Here, the single-stranded DNA curls around in a long circle stabilized by telomere-binding proteins. At the very end of the T-loop, the single-stranded telomere DNA is held onto a region of double-stranded DNA by the telomere strand disrupting the double-helical DNA and base pairing to one of the two strands. This triple-stranded structure is called a displacement loop or D-loop.
Branched DNA
Further information: Branched DNA and DNA nanotechnology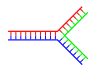
|

|
| Single branch | Multiple branches |
In DNA, fraying occurs when non-complementary regions exist at the end of an otherwise complementary double-strand of DNA. However, branched DNA can occur if a third strand of DNA is introduced and contains adjoining regions able to hybridize with the frayed regions of the pre-existing double-strand. Although the simplest example of branched DNA involves only three strands of DNA, complexes involving additional strands and multiple branches are also possible. Branched DNA can be used in nanotechnology to construct geometric shapes, see the section on uses in technology below.
Artificial bases
Main article: Nucleic acid analogueSeveral artificial nucleobases have been synthesized, and successfully incorporated in the eight-base DNA analogue named Hachimoji DNA. Dubbed S, B, P, and Z, these artificial bases are capable of bonding with each other in a predictable way (S–B and P–Z), maintain the double helix structure of DNA, and be transcribed to RNA. Their existence could be seen as an indication that there is nothing special about the four natural nucleobases that evolved on Earth. On the other hand, DNA is tightly related to RNA which does not only act as a transcript of DNA but also performs as molecular machines many tasks in cells. For this purpose it has to fold into a structure. It has been shown that to allow to create all possible structures at least four bases are required for the corresponding RNA, while a higher number is also possible but this would be against the natural principle of least effort.
Acidity
The phosphate groups of DNA give it similar acidic properties to phosphoric acid and it can be considered as a strong acid. It will be fully ionized at a normal cellular pH, releasing protons which leave behind negative charges on the phosphate groups. These negative charges protect DNA from breakdown by hydrolysis by repelling nucleophiles which could hydrolyze it.
Macroscopic appearance

Pure DNA extracted from cells forms white, stringy clumps.
Chemical modifications and altered DNA packaging
Base modifications and DNA packaging
Further information: DNA methylation and Chromatin remodeling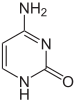
|

|

|
| cytosine | 5-methylcytosine | thymine |
The expression of genes is influenced by how the DNA is packaged in chromosomes, in a structure called chromatin. Base modifications can be involved in packaging, with regions that have low or no gene expression usually containing high levels of methylation of cytosine bases. DNA packaging and its influence on gene expression can also occur by covalent modifications of the histone protein core around which DNA is wrapped in the chromatin structure or else by remodeling carried out by chromatin remodeling complexes (see Chromatin remodeling). There is, further, crosstalk between DNA methylation and histone modification, so they can coordinately affect chromatin and gene expression.
For one example, cytosine methylation produces 5-methylcytosine, which is important for X-inactivation of chromosomes. The average level of methylation varies between organisms—the worm Caenorhabditis elegans lacks cytosine methylation, while vertebrates have higher levels, with up to 1% of their DNA containing 5-methylcytosine. Despite the importance of 5-methylcytosine, it can deaminate to leave a thymine base, so methylated cytosines are particularly prone to mutations. Other base modifications include adenine methylation in bacteria, the presence of 5-hydroxymethylcytosine in the brain, and the glycosylation of uracil to produce the "J-base" in kinetoplastids.
Damage
Further information: DNA damage (naturally occurring), Mutation, and DNA damage theory of aging
DNA can be damaged by many sorts of mutagens, which change the DNA sequence. Mutagens include oxidizing agents, alkylating agents and also high-energy electromagnetic radiation such as ultraviolet light and X-rays. The type of DNA damage produced depends on the type of mutagen. For example, UV light can damage DNA by producing thymine dimers, which are cross-links between pyrimidine bases. On the other hand, oxidants such as free radicals or hydrogen peroxide produce multiple forms of damage, including base modifications, particularly of guanosine, and double-strand breaks. A typical human cell contains about 150,000 bases that have suffered oxidative damage. Of these oxidative lesions, the most dangerous are double-strand breaks, as these are difficult to repair and can produce point mutations, insertions, deletions from the DNA sequence, and chromosomal translocations. These mutations can cause cancer. Because of inherent limits in the DNA repair mechanisms, if humans lived long enough, they would all eventually develop cancer. DNA damages that are naturally occurring, due to normal cellular processes that produce reactive oxygen species, the hydrolytic activities of cellular water, etc., also occur frequently. Although most of these damages are repaired, in any cell some DNA damage may remain despite the action of repair processes. These remaining DNA damages accumulate with age in mammalian postmitotic tissues. This accumulation appears to be an important underlying cause of aging.
Many mutagens fit into the space between two adjacent base pairs, this is called intercalation. Most intercalators are aromatic and planar molecules; examples include ethidium bromide, acridines, daunomycin, and doxorubicin. For an intercalator to fit between base pairs, the bases must separate, distorting the DNA strands by unwinding of the double helix. This inhibits both transcription and DNA replication, causing toxicity and mutations. As a result, DNA intercalators may be carcinogens, and in the case of thalidomide, a teratogen. Others such as benzopyrene diol epoxide and aflatoxin form DNA adducts that induce errors in replication. Nevertheless, due to their ability to inhibit DNA transcription and replication, other similar toxins are also used in chemotherapy to inhibit rapidly growing cancer cells.
Biological functions
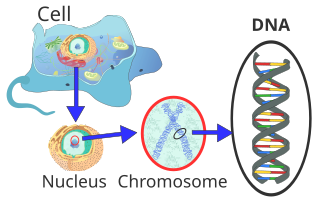
DNA usually occurs as linear chromosomes in eukaryotes, and circular chromosomes in prokaryotes. The set of chromosomes in a cell makes up its genome; the human genome has approximately 3 billion base pairs of DNA arranged into 46 chromosomes. The information carried by DNA is held in the sequence of pieces of DNA called genes. Transmission of genetic information in genes is achieved via complementary base pairing. For example, in transcription, when a cell uses the information in a gene, the DNA sequence is copied into a complementary RNA sequence through the attraction between the DNA and the correct RNA nucleotides. Usually, this RNA copy is then used to make a matching protein sequence in a process called translation, which depends on the same interaction between RNA nucleotides. In an alternative fashion, a cell may copy its genetic information in a process called DNA replication. The details of these functions are covered in other articles; here the focus is on the interactions between DNA and other molecules that mediate the function of the genome.
Genes and genomes
Further information: Cell nucleus, Chromatin, Chromosome, Gene, and Noncoding DNAGenomic DNA is tightly and orderly packed in the process called DNA condensation, to fit the small available volumes of the cell. In eukaryotes, DNA is located in the cell nucleus, with small amounts in mitochondria and chloroplasts. In prokaryotes, the DNA is held within an irregularly shaped body in the cytoplasm called the nucleoid. The genetic information in a genome is held within genes, and the complete set of this information in an organism is called its genotype. A gene is a unit of heredity and is a region of DNA that influences a particular characteristic in an organism. Genes contain an open reading frame that can be transcribed, and regulatory sequences such as promoters and enhancers, which control transcription of the open reading frame.
In many species, only a small fraction of the total sequence of the genome encodes protein. For example, only about 1.5% of the human genome consists of protein-coding exons, with over 50% of human DNA consisting of non-coding repetitive sequences. The reasons for the presence of so much noncoding DNA in eukaryotic genomes and the extraordinary differences in genome size, or C-value, among species, represent a long-standing puzzle known as the "C-value enigma". However, some DNA sequences that do not code protein may still encode functional non-coding RNA molecules, which are involved in the regulation of gene expression.

Some noncoding DNA sequences play structural roles in chromosomes. Telomeres and centromeres typically contain few genes but are important for the function and stability of chromosomes. An abundant form of noncoding DNA in humans are pseudogenes, which are copies of genes that have been disabled by mutation. These sequences are usually just molecular fossils, although they can occasionally serve as raw genetic material for the creation of new genes through the process of gene duplication and divergence.
Transcription and translation
Further information: Genetic code, Transcription (genetics), and Protein biosynthesisA gene is a sequence of DNA that contains genetic information and can influence the phenotype of an organism. Within a gene, the sequence of bases along a DNA strand defines a messenger RNA sequence, which then defines one or more protein sequences. The relationship between the nucleotide sequences of genes and the amino-acid sequences of proteins is determined by the rules of translation, known collectively as the genetic code. The genetic code consists of three-letter 'words' called codons formed from a sequence of three nucleotides (e.g. ACT, CAG, TTT).
In transcription, the codons of a gene are copied into messenger RNA by RNA polymerase. This RNA copy is then decoded by a ribosome that reads the RNA sequence by base-pairing the messenger RNA to transfer RNA, which carries amino acids. Since there are 4 bases in 3-letter combinations, there are 64 possible codons (4 combinations). These encode the twenty standard amino acids, giving most amino acids more than one possible codon. There are also three 'stop' or 'nonsense' codons signifying the end of the coding region; these are the TAG, TAA, and TGA codons, (UAG, UAA, and UGA on the mRNA).
Replication
Further information: DNA replication
Cell division is essential for an organism to grow, but, when a cell divides, it must replicate the DNA in its genome so that the two daughter cells have the same genetic information as their parent. The double-stranded structure of DNA provides a simple mechanism for DNA replication. Here, the two strands are separated and then each strand's complementary DNA sequence is recreated by an enzyme called DNA polymerase. This enzyme makes the complementary strand by finding the correct base through complementary base pairing and bonding it onto the original strand. As DNA polymerases can only extend a DNA strand in a 5′ to 3′ direction, different mechanisms are used to copy the antiparallel strands of the double helix. In this way, the base on the old strand dictates which base appears on the new strand, and the cell ends up with a perfect copy of its DNA.
Extracellular nucleic acids
Naked extracellular DNA (eDNA), most of it released by cell death, is nearly ubiquitous in the environment. Its concentration in soil may be as high as 2 μg/L, and its concentration in natural aquatic environments may be as high at 88 μg/L. Various possible functions have been proposed for eDNA: it may be involved in horizontal gene transfer; it may provide nutrients; and it may act as a buffer to recruit or titrate ions or antibiotics. Extracellular DNA acts as a functional extracellular matrix component in the biofilms of several bacterial species. It may act as a recognition factor to regulate the attachment and dispersal of specific cell types in the biofilm; it may contribute to biofilm formation; and it may contribute to the biofilm's physical strength and resistance to biological stress.
Cell-free fetal DNA is found in the blood of the mother, and can be sequenced to determine a great deal of information about the developing fetus.
Under the name of environmental DNA eDNA has seen increased use in the natural sciences as a survey tool for ecology, monitoring the movements and presence of species in water, air, or on land, and assessing an area's biodiversity.
Neutrophil extracellular traps
Main article: Neutrophil extracellular trapsNeutrophil extracellular traps (NETs) are networks of extracellular fibers, primarily composed of DNA, which allow neutrophils, a type of white blood cell, to kill extracellular pathogens while minimizing damage to the host cells.
Interactions with proteins
All the functions of DNA depend on interactions with proteins. These protein interactions can be non-specific, or the protein can bind specifically to a single DNA sequence. Enzymes can also bind to DNA and of these, the polymerases that copy the DNA base sequence in transcription and DNA replication are particularly important.
DNA-binding proteins
Further information: DNA-binding protein
Structural proteins that bind DNA are well-understood examples of non-specific DNA-protein interactions. Within chromosomes, DNA is held in complexes with structural proteins. These proteins organize the DNA into a compact structure called chromatin. In eukaryotes, this structure involves DNA binding to a complex of small basic proteins called histones, while in prokaryotes multiple types of proteins are involved. The histones form a disk-shaped complex called a nucleosome, which contains two complete turns of double-stranded DNA wrapped around its surface. These non-specific interactions are formed through basic residues in the histones, making ionic bonds to the acidic sugar-phosphate backbone of the DNA, and are thus largely independent of the base sequence. Chemical modifications of these basic amino acid residues include methylation, phosphorylation, and acetylation. These chemical changes alter the strength of the interaction between the DNA and the histones, making the DNA more or less accessible to transcription factors and changing the rate of transcription. Other non-specific DNA-binding proteins in chromatin include the high-mobility group proteins, which bind to bent or distorted DNA. These proteins are important in bending arrays of nucleosomes and arranging them into the larger structures that make up chromosomes.
A distinct group of DNA-binding proteins is the DNA-binding proteins that specifically bind single-stranded DNA. In humans, replication protein A is the best-understood member of this family and is used in processes where the double helix is separated, including DNA replication, recombination, and DNA repair. These binding proteins seem to stabilize single-stranded DNA and protect it from forming stem-loops or being degraded by nucleases.

In contrast, other proteins have evolved to bind to particular DNA sequences. The most intensively studied of these are the various transcription factors, which are proteins that regulate transcription. Each transcription factor binds to one particular set of DNA sequences and activates or inhibits the transcription of genes that have these sequences close to their promoters. The transcription factors do this in two ways. Firstly, they can bind the RNA polymerase responsible for transcription, either directly or through other mediator proteins; this locates the polymerase at the promoter and allows it to begin transcription. Alternatively, transcription factors can bind enzymes that modify the histones at the promoter. This changes the accessibility of the DNA template to the polymerase.
As these DNA targets can occur throughout an organism's genome, changes in the activity of one type of transcription factor can affect thousands of genes. Consequently, these proteins are often the targets of the signal transduction processes that control responses to environmental changes or cellular differentiation and development. The specificity of these transcription factors' interactions with DNA come from the proteins making multiple contacts to the edges of the DNA bases, allowing them to "read" the DNA sequence. Most of these base-interactions are made in the major groove, where the bases are most accessible.
DNA-modifying enzymes
Nucleases and ligases
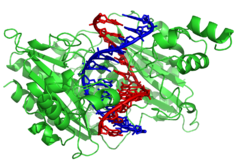
Nucleases are enzymes that cut DNA strands by catalyzing the hydrolysis of the phosphodiester bonds. Nucleases that hydrolyse nucleotides from the ends of DNA strands are called exonucleases, while endonucleases cut within strands. The most frequently used nucleases in molecular biology are the restriction endonucleases, which cut DNA at specific sequences. For instance, the EcoRV enzyme shown to the left recognizes the 6-base sequence 5′-GATATC-3′ and makes a cut at the horizontal line. In nature, these enzymes protect bacteria against phage infection by digesting the phage DNA when it enters the bacterial cell, acting as part of the restriction modification system. In technology, these sequence-specific nucleases are used in molecular cloning and DNA fingerprinting.
Enzymes called DNA ligases can rejoin cut or broken DNA strands. Ligases are particularly important in lagging strand DNA replication, as they join the short segments of DNA produced at the replication fork into a complete copy of the DNA template. They are also used in DNA repair and genetic recombination.
Topoisomerases and helicases
Topoisomerases are enzymes with both nuclease and ligase activity. These proteins change the amount of supercoiling in DNA. Some of these enzymes work by cutting the DNA helix and allowing one section to rotate, thereby reducing its level of supercoiling; the enzyme then seals the DNA break. Other types of these enzymes are capable of cutting one DNA helix and then passing a second strand of DNA through this break, before rejoining the helix. Topoisomerases are required for many processes involving DNA, such as DNA replication and transcription.
Helicases are proteins that are a type of molecular motor. They use the chemical energy in nucleoside triphosphates, predominantly adenosine triphosphate (ATP), to break hydrogen bonds between bases and unwind the DNA double helix into single strands. These enzymes are essential for most processes where enzymes need to access the DNA bases.
Polymerases
Polymerases are enzymes that synthesize polynucleotide chains from nucleoside triphosphates. The sequence of their products is created based on existing polynucleotide chains—which are called templates. These enzymes function by repeatedly adding a nucleotide to the 3′ hydroxyl group at the end of the growing polynucleotide chain. As a consequence, all polymerases work in a 5′ to 3′ direction. In the active site of these enzymes, the incoming nucleoside triphosphate base-pairs to the template: this allows polymerases to accurately synthesize the complementary strand of their template. Polymerases are classified according to the type of template that they use.
In DNA replication, DNA-dependent DNA polymerases make copies of DNA polynucleotide chains. To preserve biological information, it is essential that the sequence of bases in each copy are precisely complementary to the sequence of bases in the template strand. Many DNA polymerases have a proofreading activity. Here, the polymerase recognizes the occasional mistakes in the synthesis reaction by the lack of base pairing between the mismatched nucleotides. If a mismatch is detected, a 3′ to 5′ exonuclease activity is activated and the incorrect base removed. In most organisms, DNA polymerases function in a large complex called the replisome that contains multiple accessory subunits, such as the DNA clamp or helicases.
RNA-dependent DNA polymerases are a specialized class of polymerases that copy the sequence of an RNA strand into DNA. They include reverse transcriptase, which is a viral enzyme involved in the infection of cells by retroviruses, and telomerase, which is required for the replication of telomeres. For example, HIV reverse transcriptase is an enzyme for AIDS virus replication. Telomerase is an unusual polymerase because it contains its own RNA template as part of its structure. It synthesizes telomeres at the ends of chromosomes. Telomeres prevent fusion of the ends of neighboring chromosomes and protect chromosome ends from damage.
Transcription is carried out by a DNA-dependent RNA polymerase that copies the sequence of a DNA strand into RNA. To begin transcribing a gene, the RNA polymerase binds to a sequence of DNA called a promoter and separates the DNA strands. It then copies the gene sequence into a messenger RNA transcript until it reaches a region of DNA called the terminator, where it halts and detaches from the DNA. As with human DNA-dependent DNA polymerases, RNA polymerase II, the enzyme that transcribes most of the genes in the human genome, operates as part of a large protein complex with multiple regulatory and accessory subunits.
Genetic recombination
Further information: Genetic recombination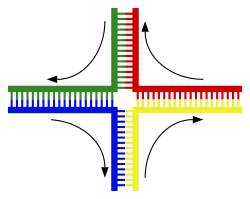
|

|
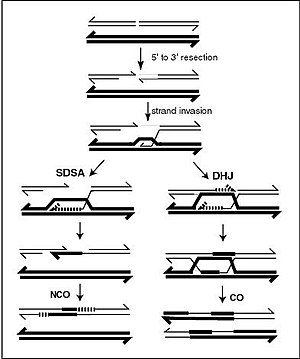
A DNA helix usually does not interact with other segments of DNA, and in human cells, the different chromosomes even occupy separate areas in the nucleus called "chromosome territories". This physical separation of different chromosomes is important for the ability of DNA to function as a stable repository for information, as one of the few times chromosomes interact is in chromosomal crossover which occurs during sexual reproduction, when genetic recombination occurs. Chromosomal crossover is when two DNA helices break, swap a section and then rejoin.
Recombination allows chromosomes to exchange genetic information and produces new combinations of genes, which increases the efficiency of natural selection and can be important in the rapid evolution of new proteins. Genetic recombination can also be involved in DNA repair, particularly in the cell's response to double-strand breaks.
The most common form of chromosomal crossover is homologous recombination, where the two chromosomes involved share very similar sequences. Non-homologous recombination can be damaging to cells, as it can produce chromosomal translocations and genetic abnormalities. The recombination reaction is catalyzed by enzymes known as recombinases, such as RAD51. The first step in recombination is a double-stranded break caused by either an endonuclease or damage to the DNA. A series of steps catalyzed in part by the recombinase then leads to joining of the two helices by at least one Holliday junction, in which a segment of a single strand in each helix is annealed to the complementary strand in the other helix. The Holliday junction is a tetrahedral junction structure that can be moved along the pair of chromosomes, swapping one strand for another. The recombination reaction is then halted by cleavage of the junction and re-ligation of the released DNA. Only strands of like polarity exchange DNA during recombination. There are two types of cleavage: east-west cleavage and north–south cleavage. The north–south cleavage nicks both strands of DNA, while the east–west cleavage has one strand of DNA intact. The formation of a Holliday junction during recombination makes it possible for genetic diversity, genes to exchange on chromosomes, and expression of wild-type viral genomes.
Evolution
Further information: RNA world hypothesisDNA contains the genetic information that allows all forms of life to function, grow and reproduce. However, it is unclear how long in the 4-billion-year history of life DNA has performed this function, as it has been proposed that the earliest forms of life may have used RNA as their genetic material. RNA may have acted as the central part of early cell metabolism as it can both transmit genetic information and carry out catalysis as part of ribozymes. This ancient RNA world where nucleic acid would have been used for both catalysis and genetics may have influenced the evolution of the current genetic code based on four nucleotide bases. This would occur, since the number of different bases in such an organism is a trade-off between a small number of bases increasing replication accuracy and a large number of bases increasing the catalytic efficiency of ribozymes. However, there is no direct evidence of ancient genetic systems, as recovery of DNA from most fossils is impossible because DNA survives in the environment for less than one million years, and slowly degrades into short fragments in solution. Claims for older DNA have been made, most notably a report of the isolation of a viable bacterium from a salt crystal 250 million years old, but these claims are controversial.
Building blocks of DNA (adenine, guanine, and related organic molecules) may have been formed extraterrestrially in outer space. Complex DNA and RNA organic compounds of life, including uracil, cytosine, and thymine, have also been formed in the laboratory under conditions mimicking those found in outer space, using starting chemicals, such as pyrimidine, found in meteorites. Pyrimidine, like polycyclic aromatic hydrocarbons (PAHs), the most carbon-rich chemical found in the universe, may have been formed in red giants or in interstellar cosmic dust and gas clouds.
Ancient DNA has been recovered from ancient organisms at a timescale where genome evolution can be directly observed, including from extinct organisms up to millions of years old, such as the woolly mammoth.
Uses in technology
Genetic engineering
Further information: Molecular biology, Nucleic acid methods, and Genetic engineeringMethods have been developed to purify DNA from organisms, such as phenol-chloroform extraction, and to manipulate it in the laboratory, such as restriction digests and the polymerase chain reaction. Modern biology and biochemistry make intensive use of these techniques in recombinant DNA technology. Recombinant DNA is a man-made DNA sequence that has been assembled from other DNA sequences. They can be transformed into organisms in the form of plasmids or in the appropriate format, by using a viral vector. The genetically modified organisms produced can be used to produce products such as recombinant proteins, used in medical research, or be grown in agriculture.
DNA profiling
Further information: DNA profilingForensic scientists can use DNA in blood, semen, skin, saliva or hair found at a crime scene to identify a matching DNA of an individual, such as a perpetrator. This process is formally termed DNA profiling, also called DNA fingerprinting. In DNA profiling, the lengths of variable sections of repetitive DNA, such as short tandem repeats and minisatellites, are compared between people. This method is usually an extremely reliable technique for identifying a matching DNA. However, identification can be complicated if the scene is contaminated with DNA from several people. DNA profiling was developed in 1984 by British geneticist Sir Alec Jeffreys, and first used in forensic science to convict Colin Pitchfork in the 1988 Enderby murders case.
The development of forensic science and the ability to now obtain genetic matching on minute samples of blood, skin, saliva, or hair has led to re-examining many cases. Evidence can now be uncovered that was scientifically impossible at the time of the original examination. Combined with the removal of the double jeopardy law in some places, this can allow cases to be reopened where prior trials have failed to produce sufficient evidence to convince a jury. People charged with serious crimes may be required to provide a sample of DNA for matching purposes. The most obvious defense to DNA matches obtained forensically is to claim that cross-contamination of evidence has occurred. This has resulted in meticulous strict handling procedures with new cases of serious crime.
DNA profiling is also used successfully to positively identify victims of mass casualty incidents, bodies or body parts in serious accidents, and individual victims in mass war graves, via matching to family members.
DNA profiling is also used in DNA paternity testing to determine if someone is the biological parent or grandparent of a child with the probability of parentage is typically 99.99% when the alleged parent is biologically related to the child. Normal DNA sequencing methods happen after birth, but there are new methods to test paternity while a mother is still pregnant.
DNA enzymes or catalytic DNA
Further information: DeoxyribozymeDeoxyribozymes, also called DNAzymes or catalytic DNA, were first discovered in 1994. They are mostly single stranded DNA sequences isolated from a large pool of random DNA sequences through a combinatorial approach called in vitro selection or systematic evolution of ligands by exponential enrichment (SELEX). DNAzymes catalyze variety of chemical reactions including RNA-DNA cleavage, RNA-DNA ligation, amino acids phosphorylation-dephosphorylation, carbon-carbon bond formation, etc. DNAzymes can enhance catalytic rate of chemical reactions up to 100,000,000,000-fold over the uncatalyzed reaction. The most extensively studied class of DNAzymes is RNA-cleaving types which have been used to detect different metal ions and designing therapeutic agents. Several metal-specific DNAzymes have been reported including the GR-5 DNAzyme (lead-specific), the CA1-3 DNAzymes (copper-specific), the 39E DNAzyme (uranyl-specific) and the NaA43 DNAzyme (sodium-specific). The NaA43 DNAzyme, which is reported to be more than 10,000-fold selective for sodium over other metal ions, was used to make a real-time sodium sensor in cells.
Bioinformatics
Further information: BioinformaticsBioinformatics involves the development of techniques to store, data mine, search and manipulate biological data, including DNA nucleic acid sequence data. These have led to widely applied advances in computer science, especially string searching algorithms, machine learning, and database theory. String searching or matching algorithms, which find an occurrence of a sequence of letters inside a larger sequence of letters, were developed to search for specific sequences of nucleotides. The DNA sequence may be aligned with other DNA sequences to identify homologous sequences and locate the specific mutations that make them distinct. These techniques, especially multiple sequence alignment, are used in studying phylogenetic relationships and protein function. Data sets representing entire genomes' worth of DNA sequences, such as those produced by the Human Genome Project, are difficult to use without the annotations that identify the locations of genes and regulatory elements on each chromosome. Regions of DNA sequence that have the characteristic patterns associated with protein- or RNA-coding genes can be identified by gene finding algorithms, which allow researchers to predict the presence of particular gene products and their possible functions in an organism even before they have been isolated experimentally. Entire genomes may also be compared, which can shed light on the evolutionary history of particular organism and permit the examination of complex evolutionary events.
DNA nanotechnology
Further information: DNA nanotechnology
DNA nanotechnology uses the unique molecular recognition properties of DNA and other nucleic acids to create self-assembling branched DNA complexes with useful properties. DNA is thus used as a structural material rather than as a carrier of biological information. This has led to the creation of two-dimensional periodic lattices (both tile-based and using the DNA origami method) and three-dimensional structures in the shapes of polyhedra. Nanomechanical devices and algorithmic self-assembly have also been demonstrated, and these DNA structures have been used to template the arrangement of other molecules such as gold nanoparticles and streptavidin proteins. DNA and other nucleic acids are the basis of aptamers, synthetic oligonucleotide ligands for specific target molecules used in a range of biotechnology and biomedical applications.
History and anthropology
Further information: Phylogenetics and Genetic genealogyBecause DNA collects mutations over time, which are then inherited, it contains historical information, and, by comparing DNA sequences, geneticists can infer the evolutionary history of organisms, their phylogeny. This field of phylogenetics is a powerful tool in evolutionary biology. If DNA sequences within a species are compared, population geneticists can learn the history of particular populations. This can be used in studies ranging from ecological genetics to anthropology.
Information storage
Main article: DNA digital data storageDNA as a storage device for information has enormous potential since it has much higher storage density compared to electronic devices. However, high costs, slow read and write times (memory latency), and insufficient reliability has prevented its practical use.
History
Further information: History of molecular biology

DNA was first isolated by the Swiss physician Friedrich Miescher who, in 1869, discovered a microscopic substance in the pus of discarded surgical bandages. As it resided in the nuclei of cells, he called it "nuclein". In 1878, Albrecht Kossel isolated the non-protein component of "nuclein", nucleic acid, and later isolated its five primary nucleobases.
In 1909, Phoebus Levene identified the base, sugar, and phosphate nucleotide unit of RNA (then named "yeast nucleic acid"). In 1929, Levene identified deoxyribose sugar in "thymus nucleic acid" (DNA). Levene suggested that DNA consisted of a string of four nucleotide units linked together through the phosphate groups ("tetranucleotide hypothesis"). Levene thought the chain was short and the bases repeated in a fixed order. In 1927, Nikolai Koltsov proposed that inherited traits would be inherited via a "giant hereditary molecule" made up of "two mirror strands that would replicate in a semi-conservative fashion using each strand as a template". In 1928, Frederick Griffith in his experiment discovered that traits of the "smooth" form of Pneumococcus could be transferred to the "rough" form of the same bacteria by mixing killed "smooth" bacteria with the live "rough" form. This system provided the first clear suggestion that DNA carries genetic information.
In 1933, while studying virgin sea urchin eggs, Jean Brachet suggested that DNA is found in the cell nucleus and that RNA is present exclusively in the cytoplasm. At the time, "yeast nucleic acid" (RNA) was thought to occur only in plants, while "thymus nucleic acid" (DNA) only in animals. The latter was thought to be a tetramer, with the function of buffering cellular pH.
In 1937, William Astbury produced the first X-ray diffraction patterns that showed that DNA had a regular structure.
In 1943, Oswald Avery, along with co-workers Colin MacLeod and Maclyn McCarty, identified DNA as the transforming principle, supporting Griffith's suggestion (Avery–MacLeod–McCarty experiment). Erwin Chargaff developed and published observations now known as Chargaff's rules, stating that in DNA from any species of any organism, the amount of guanine should be equal to cytosine and the amount of adenine should be equal to thymine.

Late in 1951, Francis Crick started working with James Watson at the Cavendish Laboratory within the University of Cambridge. DNA's role in heredity was confirmed in 1952 when Alfred Hershey and Martha Chase in the Hershey–Chase experiment showed that DNA is the genetic material of the enterobacteria phage T2.
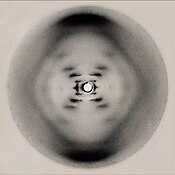
In May 1952, Raymond Gosling, a graduate student working under the supervision of Rosalind Franklin, took an X-ray diffraction image, labeled as "Photo 51", at high hydration levels of DNA. This photo was given to Watson and Crick by Maurice Wilkins and was critical to their obtaining the correct structure of DNA. Franklin told Crick and Watson that the backbones had to be on the outside. Before then, Linus Pauling, and Watson and Crick, had erroneous models with the chains inside and the bases pointing outwards. Franklin's identification of the space group for DNA crystals revealed to Crick that the two DNA strands were antiparallel. In February 1953, Linus Pauling and Robert Corey proposed a model for nucleic acids containing three intertwined chains, with the phosphates near the axis, and the bases on the outside. Watson and Crick completed their model, which is now accepted as the first correct model of the double helix of DNA. On 28 February 1953 Crick interrupted patrons' lunchtime at The Eagle pub in Cambridge, England to announce that he and Watson had "discovered the secret of life".
The 25 April 1953 issue of the journal Nature published a series of five articles giving the Watson and Crick double-helix structure DNA and evidence supporting it. The structure was reported in a letter titled "MOLECULAR STRUCTURE OF NUCLEIC ACIDS A Structure for Deoxyribose Nucleic Acid", in which they said, "It has not escaped our notice that the specific pairing we have postulated immediately suggests a possible copying mechanism for the genetic material." This letter was followed by a letter from Franklin and Gosling, which was the first publication of their own X-ray diffraction data and of their original analysis method. Then followed a letter by Wilkins and two of his colleagues, which contained an analysis of in vivo B-DNA X-ray patterns, and which supported the presence in vivo of the Watson and Crick structure.
In April 2023, scientists, based on new evidence, concluded that Rosalind Franklin was a contributor and "equal player" in the discovery process of DNA, rather than otherwise, as may have been presented subsequently after the time of the discovery.
In 1962, after Franklin's death, Watson, Crick, and Wilkins jointly received the Nobel Prize in Physiology or Medicine. Nobel Prizes are awarded only to living recipients. A debate continues about who should receive credit for the discovery.
In an influential presentation in 1957, Crick laid out the central dogma of molecular biology, which foretold the relationship between DNA, RNA, and proteins, and articulated the "adaptor hypothesis". Final confirmation of the replication mechanism that was implied by the double-helical structure followed in 1958 through the Meselson–Stahl experiment. Further work by Crick and co-workers showed that the genetic code was based on non-overlapping triplets of bases, called codons, allowing Har Gobind Khorana, Robert W. Holley, and Marshall Warren Nirenberg to decipher the genetic code. These findings represent the birth of molecular biology.
In 1986, DNA analysis was first used for criminal investigative purposes when police in the UK requested Alec Jeffreys of the University of Leicester to verify or disprove a suspect's rape-murder "confession". In this particular case, the suspect had confessed to two rape-murders, but had later retracted his confession. DNA testing at the university labs soon disproved the veracity of the suspect's original "confession", and the suspect was exonerated from the murder-rape charges.
See also
- Autosome – Any chromosome other than a sex chromosome
- Crystallography – Scientific study of crystal structures
- DNA Day – Holiday celebrated on April 25
- DNA microarray – Collection of microscopic DNA spots attached to a solid surface
- DNA sequencing – Process of determining the nucleic acid sequence
- Genetic disorder – Health problem caused by one or more abnormalities in the genome
- Genetic genealogy – DNA testing to infer relationships
- Haplotype – Group of genes from one parent
- Meiosis – Cell division producing haploid gametes
- Nucleic acid notation – Universal notation using the Roman characters A, C, G, and T to call the four DNA nucleotides
- Nucleic acid sequence – Succession of nucleotides in a nucleic acid
- Ribosomal DNA – Genes coding for ribosomal RNA
- Southern blot – DNA analysis technique
- X-ray scattering techniques – family of non-destructive analytical techniquesPages displaying wikidata descriptions as a fallback
- Xeno nucleic acid – Synthetic nucleic acid analogues
References
- "deoxyribonucleic acid". Merriam-Webster.com Dictionary. Merriam-Webster.
- Alberts B, Johnson A, Lewis J, Raff M, Roberts K, Walter P (2014). Molecular Biology of the Cell (6th ed.). Garland. p. Chapter 4: DNA, Chromosomes and Genomes. ISBN 978-0-8153-4432-2. Archived from the original on 14 July 2014.
- Purcell A. "DNA". Basic Biology. Archived from the original on 5 January 2017.
- "Uracil". Genome.gov. Retrieved 21 November 2019.
- Russell P (2001). iGenetics. New York: Benjamin Cummings. ISBN 0-8053-4553-1.
- Saenger W (1984). Principles of Nucleic Acid Structure. New York: Springer-Verlag. ISBN 0-387-90762-9.
- ^ Alberts B, Johnson A, Lewis J, Raff M, Roberts K, Peter W (2002). Molecular Biology of the Cell (Fourth ed.). New York and London: Garland Science. ISBN 0-8153-3218-1. OCLC 145080076. Archived from the original on 1 November 2016.
- Irobalieva RN, Fogg JM, Catanese DJ, Catanese DJ, Sutthibutpong T, Chen M, Barker AK, Ludtke SJ, Harris SA, Schmid MF, Chiu W, Zechiedrich L (October 2015). "Structural diversity of supercoiled DNA". Nature Communications. 6: 8440. Bibcode:2015NatCo...6.8440I. doi:10.1038/ncomms9440. ISSN 2041-1723. PMC 4608029. PMID 26455586.
- ^ Watson JD, Crick FH (April 1953). "Molecular structure of nucleic acids; a structure for deoxyribose nucleic acid" (PDF). Nature. 171 (4356): 737–38. Bibcode:1953Natur.171..737W. doi:10.1038/171737a0. ISSN 0028-0836. PMID 13054692. S2CID 4253007. Archived (PDF) from the original on 4 February 2007.
- Mandelkern M, Elias JG, Eden D, Crothers DM (October 1981). "The dimensions of DNA in solution". Journal of Molecular Biology. 152 (1): 153–61. doi:10.1016/0022-2836(81)90099-1. ISSN 0022-2836. PMID 7338906.
- Arrighi, Frances E.; Mandel, Manley; Bergendahl, Janet; Hsu, T. C. (June 1970). "Buoyant densities of DNA of mammals". Biochemical Genetics. 4 (3): 367–376. doi:10.1007/BF00485753. ISSN 0006-2928. PMID 4991030. S2CID 27950750.
- ^ Berg J, Tymoczko J, Stryer L (2002). Biochemistry. W.H. Freeman and Company. ISBN 0-7167-4955-6.
- IUPAC-IUB Commission on Biochemical Nomenclature (CBN) (December 1970). "Abbreviations and Symbols for Nucleic Acids, Polynucleotides and their Constituents. Recommendations 1970". The Biochemical Journal. 120 (3): 449–54. doi:10.1042/bj1200449. ISSN 0306-3283. PMC 1179624. PMID 5499957. Archived from the original on 5 February 2007.
- ^ Ghosh A, Bansal M (April 2003). "A glossary of DNA structures from A to Z". Acta Crystallographica Section D. 59 (Pt 4): 620–26. Bibcode:2003AcCrD..59..620G. doi:10.1107/S0907444903003251. ISSN 0907-4449. PMID 12657780.
- Edwards KJ, Brown DG, Spink N, Skelly JV, Neidle S. "RCSB PDB – 1D65: Molecular structure of the B-DNA dodecamer d(CGCAAATTTGCG)2. An examination of propeller twist and minor-groove water structure at 2.2 A resolution". www.rcsb.org. Retrieved 27 March 2023.
- Yakovchuk P, Protozanova E, Frank-Kamenetskii MD (2006). "Base-stacking and base-pairing contributions into thermal stability of the DNA double helix". Nucleic Acids Research. 34 (2): 564–74. doi:10.1093/nar/gkj454. ISSN 0305-1048. PMC 1360284. PMID 16449200.
- Tropp BE (2012). Molecular Biology (4th ed.). Sudbury, Mass.: Jones and Barlett Learning. ISBN 978-0-7637-8663-2.
- Carr S (1953). "Watson-Crick Structure of DNA". Memorial University of Newfoundland. Archived from the original on 19 July 2016. Retrieved 13 July 2016.
- Verma S, Eckstein F (1998). "Modified oligonucleotides: synthesis and strategy for users". Annual Review of Biochemistry. 67: 99–134. doi:10.1146/annurev.biochem.67.1.99. ISSN 0066-4154. PMID 9759484.
- Johnson TB, Coghill RD (1925). "Pyrimidines. CIII. The discovery of 5-methylcytosine in tuberculinic acid, the nucleic acid of the tubercle bacillus". Journal of the American Chemical Society. 47: 2838–44. doi:10.1021/ja01688a030. ISSN 0002-7863.
- Weigele P, Raleigh EA (October 2016). "Biosynthesis and Function of Modified Bases in Bacteria and Their Viruses". Chemical Reviews. 116 (20): 12655–12687. doi:10.1021/acs.chemrev.6b00114. ISSN 0009-2665. PMID 27319741.
- Kumar S, Chinnusamy V, Mohapatra T (2018). "Epigenetics of Modified DNA Bases: 5-Methylcytosine and Beyond". Frontiers in Genetics. 9: 640. doi:10.3389/fgene.2018.00640. ISSN 1664-8021. PMC 6305559. PMID 30619465.
- Carell T, Kurz MQ, Müller M, Rossa M, Spada F (April 2018). "Non-canonical Bases in the Genome: The Regulatory Information Layer in DNA". Angewandte Chemie. 57 (16): 4296–4312. doi:10.1002/anie.201708228. PMID 28941008.
- Wing R, Drew H, Takano T, Broka C, Tanaka S, Itakura K, Dickerson RE (October 1980). "Crystal structure analysis of a complete turn of B-DNA". Nature. 287 (5784): 755–58. Bibcode:1980Natur.287..755W. doi:10.1038/287755a0. PMID 7432492. S2CID 4315465.
- ^ Pabo CO, Sauer RT (1984). "Protein-DNA recognition". Annual Review of Biochemistry. 53: 293–321. doi:10.1146/annurev.bi.53.070184.001453. PMID 6236744.
- Nikolova EN, Zhou H, Gottardo FL, Alvey HS, Kimsey IJ, Al-Hashimi HM (2013). "A historical account of Hoogsteen base-pairs in duplex DNA". Biopolymers. 99 (12): 955–68. doi:10.1002/bip.22334. PMC 3844552. PMID 23818176.
- Clausen-Schaumann H, Rief M, Tolksdorf C, Gaub HE (April 2000). "Mechanical stability of single DNA molecules". Biophysical Journal. 78 (4): 1997–2007. Bibcode:2000BpJ....78.1997C. doi:10.1016/S0006-3495(00)76747-6. PMC 1300792. PMID 10733978.
- Chalikian TV, Völker J, Plum GE, Breslauer KJ (July 1999). "A more unified picture for the thermodynamics of nucleic acid duplex melting: a characterization by calorimetric and volumetric techniques". Proceedings of the National Academy of Sciences of the United States of America. 96 (14): 7853–58. Bibcode:1999PNAS...96.7853C. doi:10.1073/pnas.96.14.7853. PMC 22151. PMID 10393911.
- deHaseth PL, Helmann JD (June 1995). "Open complex formation by Escherichia coli RNA polymerase: the mechanism of polymerase-induced strand separation of double helical DNA". Molecular Microbiology. 16 (5): 817–24. doi:10.1111/j.1365-2958.1995.tb02309.x. PMID 7476180. S2CID 24479358.
- Isaksson J, Acharya S, Barman J, Cheruku P, Chattopadhyaya J (December 2004). "Single-stranded adenine-rich DNA and RNA retain structural characteristics of their respective double-stranded conformations and show directional differences in stacking pattern" (PDF). Biochemistry. 43 (51): 15996–6010. doi:10.1021/bi048221v. PMID 15609994. Archived (PDF) from the original on 10 June 2007.
- ^ Piovesan A, Pelleri MC, Antonaros F, Strippoli P, Caracausi M, Vitale L (2019). "On the length, weight and GC content of the human genome". BMC Res Notes. 12 (1): 106. doi:10.1186/s13104-019-4137-z. PMC 6391780. PMID 30813969.
- Gregory SG, Barlow KF, McLay KE, Kaul R, Swarbreck D, Dunham A, et al. (May 2006). "The DNA sequence and biological annotation of human chromosome 1". Nature. 441 (7091): 315–21. Bibcode:2006Natur.441..315G. doi:10.1038/nature04727. PMID 16710414.
- Anderson S, Bankier AT, Barrell BG, de Bruijn MH, Coulson AR, Drouin J, et al. (April 1981). "Sequence and organization of the human mitochondrial genome". Nature. 290 (5806): 457–465. Bibcode:1981Natur.290..457A. doi:10.1038/290457a0. PMID 7219534. S2CID 4355527.
- "Untitled". Archived from the original on 13 August 2011. Retrieved 13 June 2012.
- ^ Satoh M, Kuroiwa T (September 1991). "Organization of multiple nucleoids and DNA molecules in mitochondria of a human cell". Experimental Cell Research. 196 (1): 137–140. doi:10.1016/0014-4827(91)90467-9. PMID 1715276.
- Zhang D, Keilty D, Zhang ZF, Chian RC (March 2017). "Mitochondria in oocyte aging: current understanding". Facts, Views & Vision in ObGyn. 9 (1): 29–38. PMC 5506767. PMID 28721182.
- Designation of the two strands of DNA Archived 24 April 2008 at the Wayback Machine JCBN/NC-IUB Newsletter 1989. Retrieved 7 May 2008
- Hüttenhofer A, Schattner P, Polacek N (May 2005). "Non-coding RNAs: hope or hype?". Trends in Genetics. 21 (5): 289–97. doi:10.1016/j.tig.2005.03.007. PMID 15851066.
- Munroe SH (November 2004). "Diversity of antisense regulation in eukaryotes: multiple mechanisms, emerging patterns". Journal of Cellular Biochemistry. 93 (4): 664–71. doi:10.1002/jcb.20252. PMID 15389973. S2CID 23748148.
- Makalowska I, Lin CF, Makalowski W (February 2005). "Overlapping genes in vertebrate genomes". Computational Biology and Chemistry. 29 (1): 1–12. doi:10.1016/j.compbiolchem.2004.12.006. PMID 15680581.
- Johnson ZI, Chisholm SW (November 2004). "Properties of overlapping genes are conserved across microbial genomes". Genome Research. 14 (11): 2268–72. doi:10.1101/gr.2433104. PMC 525685. PMID 15520290.
- Lamb RA, Horvath CM (August 1991). "Diversity of coding strategies in influenza viruses". Trends in Genetics. 7 (8): 261–66. doi:10.1016/0168-9525(91)90326-L. PMC 7173306. PMID 1771674.
- Benham CJ, Mielke SP (2005). "DNA mechanics" (PDF). Annual Review of Biomedical Engineering. 7: 21–53. doi:10.1146/annurev.bioeng.6.062403.132016. PMID 16004565. S2CID 1427671. Archived from the original (PDF) on 1 March 2019.
- ^ Champoux JJ (2001). "DNA topoisomerases: structure, function, and mechanism". Annual Review of Biochemistry. 70: 369–413. doi:10.1146/annurev.biochem.70.1.369. PMID 11395412. S2CID 18144189.
- ^ Wang JC (June 2002). "Cellular roles of DNA topoisomerases: a molecular perspective". Nature Reviews Molecular Cell Biology. 3 (6): 430–40. doi:10.1038/nrm831. PMID 12042765. S2CID 205496065.
- Basu HS, Feuerstein BG, Zarling DA, Shafer RH, Marton LJ (October 1988). "Recognition of Z-RNA and Z-DNA determinants by polyamines in solution: experimental and theoretical studies". Journal of Biomolecular Structure & Dynamics. 6 (2): 299–309. doi:10.1080/07391102.1988.10507714. PMID 2482766.
-
- Franklin RE, Gosling RG (6 March 1953). "The Structure of Sodium Thymonucleate Fibres I. The Influence of Water Content" (PDF). Acta Crystallogr. 6 (8–9): 673–77. Bibcode:1953AcCry...6..673F. doi:10.1107/S0365110X53001939. Archived (PDF) from the original on 9 January 2016.
- Franklin RE, Gosling RG (1953). "The structure of sodium thymonucleate fibres. II. The cylindrically symmetrical Patterson function" (PDF). Acta Crystallogr. 6 (8–9): 678–85. Bibcode:1953AcCry...6..678F. doi:10.1107/S0365110X53001940. Archived (PDF) from the original on 29 June 2017.
- ^ Franklin RE, Gosling RG (April 1953). "Molecular configuration in sodium thymonucleate" (PDF). Nature. 171 (4356): 740–41. Bibcode:1953Natur.171..740F. doi:10.1038/171740a0. PMID 13054694. S2CID 4268222. Archived (PDF) from the original on 3 January 2011.
- ^ Wilkins MH, Stokes AR, Wilson HR (April 1953). "Molecular structure of deoxypentose nucleic acids" (PDF). Nature. 171 (4356): 738–40. Bibcode:1953Natur.171..738W. doi:10.1038/171738a0. PMID 13054693. S2CID 4280080. Archived (PDF) from the original on 13 May 2011.
- Leslie AG, Arnott S, Chandrasekaran R, Ratliff RL (October 1980). "Polymorphism of DNA double helices". Journal of Molecular Biology. 143 (1): 49–72. doi:10.1016/0022-2836(80)90124-2. PMID 7441761.
- Baianu IC (1980). "Structural Order and Partial Disorder in Biological systems". Bull. Math. Biol. 42 (4): 137–41. doi:10.1007/BF02462372. S2CID 189888972.
- Hosemann R, Bagchi RN (1962). Direct analysis of diffraction by matter. Amsterdam – New York: North-Holland Publishers.
- Baianu IC (1978). "X-ray scattering by partially disordered membrane systems" (PDF). Acta Crystallogr A. 34 (5): 751–53. Bibcode:1978AcCrA..34..751B. doi:10.1107/S0567739478001540. Archived from the original (PDF) on 14 March 2020. Retrieved 29 August 2019.
- Wahl MC, Sundaralingam M (1997). "Crystal structures of A-DNA duplexes". Biopolymers. 44 (1): 45–63. doi:10.1002/(SICI)1097-0282(1997)44:1<45::AID-BIP4>3.0.CO;2-#. PMID 9097733.
- Lu XJ, Shakked Z, Olson WK (July 2000). "A-form conformational motifs in ligand-bound DNA structures". Journal of Molecular Biology. 300 (4): 819–40. doi:10.1006/jmbi.2000.3690. PMID 10891271.
- Rothenburg S, Koch-Nolte F, Haag F (December 2001). "DNA methylation and Z-DNA formation as mediators of quantitative differences in the expression of alleles". Immunological Reviews. 184: 286–98. doi:10.1034/j.1600-065x.2001.1840125.x. PMID 12086319. S2CID 20589136.
- Oh DB, Kim YG, Rich A (December 2002). "Z-DNA-binding proteins can act as potent effectors of gene expression in vivo". Proceedings of the National Academy of Sciences of the United States of America. 99 (26): 16666–71. Bibcode:2002PNAS...9916666O. doi:10.1073/pnas.262672699. PMC 139201. PMID 12486233.
- Palmer J (2 December 2010). "Arsenic-loving bacteria may help in hunt for alien life". BBC News. Archived from the original on 3 December 2010. Retrieved 2 December 2010.
- ^ Bortman H (2 December 2010). "Arsenic-Eating Bacteria Opens New Possibilities for Alien Life". Space.com. Archived from the original on 4 December 2010. Retrieved 2 December 2010.
- Katsnelson A (2 December 2010). "Arsenic-eating microbe may redefine chemistry of life". Nature News. doi:10.1038/news.2010.645. Archived from the original on 12 February 2012.
- Cressey D (3 October 2012). "'Arsenic-life' Bacterium Prefers Phosphorus after all". Nature News. doi:10.1038/nature.2012.11520. S2CID 87341731.
- "Structure and packing of human telomeric DNA". ndbserver.rutgers.edu. Retrieved 18 May 2023.
- ^ Greider CW, Blackburn EH (December 1985). "Identification of a specific telomere terminal transferase activity in Tetrahymena extracts". Cell. 43 (2 Pt 1): 405–13. doi:10.1016/0092-8674(85)90170-9. PMID 3907856.
- ^ Nugent CI, Lundblad V (April 1998). "The telomerase reverse transcriptase: components and regulation". Genes & Development. 12 (8): 1073–85. doi:10.1101/gad.12.8.1073. PMID 9553037.
- Wright WE, Tesmer VM, Huffman KE, Levene SD, Shay JW (November 1997). "Normal human chromosomes have long G-rich telomeric overhangs at one end". Genes & Development. 11 (21): 2801–09. doi:10.1101/gad.11.21.2801. PMC 316649. PMID 9353250.
- ^ Burge S, Parkinson GN, Hazel P, Todd AK, Neidle S (2006). "Quadruplex DNA: sequence, topology and structure". Nucleic Acids Research. 34 (19): 5402–15. doi:10.1093/nar/gkl655. PMC 1636468. PMID 17012276.
- Parkinson GN, Lee MP, Neidle S (June 2002). "Crystal structure of parallel quadruplexes from human telomeric DNA". Nature. 417 (6891): 876–80. Bibcode:2002Natur.417..876P. doi:10.1038/nature755. PMID 12050675. S2CID 4422211.
- Griffith JD, Comeau L, Rosenfield S, Stansel RM, Bianchi A, Moss H, de Lange T (May 1999). "Mammalian telomeres end in a large duplex loop". Cell. 97 (4): 503–14. CiteSeerX 10.1.1.335.2649. doi:10.1016/S0092-8674(00)80760-6. PMID 10338214. S2CID 721901.
- Seeman NC (November 2005). "DNA enables nanoscale control of the structure of matter". Quarterly Reviews of Biophysics. 38 (4): 363–71. doi:10.1017/S0033583505004087. PMC 3478329. PMID 16515737.
- Warren M (21 February 2019). "Four new DNA letters double life's alphabet". Nature. 566 (7745): 436. Bibcode:2019Natur.566..436W. doi:10.1038/d41586-019-00650-8. PMID 30809059.
- Hoshika S, Leal NA, Kim MJ, Kim MS, Karalkar NB, Kim HJ, et al. (22 February 2019). "Hachimoji DNA and RNA: A genetic system with eight building blocks (paywall)". Science. 363 (6429): 884–887. Bibcode:2019Sci...363..884H. doi:10.1126/science.aat0971. PMC 6413494. PMID 30792304.
- Burghardt B, Hartmann AK (February 2007). "RNA secondary structure design". Physical Review E. 75 (2): 021920. arXiv:physics/0609135. Bibcode:2007PhRvE..75b1920B. doi:10.1103/PhysRevE.75.021920. PMID 17358380. S2CID 17574854.
- Reusch W. "Nucleic Acids". Michigan State University. Retrieved 30 June 2022.
- "How To Extract DNA From Anything Living". University of Utah. Retrieved 30 June 2022.
- Hu Q, Rosenfeld MG (2012). "Epigenetic regulation of human embryonic stem cells". Frontiers in Genetics. 3: 238. doi:10.3389/fgene.2012.00238. PMC 3488762. PMID 23133442.
- Klose RJ, Bird AP (February 2006). "Genomic DNA methylation: the mark and its mediators". Trends in Biochemical Sciences. 31 (2): 89–97. doi:10.1016/j.tibs.2005.12.008. PMID 16403636.
- Bird A (January 2002). "DNA methylation patterns and epigenetic memory". Genes & Development. 16 (1): 6–21. doi:10.1101/gad.947102. PMID 11782440.
- Walsh CP, Xu GL (2006). "Cytosine methylation and DNA repair". DNA Methylation: Basic Mechanisms. Current Topics in Microbiology and Immunology. Vol. 301. pp. 283–315. doi:10.1007/3-540-31390-7_11. ISBN 3-540-29114-8. PMID 16570853.
- Kriaucionis S, Heintz N (May 2009). "The nuclear DNA base 5-hydroxymethylcytosine is present in Purkinje neurons and the brain". Science. 324 (5929): 929–30. Bibcode:2009Sci...324..929K. doi:10.1126/science.1169786. PMC 3263819. PMID 19372393.
- Ratel D, Ravanat JL, Berger F, Wion D (March 2006). "N6-methyladenine: the other methylated base of DNA". BioEssays. 28 (3): 309–15. doi:10.1002/bies.20342. PMC 2754416. PMID 16479578.
- Gommers-Ampt JH, Van Leeuwen F, de Beer AL, Vliegenthart JF, Dizdaroglu M, Kowalak JA, Crain PF, Borst P (December 1993). "beta-D-glucosyl-hydroxymethyluracil: a novel modified base present in the DNA of the parasitic protozoan T. brucei". Cell. 75 (6): 1129–36. doi:10.1016/0092-8674(93)90322-H. hdl:1874/5219. PMID 8261512. S2CID 24801094.
- Created from PDB 1JDG Archived 22 September 2008 at the Wayback Machine
- Douki T, Reynaud-Angelin A, Cadet J, Sage E (August 2003). "Bipyrimidine photoproducts rather than oxidative lesions are the main type of DNA damage involved in the genotoxic effect of solar UVA radiation". Biochemistry. 42 (30): 9221–26. doi:10.1021/bi034593c. PMID 12885257.
- Cadet J, Delatour T, Douki T, Gasparutto D, Pouget JP, Ravanat JL, Sauvaigo S (March 1999). "Hydroxyl radicals and DNA base damage". Mutation Research. 424 (1–2): 9–21. Bibcode:1999MRFMM.424....9C. doi:10.1016/S0027-5107(99)00004-4. PMID 10064846.
- Beckman KB, Ames BN (August 1997). "Oxidative decay of DNA". The Journal of Biological Chemistry. 272 (32): 19633–36. doi:10.1074/jbc.272.32.19633. PMID 9289489.
- Valerie K, Povirk LF (September 2003). "Regulation and mechanisms of mammalian double-strand break repair". Oncogene. 22 (37): 5792–812. doi:10.1038/sj.onc.1206679. PMID 12947387.
- Johnson G (28 December 2010). "Unearthing Prehistoric Tumors, and Debate". The New York Times. Archived from the original on 24 June 2017.
If we lived long enough, sooner or later we all would get cancer.
- Alberts B, Johnson A, Lewis J, et al. (2002). "The Preventable Causes of Cancer". Molecular biology of the cell (4th ed.). New York: Garland Science. ISBN 0-8153-4072-9. Archived from the original on 2 January 2016.
A certain irreducible background incidence of cancer is to be expected regardless of circumstances: mutations can never be absolutely avoided, because they are an inescapable consequence of fundamental limitations on the accuracy of DNA replication, as discussed in Chapter 5. If a human could live long enough, it is inevitable that at least one of his or her cells would eventually accumulate a set of mutations sufficient for cancer to develop.
- Bernstein H, Payne CM, Bernstein C, Garewal H, Dvorak K (2008). "Cancer and aging as consequences of un-repaired DNA damage". In Kimura H, Suzuki A (eds.). New Research on DNA Damage. New York: Nova Science Publishers. pp. 1–47. ISBN 978-1-60456-581-2. Archived from the original on 25 October 2014.
- Hoeijmakers JH (October 2009). "DNA damage, aging, and cancer". The New England Journal of Medicine. 361 (15): 1475–85. doi:10.1056/NEJMra0804615. PMID 19812404.
- Freitas AA, de Magalhães JP (2011). "A review and appraisal of the DNA damage theory of ageing". Mutation Research. 728 (1–2): 12–22. Bibcode:2011MRRMR.728...12F. doi:10.1016/j.mrrev.2011.05.001. PMID 21600302.
- Ferguson LR, Denny WA (September 1991). "The genetic toxicology of acridines". Mutation Research. 258 (2): 123–60. doi:10.1016/0165-1110(91)90006-H. PMID 1881402.
- Stephens TD, Bunde CJ, Fillmore BJ (June 2000). "Mechanism of action in thalidomide teratogenesis". Biochemical Pharmacology. 59 (12): 1489–99. doi:10.1016/S0006-2952(99)00388-3. PMID 10799645.
- Jeffrey AM (1985). "DNA modification by chemical carcinogens". Pharmacology & Therapeutics. 28 (2): 237–72. doi:10.1016/0163-7258(85)90013-0. PMID 3936066.
- Braña MF, Cacho M, Gradillas A, de Pascual-Teresa B, Ramos A (November 2001). "Intercalators as anticancer drugs". Current Pharmaceutical Design. 7 (17): 1745–80. doi:10.2174/1381612013397113. PMID 11562309.
- Venter JC, Adams MD, Myers EW, Li PW, Mural RJ, Sutton GG, et al. (February 2001). "The sequence of the human genome". Science. 291 (5507): 1304–51. Bibcode:2001Sci...291.1304V. doi:10.1126/science.1058040. PMID 11181995.
- Thanbichler M, Wang SC, Shapiro L (October 2005). "The bacterial nucleoid: a highly organized and dynamic structure". Journal of Cellular Biochemistry. 96 (3): 506–21. doi:10.1002/jcb.20519. PMID 15988757.
- Wolfsberg TG, McEntyre J, Schuler GD (February 2001). "Guide to the draft human genome". Nature. 409 (6822): 824–26. Bibcode:2001Natur.409..824W. doi:10.1038/35057000. PMID 11236998.
- Gregory TR (January 2005). "The C-value enigma in plants and animals: a review of parallels and an appeal for partnership". Annals of Botany. 95 (1): 133–46. doi:10.1093/aob/mci009. PMC 4246714. PMID 15596463.
- Birney E, Stamatoyannopoulos JA, Dutta A, Guigó R, Gingeras TR, Margulies EH, et al. (June 2007). "Identification and analysis of functional elements in 1% of the human genome by the ENCODE pilot project". Nature. 447 (7146): 799–816. Bibcode:2007Natur.447..799B. doi:10.1038/nature05874. PMC 2212820. PMID 17571346.
- Yin YW, Steitz TA. "RCSB PDB – 1MSW: Structural basis for the transition from initiation to elongation transcription in T7 RNA polymerase". www.rcsb.org. Retrieved 27 March 2023.
- Pidoux AL, Allshire RC (March 2005). "The role of heterochromatin in centromere function". Philosophical Transactions of the Royal Society of London. Series B, Biological Sciences. 360 (1455): 569–79. doi:10.1098/rstb.2004.1611. PMC 1569473. PMID 15905142.
- Harrison PM, Hegyi H, Balasubramanian S, Luscombe NM, Bertone P, Echols N, Johnson T, Gerstein M (February 2002). "Molecular fossils in the human genome: identification and analysis of the pseudogenes in chromosomes 21 and 22". Genome Research. 12 (2): 272–80. doi:10.1101/gr.207102. PMC 155275. PMID 11827946.
- Harrison PM, Gerstein M (May 2002). "Studying genomes through the aeons: protein families, pseudogenes and proteome evolution". Journal of Molecular Biology. 318 (5): 1155–74. doi:10.1016/S0022-2836(02)00109-2. PMID 12083509.
- Albà M (2001). "Replicative DNA polymerases". Genome Biology. 2 (1): REVIEWS3002. doi:10.1186/gb-2001-2-1-reviews3002. PMC 150442. PMID 11178285.
- Tani K, Nasu M (2010). "Roles of Extracellular DNA in Bacterial Ecosystems". In Kikuchi Y, Rykova EY (eds.). Extracellular Nucleic Acids. Springer. pp. 25–38. ISBN 978-3-642-12616-1.
- Vlassov VV, Laktionov PP, Rykova EY (July 2007). "Extracellular nucleic acids". BioEssays. 29 (7): 654–67. doi:10.1002/bies.20604. PMID 17563084. S2CID 32463239.
- Finkel SE, Kolter R (November 2001). "DNA as a nutrient: novel role for bacterial competence gene homologs". Journal of Bacteriology. 183 (21): 6288–93. doi:10.1128/JB.183.21.6288-6293.2001. PMC 100116. PMID 11591672.
- Mulcahy H, Charron-Mazenod L, Lewenza S (November 2008). "Extracellular DNA chelates cations and induces antibiotic resistance in Pseudomonas aeruginosa biofilms". PLOS Pathogens. 4 (11): e1000213. doi:10.1371/journal.ppat.1000213. PMC 2581603. PMID 19023416.
- Berne C, Kysela DT, Brun YV (August 2010). "A bacterial extracellular DNA inhibits settling of motile progeny cells within a biofilm". Molecular Microbiology. 77 (4): 815–29. doi:10.1111/j.1365-2958.2010.07267.x. PMC 2962764. PMID 20598083.
- Whitchurch CB, Tolker-Nielsen T, Ragas PC, Mattick JS (February 2002). "Extracellular DNA required for bacterial biofilm formation". Science. 295 (5559): 1487. doi:10.1126/science.295.5559.1487. PMID 11859186.
- Hu W, Li L, Sharma S, Wang J, McHardy I, Lux R, Yang Z, He X, Gimzewski JK, Li Y, Shi W (2012). "DNA builds and strengthens the extracellular matrix in Myxococcus xanthus biofilms by interacting with exopolysaccharides". PLOS ONE. 7 (12): e51905. Bibcode:2012PLoSO...751905H. doi:10.1371/journal.pone.0051905. PMC 3530553. PMID 23300576.
- Hui L, Bianchi DW (February 2013). "Recent advances in the prenatal interrogation of the human fetal genome". Trends in Genetics. 29 (2): 84–91. doi:10.1016/j.tig.2012.10.013. PMC 4378900. PMID 23158400.
- Foote AD, Thomsen PF, Sveegaard S, Wahlberg M, Kielgast J, Kyhn LA, et al. (2012). "Investigating the potential use of environmental DNA (eDNA) for genetic monitoring of marine mammals". PLOS ONE. 7 (8): e41781. Bibcode:2012PLoSO...741781F. doi:10.1371/journal.pone.0041781. PMC 3430683. PMID 22952587.
- "Researchers Detect Land Animals Using DNA in Nearby Water Bodies".
- Sandman K, Pereira SL, Reeve JN (December 1998). "Diversity of prokaryotic chromosomal proteins and the origin of the nucleosome". Cellular and Molecular Life Sciences. 54 (12): 1350–64. doi:10.1007/s000180050259. PMC 11147202. PMID 9893710. S2CID 21101836.
- Dame RT (May 2005). "The role of nucleoid-associated proteins in the organization and compaction of bacterial chromatin". Molecular Microbiology. 56 (4): 858–70. doi:10.1111/j.1365-2958.2005.04598.x. PMID 15853876. S2CID 26965112.
- Luger K, Mäder AW, Richmond RK, Sargent DF, Richmond TJ (September 1997). "Crystal structure of the nucleosome core particle at 2.8 A resolution". Nature. 389 (6648): 251–60. Bibcode:1997Natur.389..251L. doi:10.1038/38444. PMID 9305837. S2CID 4328827.
- Jenuwein T, Allis CD (August 2001). "Translating the histone code" (PDF). Science. 293 (5532): 1074–80. doi:10.1126/science.1063127. PMID 11498575. S2CID 1883924. Archived (PDF) from the original on 8 August 2017.
- Ito T (2003). "Nucleosome Assembly and Remodeling". Protein Complexes that Modify Chromatin. Current Topics in Microbiology and Immunology. Vol. 274. pp. 1–22. doi:10.1007/978-3-642-55747-7_1. ISBN 978-3-540-44208-0. PMID 12596902.
- Thomas JO (August 2001). "HMG1 and 2: architectural DNA-binding proteins". Biochemical Society Transactions. 29 (Pt 4): 395–401. doi:10.1042/BST0290395. PMID 11497996.
- Grosschedl R, Giese K, Pagel J (March 1994). "HMG domain proteins: architectural elements in the assembly of nucleoprotein structures". Trends in Genetics. 10 (3): 94–100. doi:10.1016/0168-9525(94)90232-1. PMID 8178371.
- Iftode C, Daniely Y, Borowiec JA (1999). "Replication protein A (RPA): the eukaryotic SSB". Critical Reviews in Biochemistry and Molecular Biology. 34 (3): 141–80. doi:10.1080/10409239991209255. PMID 10473346.
- Beamer LJ, Pabo CO. "RCSB PDB – 1LMB: Refined 1.8 Å crystal structure of the lambda repressor-operator complex". www.rcsb.org. Retrieved 27 March 2023.
- Myers LC, Kornberg RD (2000). "Mediator of transcriptional regulation". Annual Review of Biochemistry. 69: 729–49. doi:10.1146/annurev.biochem.69.1.729. PMID 10966474.
- Spiegelman BM, Heinrich R (October 2004). "Biological control through regulated transcriptional coactivators". Cell. 119 (2): 157–67. doi:10.1016/j.cell.2004.09.037. PMID 15479634.
- Li Z, Van Calcar S, Qu C, Cavenee WK, Zhang MQ, Ren B (July 2003). "A global transcriptional regulatory role for c-Myc in Burkitt's lymphoma cells". Proceedings of the National Academy of Sciences of the United States of America. 100 (14): 8164–69. Bibcode:2003PNAS..100.8164L. doi:10.1073/pnas.1332764100. PMC 166200. PMID 12808131.
- Kostrewa D, Winkler FK. "RCSB PDB – 1RVA: Mg2+ binding to the active site of EcoRV endonuclease: a crystallographic study of complexes with substrate and product DNA at 2 Å resolution". www.rcsb.org. Retrieved 27 March 2023.
- Bickle TA, Krüger DH (June 1993). "Biology of DNA restriction". Microbiological Reviews. 57 (2): 434–50. doi:10.1128/MMBR.57.2.434-450.1993. PMC 372918. PMID 8336674.
- ^ Doherty AJ, Suh SW (November 2000). "Structural and mechanistic conservation in DNA ligases". Nucleic Acids Research. 28 (21): 4051–58. doi:10.1093/nar/28.21.4051. PMC 113121. PMID 11058099.
- Schoeffler AJ, Berger JM (December 2005). "Recent advances in understanding structure-function relationships in the type II topoisomerase mechanism". Biochemical Society Transactions. 33 (Pt 6): 1465–70. doi:10.1042/BST20051465. PMID 16246147.
- Tuteja N, Tuteja R (May 2004). "Unraveling DNA helicases. Motif, structure, mechanism and function" (PDF). European Journal of Biochemistry. 271 (10): 1849–63. doi:10.1111/j.1432-1033.2004.04094.x. PMID 15128295.
- Joyce CM, Steitz TA (November 1995). "Polymerase structures and function: variations on a theme?". Journal of Bacteriology. 177 (22): 6321–29. doi:10.1128/jb.177.22.6321-6329.1995. PMC 177480. PMID 7592405.
- Hubscher U, Maga G, Spadari S (2002). "Eukaryotic DNA polymerases" (PDF). Annual Review of Biochemistry. 71: 133–63. doi:10.1146/annurev.biochem.71.090501.150041. PMID 12045093. S2CID 26171993. Archived from the original (PDF) on 26 January 2021.
- Johnson A, O'Donnell M (2005). "Cellular DNA replicases: components and dynamics at the replication fork". Annual Review of Biochemistry. 74: 283–315. doi:10.1146/annurev.biochem.73.011303.073859. PMID 15952889.
- ^ Tarrago-Litvak L, Andréola ML, Nevinsky GA, Sarih-Cottin L, Litvak S (May 1994). "The reverse transcriptase of HIV-1: from enzymology to therapeutic intervention". FASEB Journal. 8 (8): 497–503. doi:10.1096/fasebj.8.8.7514143. PMID 7514143. S2CID 39614573.
- Martinez E (December 2002). "Multi-protein complexes in eukaryotic gene transcription". Plant Molecular Biology. 50 (6): 925–47. doi:10.1023/A:1021258713850. PMID 12516863. S2CID 24946189.
- Thorpe JH, Gale BC, Teixeira SC, Cardin CJ. "RCSB PDB – 1M6G: Structural Characterisation of the Holliday Junction TCGGTACCGA". www.rcsb.org. Retrieved 27 March 2023.
- Cremer T, Cremer C (April 2001). "Chromosome territories, nuclear architecture and gene regulation in mammalian cells". Nature Reviews Genetics. 2 (4): 292–301. doi:10.1038/35066075. PMID 11283701. S2CID 8547149.
- Pál C, Papp B, Lercher MJ (May 2006). "An integrated view of protein evolution". Nature Reviews Genetics. 7 (5): 337–48. doi:10.1038/nrg1838. PMID 16619049. S2CID 23225873.
- O'Driscoll M, Jeggo PA (January 2006). "The role of double-strand break repair – insights from human genetics". Nature Reviews Genetics. 7 (1): 45–54. doi:10.1038/nrg1746. PMID 16369571. S2CID 7779574.
- Vispé S, Defais M (October 1997). "Mammalian Rad51 protein: a RecA homologue with pleiotropic functions". Biochimie. 79 (9–10): 587–92. doi:10.1016/S0300-9084(97)82007-X. PMID 9466696.
- Neale MJ, Keeney S (July 2006). "Clarifying the mechanics of DNA strand exchange in meiotic recombination". Nature. 442 (7099): 153–58. Bibcode:2006Natur.442..153N. doi:10.1038/nature04885. PMC 5607947. PMID 16838012.
- Dickman MJ, Ingleston SM, Sedelnikova SE, Rafferty JB, Lloyd RG, Grasby JA, Hornby DP (November 2002). "The RuvABC resolvasome". European Journal of Biochemistry. 269 (22): 5492–501. doi:10.1046/j.1432-1033.2002.03250.x. PMID 12423347. S2CID 39505263.
- Joyce GF (July 2002). "The antiquity of RNA-based evolution". Nature. 418 (6894): 214–21. Bibcode:2002Natur.418..214J. doi:10.1038/418214a. PMID 12110897. S2CID 4331004.
- Orgel LE (2004). "Prebiotic chemistry and the origin of the RNA world". Critical Reviews in Biochemistry and Molecular Biology. 39 (2): 99–123. CiteSeerX 10.1.1.537.7679. doi:10.1080/10409230490460765. PMID 15217990. S2CID 4939632.
- Davenport RJ (May 2001). "Ribozymes. Making copies in the RNA world". Science. 292 (5520): 1278a–1278. doi:10.1126/science.292.5520.1278a. PMID 11360970. S2CID 85976762.
- Szathmáry E (April 1992). "What is the optimum size for the genetic alphabet?". Proceedings of the National Academy of Sciences of the United States of America. 89 (7): 2614–18. Bibcode:1992PNAS...89.2614S. doi:10.1073/pnas.89.7.2614. PMC 48712. PMID 1372984.
- Lindahl T (April 1993). "Instability and decay of the primary structure of DNA". Nature. 362 (6422): 709–15. Bibcode:1993Natur.362..709L. doi:10.1038/362709a0. PMID 8469282. S2CID 4283694.
- Vreeland RH, Rosenzweig WD, Powers DW (October 2000). "Isolation of a 250 million-year-old halotolerant bacterium from a primary salt crystal". Nature. 407 (6806): 897–900. Bibcode:2000Natur.407..897V. doi:10.1038/35038060. PMID 11057666. S2CID 9879073.
- Hebsgaard MB, Phillips MJ, Willerslev E (May 2005). "Geologically ancient DNA: fact or artefact?". Trends in Microbiology. 13 (5): 212–20. doi:10.1016/j.tim.2005.03.010. PMID 15866038.
- Nickle DC, Learn GH, Rain MW, Mullins JI, Mittler JE (January 2002). "Curiously modern DNA for a "250 million-year-old" bacterium". Journal of Molecular Evolution. 54 (1): 134–37. Bibcode:2002JMolE..54..134N. doi:10.1007/s00239-001-0025-x. PMID 11734907. S2CID 24740859.
- Callahan MP, Smith KE, Cleaves HJ, Ruzicka J, Stern JC, Glavin DP, House CH, Dworkin JP (August 2011). "Carbonaceous meteorites contain a wide range of extraterrestrial nucleobases". Proceedings of the National Academy of Sciences of the United States of America. 108 (34): 13995–98. Bibcode:2011PNAS..10813995C. doi:10.1073/pnas.1106493108. PMC 3161613. PMID 21836052.
- Steigerwald J (8 August 2011). "NASA Researchers: DNA Building Blocks Can Be Made in Space". NASA. Archived from the original on 23 June 2015. Retrieved 10 August 2011.
- ScienceDaily Staff (9 August 2011). "DNA Building Blocks Can Be Made in Space, NASA Evidence Suggests". ScienceDaily. Archived from the original on 5 September 2011. Retrieved 9 August 2011.
- Marlaire R (3 March 2015). "NASA Ames Reproduces the Building Blocks of Life in Laboratory". NASA. Archived from the original on 5 March 2015. Retrieved 5 March 2015.
- Hunt K (17 February 2021). "World's oldest DNA sequenced from a mammoth that lived more than a million years ago". CNN News. Retrieved 17 February 2021.
- Callaway E (17 February 2021). "Million-year-old mammoth genomes shatter record for oldest ancient DNA – Permafrost-preserved teeth, up to 1.6 million years old, identify a new kind of mammoth in Siberia". Nature. 590 (7847): 537–538. Bibcode:2021Natur.590..537C. doi:10.1038/d41586-021-00436-x. ISSN 0028-0836. PMID 33597786.
- Goff SP, Berg P (December 1976). "Construction of hybrid viruses containing SV40 and lambda phage DNA segments and their propagation in cultured monkey cells". Cell. 9 (4 PT 2): 695–705. doi:10.1016/0092-8674(76)90133-1. PMID 189942. S2CID 41788896.
- Houdebine LM (2007). "Transgenic animal models in biomedical research". Target Discovery and Validation Reviews and Protocols. Methods in Molecular Biology. Vol. 360. pp. 163–202. doi:10.1385/1-59745-165-7:163. ISBN 978-1-59745-165-9. PMID 17172731.
- Daniell H, Dhingra A (April 2002). "Multigene engineering: dawn of an exciting new era in biotechnology". Current Opinion in Biotechnology. 13 (2): 136–41. doi:10.1016/S0958-1669(02)00297-5. PMC 3481857. PMID 11950565.
- Job D (November 2002). "Plant biotechnology in agriculture". Biochimie. 84 (11): 1105–10. doi:10.1016/S0300-9084(02)00013-5. PMID 12595138.
- Curtis C, Hereward J (29 August 2017). "From the crime scene to the courtroom: the journey of a DNA sample". The Conversation. Archived from the original on 22 October 2017. Retrieved 22 October 2017.
- Collins A, Morton NE (June 1994). "Likelihood ratios for DNA identification". Proceedings of the National Academy of Sciences of the United States of America. 91 (13): 6007–11. Bibcode:1994PNAS...91.6007C. doi:10.1073/pnas.91.13.6007. PMC 44126. PMID 8016106.
- Weir BS, Triggs CM, Starling L, Stowell LI, Walsh KA, Buckleton J (March 1997). "Interpreting DNA mixtures". Journal of Forensic Sciences. 42 (2): 213–22. doi:10.1520/JFS14100J. PMID 9068179. S2CID 14511630.
- Jeffreys AJ, Wilson V, Thein SL (1985). "Individual-specific 'fingerprints' of human DNA". Nature. 316 (6023): 76–79. Bibcode:1985Natur.316...76J. doi:10.1038/316076a0. PMID 2989708. S2CID 4229883.
- "Colin Pitchfork". 14 December 2006. Archived from the original on 14 December 2006. Retrieved 27 March 2023.
- "DNA Identification in Mass Fatality Incidents". National Institute of Justice. September 2006. Archived from the original on 12 November 2006.
- Pollack A (19 June 2012). "Before Birth, Dad's ID". The New York Times. ISSN 0362-4331. Archived from the original on 24 June 2017. Retrieved 27 March 2023.
- ^ Breaker RR, Joyce GF (December 1994). "A DNA enzyme that cleaves RNA". Chemistry & Biology. 1 (4): 223–29. doi:10.1016/1074-5521(94)90014-0. PMID 9383394.
- Chandra M, Sachdeva A, Silverman SK (October 2009). "DNA-catalyzed sequence-specific hydrolysis of DNA". Nature Chemical Biology. 5 (10): 718–20. doi:10.1038/nchembio.201. PMC 2746877. PMID 19684594.
- Carmi N, Shultz LA, Breaker RR (December 1996). "In vitro selection of self-cleaving DNAs". Chemistry & Biology. 3 (12): 1039–46. doi:10.1016/S1074-5521(96)90170-2. PMID 9000012.
- Torabi SF, Wu P, McGhee CE, Chen L, Hwang K, Zheng N, Cheng J, Lu Y (May 2015). "In vitro selection of a sodium-specific DNAzyme and its application in intracellular sensing". Proceedings of the National Academy of Sciences of the United States of America. 112 (19): 5903–08. Bibcode:2015PNAS..112.5903T. doi:10.1073/pnas.1420361112. PMC 4434688. PMID 25918425.
- Baldi P, Brunak S (2001). Bioinformatics: The Machine Learning Approach. MIT Press. ISBN 978-0-262-02506-5. OCLC 45951728.
- Gusfield D (15 January 1997). Algorithms on Strings, Trees, and Sequences: Computer Science and Computational Biology. Cambridge University Press. ISBN 978-0-521-58519-4.
- Sjölander K (January 2004). "Phylogenomic inference of protein molecular function: advances and challenges". Bioinformatics. 20 (2): 170–79. CiteSeerX 10.1.1.412.943. doi:10.1093/bioinformatics/bth021. PMID 14734307.
- Mount DM (2004). Bioinformatics: Sequence and Genome Analysis (2nd ed.). Cold Spring Harbor, NY: Cold Spring Harbor Laboratory Press. ISBN 0-87969-712-1. OCLC 55106399.
- Strong M (March 2004). "Protein nanomachines". PLOS Biology. 2 (3): E73. doi:10.1371/journal.pbio.0020073. PMC 368168. PMID 15024422. S2CID 13222080.
- Rothemund PW (March 2006). "Folding DNA to create nanoscale shapes and patterns" (PDF). Nature. 440 (7082): 297–302. Bibcode:2006Natur.440..297R. doi:10.1038/nature04586. PMID 16541064. S2CID 4316391.
- Andersen ES, Dong M, Nielsen MM, Jahn K, Subramani R, Mamdouh W, Golas MM, Sander B, Stark H, Oliveira CL, Pedersen JS, Birkedal V, Besenbacher F, Gothelf KV, Kjems J (May 2009). "Self-assembly of a nanoscale DNA box with a controllable lid". Nature. 459 (7243): 73–76. Bibcode:2009Natur.459...73A. doi:10.1038/nature07971. hdl:11858/00-001M-0000-0010-9362-B. PMID 19424153. S2CID 4430815.
- Ishitsuka Y, Ha T (May 2009). "DNA nanotechnology: a nanomachine goes live". Nature Nanotechnology. 4 (5): 281–82. Bibcode:2009NatNa...4..281I. doi:10.1038/nnano.2009.101. PMID 19421208.
- Aldaye FA, Palmer AL, Sleiman HF (September 2008). "Assembling materials with DNA as the guide". Science. 321 (5897): 1795–99. Bibcode:2008Sci...321.1795A. doi:10.1126/science.1154533. PMID 18818351. S2CID 2755388.
- Dunn MR, Jimenez RM, Chaput JC (2017). "Analysis of aptamer discovery and technology". Nature Reviews Chemistry. 1 (10). doi:10.1038/s41570-017-0076. Retrieved 30 June 2022.
- Wray GA (2002). "Dating branches on the tree of life using DNA". Genome Biology. 3 (1): REVIEWS0001. doi:10.1186/gb-2001-3-1-reviews0001. PMC 150454. PMID 11806830.
- Panda D, Molla KA, Baig MJ, Swain A, Behera D, Dash M (May 2018). "DNA as a digital information storage device: hope or hype?". 3 Biotech. 8 (5): 239. doi:10.1007/s13205-018-1246-7. PMC 5935598. PMID 29744271.
- Akram F, Haq IU, Ali H, Laghari AT (October 2018). "Trends to store digital data in DNA: an overview". Molecular Biology Reports. 45 (5): 1479–1490. doi:10.1007/s11033-018-4280-y. PMID 30073589. S2CID 51905843.
- Miescher F (1871). "Ueber die chemische Zusammensetzung der Eiterzellen" [On the chemical composition of pus cells]. Medicinisch-chemische Untersuchungen (in German). 4: 441–60.
Ich habe mich daher später mit meinen Versuchen an die ganzen Kerne gehalten, die Trennung der Körper, die ich einstweilen ohne weiteres Präjudiz als lösliches und unlösliches Nuclein bezeichnen will, einem günstigeren Material überlassend. (Therefore, in my experiments I subsequently limited myself to the whole nucleus, leaving to a more favorable material the separation of the substances, that for the present, without further prejudice, I will designate as soluble and insoluble nuclear material ("Nuclein"))
- Dahm R (January 2008). "Discovering DNA: Friedrich Miescher and the early years of nucleic acid research". Human Genetics. 122 (6): 565–81. doi:10.1007/s00439-007-0433-0. PMID 17901982. S2CID 915930.
- See:
- Kossel A (1879). "Ueber Nucleïn der Hefe" [On nuclein in yeast]. Zeitschrift für physiologische Chemie (in German). 3: 284–91.
- Kossel A (1880). "Ueber Nucleïn der Hefe II" [On nuclein in yeast, Part 2]. Zeitschrift für physiologische Chemie (in German). 4: 290–95.
- Kossel A (1881). "Ueber die Verbreitung des Hypoxanthins im Thier- und Pflanzenreich" [On the distribution of hypoxanthins in the animal and plant kingdoms]. Zeitschrift für physiologische Chemie (in German). 5: 267–71.
- Kossel A (1881). Untersuchungen über die Nucleine und ihre Spaltungsprodukte [Investigations into nuclein and its cleavage products] (in German). Strassburg, Germany: K.J. Trübner. p. 19.
- Kossel A (1882). "Ueber Xanthin und Hypoxanthin" [On xanthin and hypoxanthin]. Zeitschrift für physiologische Chemie. 6: 422–31.
- Albrect Kossel (1883) "Zur Chemie des Zellkerns" Archived 17 November 2017 at the Wayback Machine (On the chemistry of the cell nucleus), Zeitschrift für physiologische Chemie, 7: 7–22.
- Kossel A (1886). "Weitere Beiträge zur Chemie des Zellkerns" [Further contributions to the chemistry of the cell nucleus]. Zeitschrift für Physiologische Chemie (in German). 10: 248–64.
On p. 264, Kossel remarked presciently: Der Erforschung der quantitativen Verhältnisse der vier stickstoffreichen Basen, der Abhängigkeit ihrer Menge von den physiologischen Zuständen der Zelle, verspricht wichtige Aufschlüsse über die elementaren physiologisch-chemischen Vorgänge. (The study of the quantitative relations of the four nitrogenous bases— of the dependence of their quantity on the physiological states of the cell—promises important insights into the fundamental physiological-chemical processes.)
- Jones ME (September 1953). "Albrecht Kossel, a biographical sketch". The Yale Journal of Biology and Medicine. 26 (1): 80–97. PMC 2599350. PMID 13103145.
- Levene PA, Jacobs WA (1909). "Über Inosinsäure". Berichte der Deutschen Chemischen Gesellschaft (in German). 42: 1198–203. doi:10.1002/cber.190904201196.
- Levene PA, Jacobs WA (1909). "Über die Hefe-Nucleinsäure". Berichte der Deutschen Chemischen Gesellschaft (in German). 42 (2): 2474–78. doi:10.1002/cber.190904202148.
- Levene P (1919). "The structure of yeast nucleic acid". J Biol Chem. 40 (2): 415–24. doi:10.1016/S0021-9258(18)87254-4.
- Cohen JS, Portugal FH (1974). "The search for the chemical structure of DNA" (PDF). Connecticut Medicine. 38 (10): 551–52, 554–57. PMID 4609088.
- Koltsov proposed that a cell's genetic information was encoded in a long chain of amino acids. See:
- Koltsov HK (12 December 1927). Физико-химические основы морфологии [The physical-chemical basis of morphology] (Speech). 3rd All-Union Meeting of Zoologist, Anatomists, and Histologists (in Russian). Leningrad, U.S.S.R.
- Reprinted in: Koltsov HK (1928). "Физико-химические основы морфологии" [The physical-chemical basis of morphology]. Успехи экспериментальной биологии (Advances in Experimental Biology) series B (in Russian). 7 (1): ?.
- Reprinted in German as: Koltzoff NK (1928). "Physikalisch-chemische Grundlagen der Morphologie" [The physical-chemical basis of morphology]. Biologisches Zentralblatt (in German). 48 (6): 345–69.
- In 1934, Koltsov contended that the proteins that contain a cell's genetic information replicate. See: Koltzoff N (October 1934). "The structure of the chromosomes in the salivary glands of Drosophila". Science. 80 (2075): 312–13. Bibcode:1934Sci....80..312K. doi:10.1126/science.80.2075.312. PMID 17769043.
From page 313: "I think that the size of the chromosomes in the salivary glands is determined through the multiplication of genonemes. By this term I designate the axial thread of the chromosome, in which the geneticists locate the linear combination of genes; … In the normal chromosome there is usually only one genoneme; before cell-division this genoneme has become divided into two strands."
- Soyfer VN (September 2001). "The consequences of political dictatorship for Russian science". Nature Reviews Genetics. 2 (9): 723–29. doi:10.1038/35088598. PMID 11533721. S2CID 46277758.
- Griffith F (January 1928). "The Significance of Pneumococcal Types". The Journal of Hygiene. 27 (2): 113–59. doi:10.1017/S0022172400031879. PMC 2167760. PMID 20474956.
- Lorenz MG, Wackernagel W (September 1994). "Bacterial gene transfer by natural genetic transformation in the environment". Microbiological Reviews. 58 (3): 563–602. doi:10.1128/MMBR.58.3.563-602.1994. PMC 372978. PMID 7968924.
- Brachet J (1933). "Recherches sur la synthese de l'acide thymonucleique pendant le developpement de l'oeuf d'Oursin". Archives de Biologie (in Italian). 44: 519–76.
- Burian R (1994). "Jean Brachet's Cytochemical Embryology: Connections with the Renovation of Biology in France?" (PDF). In Debru C, Gayon J, Picard JF (eds.). Les sciences biologiques et médicales en France 1920–1950. Cahiers pour I'histoire de la recherche. Vol. 2. Paris: CNRS Editions. pp. 207–20.
- See:
- Astbury WT, Bell FO (1938). "Some recent developments in the X-ray study of proteins and related structures" (PDF). Cold Spring Harbor Symposia on Quantitative Biology. 6: 109–21. doi:10.1101/sqb.1938.006.01.013. Archived from the original (PDF) on 14 July 2014.
- Astbury WT (1947). "X-ray studies of nucleic acids". Symposia of the Society for Experimental Biology (1): 66–76. PMID 20257017. Archived from the original on 5 July 2014.
- Avery OT, Macleod CM, McCarty M (February 1944). "Studies on the Chemical Nature of the Substance Inducing Transformation of Pneumococcal Types: Induction of Transformation by a Desoxyribonucleic Acid Fraction Isolated from Pneumococcus Type III". The Journal of Experimental Medicine. 79 (2): 137–158. doi:10.1084/jem.79.2.137. PMC 2135445. PMID 19871359.
- Chargaff E (June 1950). "Chemical specificity of nucleic acids and mechanism of their enzymatic degradation". Experientia. 6 (6): 201–209. doi:10.1007/BF02173653. PMID 15421335. S2CID 2522535.
- Kresge N, Simoni RD, Hill RL (June 2005). "Chargaff's Rules: the Work of Erwin Chargaff". Journal of Biological Chemistry. 280 (24): 172–174. doi:10.1016/S0021-9258(20)61522-8.
- Hershey AD, Chase M (May 1952). "Independent functions of viral protein and nucleic acid in growth of bacteriophage". The Journal of General Physiology. 36 (1): 39–56. doi:10.1085/jgp.36.1.39. PMC 2147348. PMID 12981234.
- "Pictures and Illustrations: Crystallographic photo of Sodium Thymonucleate, Type B. "Photo 51." May 1952". scarc.library.oregonstate.edu. Retrieved 18 May 2023.
- Schwartz J (2008). In pursuit of the gene: from Darwin to DNA. Cambridge, Mass.: Harvard University Press. ISBN 978-0-674-02670-4.
- Pauling L, Corey RB (February 1953). "A Proposed Structure For The Nucleic Acids". Proceedings of the National Academy of Sciences of the United States of America. 39 (2): 84–97. Bibcode:1953PNAS...39...84P. doi:10.1073/pnas.39.2.84. PMC 1063734. PMID 16578429.
- Regis E (2009). What Is Life?: investigating the nature of life in the age of synthetic biology. Oxford: Oxford University Press. p. 52. ISBN 978-0-19-538341-6.
- "Double Helix of DNA: 50 Years". Nature Archives. Archived from the original on 5 April 2015.
- "Original X-ray diffraction image". Oregon State Library. Archived from the original on 30 January 2009. Retrieved 6 February 2011.
- Burakoff M (25 April 2023). "Rosalind Franklin's role in DNA discovery gets a new twist". AP News. Retrieved 25 April 2023.
- Anthes E (25 April 2023). "Untangling Rosalind Franklin's Role in DNA Discovery, 70 Years On – Historians have long debated the role that Dr. Franklin played in identifying the double helix. A new opinion essay argues that she was an "equal contributor."". The New York Times. Archived from the original on 25 April 2023. Retrieved 26 April 2023.
- Cobb M, Comfort N (25 April 2023). "What Rosalind Franklin truly contributed to the discovery of DNA's structure – Franklin was no victim in how the DNA double helix was solved. An overlooked letter and an unpublished news article, both written in 1953, reveal that she was an equal player". Nature. 616 (7958): 657–660. Bibcode:2023Natur.616..657C. doi:10.1038/d41586-023-01313-5. PMID 37100935. S2CID 258314143.
- "The Nobel Prize in Physiology or Medicine 1962". Nobelprize.org.
- Maddox B (January 2003). "The double helix and the 'wronged heroine'" (PDF). Nature. 421 (6921): 407–08. Bibcode:2003Natur.421..407M. doi:10.1038/nature01399. PMID 12540909. S2CID 4428347. Archived (PDF) from the original on 17 October 2016.
- Crick FH (1955). A Note for the RNA Tie Club (PDF) (Speech). Cambridge, England. Archived from the original (PDF) on 1 October 2008.
- Meselson M, Stahl FW (July 1958). "The Replication of DNA in Escherichia Coli". Proceedings of the National Academy of Sciences of the United States of America. 44 (7): 671–82. Bibcode:1958PNAS...44..671M. doi:10.1073/pnas.44.7.671. PMC 528642. PMID 16590258.
- "The Nobel Prize in Physiology or Medicine 1968". Nobelprize.org.
- Pray L (2008). "Discovery of DNA structure and function: Watson and Crick". Nature Education. 1 (1): 100.
- Panneerchelvam S, Norazmi MN (2003). "Forensic DNA Profiling and Database". The Malaysian Journal of Medical Sciences. 10 (2): 20–26. PMC 3561883. PMID 23386793.
Further reading
- Berry A, Watson J (2003). DNA: the secret of life. New York: Alfred A. Knopf. ISBN 0-375-41546-7.
- Calladine CR, Drew HR, Luisi BF, Travers AA (2003). Understanding DNA: the molecule & how it works. Amsterdam: Elsevier Academic Press. ISBN 0-12-155089-3.
- Carina D, Clayton J (2003). 50 years of DNA. Basingstoke: Palgrave Macmillan. ISBN 1-4039-1479-6.
- Judson HF (1979). The Eighth Day of Creation: Makers of the Revolution in Biology (2nd ed.). Cold Spring Harbor Laboratory Press. ISBN 0-671-22540-5.
- Olby RC (1994). The path to the double helix: the discovery of DNA. New York: Dover Publications. ISBN 0-486-68117-3. First published in October 1974 by MacMillan, with foreword by Francis Crick; the definitive DNA textbook, revised in 1994 with a nine-page postscript.
- Olby R (January 2003). "Quiet debut for the double helix". Nature. 421 (6921): 402–05. Bibcode:2003Natur.421..402O. doi:10.1038/nature01397. PMID 12540907.
- Olby RC (2009). Francis Crick: A Biography. Plainview, N.Y: Cold Spring Harbor Laboratory Press. ISBN 978-0-87969-798-3.
- Micklas D (2003). DNA Science: A First Course. Cold Spring Harbor Press. ISBN 978-0-87969-636-8.
- Ridley M (2006). Francis Crick: discoverer of the genetic code. Ashland, OH: Eminent Lives, Atlas Books. ISBN 0-06-082333-X.
- Rosenfeld I (2010). DNA: A Graphic Guide to the Molecule that Shook the World. Columbia University Press. ISBN 978-0-231-14271-7.
- Schultz M, Cannon Z (2009). The Stuff of Life: A Graphic Guide to Genetics and DNA. Hill and Wang. ISBN 978-0-8090-8947-5.
- Stent GS, Watson J (1980). The Double Helix: A Personal Account of the Discovery of the Structure of DNA. New York: Norton. ISBN 0-393-95075-1.
- Watson J (2004). DNA: The Secret of Life. Random House. ISBN 978-0-09-945184-6.
- Wilkins M (2003). The third man of the double helix the autobiography of Maurice Wilkins. Cambridge, England: University Press. ISBN 0-19-860665-6.
External links
Library resources aboutDNA
Listen to this article (34 minutes)
- DNA binding site prediction on protein
- DNA the Double Helix Game From the official Nobel Prize web site
- DNA under electron microscope
- Dolan DNA Learning Center
- Double Helix: 50 years of DNA, Nature
- Proteopedia DNA
- Proteopedia Forms_of_DNA
- ENCODE threads explorer ENCODE home page at Nature
- Double Helix 1953–2003 National Centre for Biotechnology Education
- Genetic Education Modules for Teachers – DNA from the Beginning Study Guide
- PDB Molecule of the Month DNA
- "Clue to chemistry of heredity found". The New York Times, June 1953. First American newspaper coverage of the discovery of the DNA structure
- DNA from the Beginning Another DNA Learning Center site on DNA, genes, and heredity from Mendel to the human genome project.
- The Register of Francis Crick Personal Papers 1938 – 2007 at Mandeville Special Collections Library, University of California, San Diego
- Seven-page, handwritten letter that Crick sent to his 12-year-old son Michael in 1953 describing the structure of DNA. See Crick's medal goes under the hammer, Nature, 5 April 2013.
| Gene expression | |||||||
|---|---|---|---|---|---|---|---|
| Introduction to genetics | |||||||
| Transcription |
| ||||||
| Translation |
| ||||||
| Regulation | |||||||
| Influential people | |||||||
| Types of nucleic acids | |||||||
|---|---|---|---|---|---|---|---|
| Constituents | |||||||
| Ribonucleic acids (coding, non-coding) |
| ||||||
| Deoxyribonucleic acids | |||||||
| Analogues | |||||||
| Cloning vectors | |||||||
 Media from Commons
Media from Commons Quotations from Wikiquote
Quotations from Wikiquote Resources from Wikiversity
Resources from Wikiversity Data from Wikidata
Data from Wikidata