| Revision as of 11:22, 15 April 2008 editTouchstone42 (talk | contribs)Extended confirmed users687 edits →Further reading: Added current citation← Previous edit | Latest revision as of 01:38, 26 December 2024 edit undoCitation bot (talk | contribs)Bots5,406,500 edits Altered template type. Add: doi, pages, issue, volume, journal, pmid, bibcode. Removed URL that duplicated identifier. | Use this bot. Report bugs. | Suggested by Dominic3203 | #UCB_webform 173/3850 | ||
| (788 intermediate revisions by more than 100 users not shown) | |||
| Line 1: | Line 1: | ||
| {{Short description|Transfer of genes from unrelated organisms}} | |||
| {{redirect|HGT}} | |||
| {{cs1 config|name-list-style=vanc|display-authors=6}} | |||
| ] showing high rates of horizontal gene transfer between organisms.]] | |||
| {{Redirect|HGT}} | |||
| '''Horizontal gene transfer (HGT)''', also '''Lateral gene transfer (LGT)''', is any process in which an organism transfers genetic material to another cell that is not its offspring. By contrast, ''vertical transfer'' occurs when an organism receives genetic material from its ancestor, e.g. its parent or a species from which it evolved. Most thinking in ] has focused on the more prevalent vertical transfer, but there is a recent awareness that horizontal gene transfer is a significant phenomenon. | |||
| {{About|the natural process|artificial gene transfer|Gene delivery}} | |||
| ] | |||
| '''Horizontal gene transfer''' ('''HGT''') or '''lateral gene transfer''' ('''LGT''')<ref>{{cite journal | vauthors = Ochman H, Lawrence JG, Groisman EA | title = Lateral gene transfer and the nature of bacterial innovation | journal = Nature | volume = 405 | issue = 6784 | pages = 299–304 | date = May 2000 | pmid = 10830951 | doi = 10.1038/35012500 | s2cid = 85739173 | bibcode = 2000Natur.405..299O }}</ref><ref>{{cite journal | vauthors = Dunning Hotopp JC | title = Horizontal gene transfer between bacteria and animals | journal = Trends in Genetics | volume = 27 | issue = 4 | pages = 157–63 | date = April 2011 | pmid = 21334091 | pmc = 3068243 | doi = 10.1016/j.tig.2011.01.005 }}</ref><ref>{{cite journal | vauthors = Robinson KM, Sieber KB, Dunning Hotopp JC | title = A review of bacteria-animal lateral gene transfer may inform our understanding of diseases like cancer | journal = PLOS Genetics | volume = 9 | issue = 10 | pages = e1003877 | date = October 2013 | pmid = 24146634 | pmc = 3798261 | doi = 10.1371/journal.pgen.1003877 | doi-access = free }}</ref> is the movement of genetic material between ]s other than by the ("vertical") transmission of ] from parent to offspring (]).<ref>{{cite journal | vauthors = Keeling PJ, Palmer JD | title = Horizontal gene transfer in eukaryotic evolution | journal = Nature Reviews. Genetics | volume = 9 | issue = 8 | pages = 605–18 | date = August 2008 | pmid = 18591983 | doi = 10.1038/nrg2386 | s2cid = 213613 | author-link2 = Jeffrey D. Palmer | author-link1 = Patrick J. Keeling }}</ref> HGT is an important factor in the evolution of many organisms.<ref name="Gyles_2014">{{cite journal | vauthors = Gyles C, Boerlin P | title = Horizontally transferred genetic elements and their role in pathogenesis of bacterial disease | journal = Veterinary Pathology | volume = 51 | issue = 2 | pages = 328–40 | date = March 2014 | pmid = 24318976 | doi = 10.1177/0300985813511131 | s2cid = 206510894 | doi-access = free }}</ref><ref>{{cite journal |doi=10.1111/bij.12872 |title=Speciation through the looking-glass |journal=Biological Journal of the Linnean Society |volume=120 |issue=2 |pages=480–488 |year=2017 | vauthors = Vaux F, Trewick SA, Morgan-Richards M |doi-access=free }}</ref> HGT is influencing scientific understanding of higher-order evolution while more significantly shifting perspectives on bacterial evolution.<ref name="Ochman_2005">{{cite journal | vauthors = Ochman H, Lerat E, Daubin V | title = Examining bacterial species under the specter of gene transfer and exchange | journal = Proceedings of the National Academy of Sciences of the United States of America | volume = 102 | issue = Suppl 1 | pages = 6595–6599 | date = May 2005 | pmid = 15851673 | pmc = 1131874 | doi = 10.1073/pnas.0502035102 | bibcode = 2005PNAS..102.6595O | doi-access = free }}</ref> | |||
| Horizontal gene transfer is the primary mechanism for the spread of ] in bacteria,<ref>{{cite journal | vauthors = Huddleston JR | title = Horizontal gene transfer in the human gastrointestinal tract: potential spread of antibiotic resistance genes | journal = Infection and Drug Resistance | volume = 7 | pages = 167–176 | date = 2014 | pmid = 25018641 | pmc = 4073975 | doi = 10.2147/idr.s48820 | doi-access = free }}</ref><ref name="Gyles_2014"/><ref>{{cite journal | vauthors = Koonin EV, Makarova KS, Aravind L | title = Horizontal gene transfer in prokaryotes: quantification and classification | journal = Annual Review of Microbiology | volume = 55 | issue = 1 | pages = 709–42 | year = 2001 | pmid = 11544372 | pmc = 4781227 | doi = 10.1146/annurev.micro.55.1.709 }}</ref><ref>{{cite journal | vauthors = Nielsen KM | title = Barriers to horizontal gene transfer by natural transformation in soil bacteria | journal = APMIS | volume = 84 | issue = S84 | pages = 77–84 | year = 1998 | pmid = 9850687 | doi = 10.1111/j.1600-0463.1998.tb05653.x | s2cid = 26490197 }}</ref> and plays an important role in the evolution of ] that can degrade novel compounds such as human-created ]<ref>{{cite journal | vauthors = McGowan C, Fulthorpe R, Wright A, Tiedje JM | title = Evidence for interspecies gene transfer in the evolution of 2,4-dichlorophenoxyacetic acid degraders | journal = Applied and Environmental Microbiology | volume = 64 | issue = 10 | pages = 4089–92 | date = October 1998 | pmid = 9758850 | pmc = 106609 | doi = 10.1128/AEM.64.10.4089-4092.1998 | bibcode = 1998ApEnM..64.4089M }}</ref> and in the evolution, maintenance, and transmission of ].<ref name="Keen_2012">{{cite journal | vauthors = Keen EC | title = Paradigms of pathogenesis: targeting the mobile genetic elements of disease | journal = Frontiers in Cellular and Infection Microbiology | volume = 2 | page = 161 | date = December 2012 | pmid = 23248780 | pmc = 3522046 | doi = 10.3389/fcimb.2012.00161 | doi-access = free }}</ref> It often involves ] ]s and ].<ref>{{cite journal |vauthors=Naik GA, Bhat LN, Chpoade BA, Lynch JM |s2cid=21015053 |title=Transfer of broad-host-range antibiotic resistance plasmids in soil microcosms |journal=Curr. Microbiol. |volume=28 |year=1994 |pages=209–215 |doi=10.1007/BF01575963 |issue=4}}</ref><ref>{{cite journal | vauthors = Varga M, Kuntová L, Pantůček R, Mašlaňová I, Růžičková V, Doškař J | title = Efficient transfer of antibiotic resistance plasmids by transduction within methicillin-resistant Staphylococcus aureus USA300 clone | journal = FEMS Microbiology Letters | volume = 332 | issue = 2 | pages = 146–52 | date = July 2012 | pmid = 22553940 | doi = 10.1111/j.1574-6968.2012.02589.x | doi-access = }}</ref><ref>{{cite journal | vauthors = Varga M, Pantu Ček R, Ru Žičková V, Doškař J | title = Molecular characterization of a new efficiently transducing bacteriophage identified in meticillin-resistant Staphylococcus aureus | journal = The Journal of General Virology | volume = 97 | issue = 1 | pages = 258–268 | date = January 2016 | pmid = 26537974 | doi = 10.1099/jgv.0.000329 | doi-access = free }}</ref> Genes responsible for antibiotic resistance in one species of bacteria can be transferred to another species of bacteria through various mechanisms of HGT such as ], ] and ], subsequently arming the antibiotic resistant genes' recipient against antibiotics. The rapid spread of antibiotic resistance genes in this manner is becoming a challenge to manage in the field of medicine. Ecological factors may also play a role in the HGT of antibiotic resistant genes.<ref>{{cite journal | vauthors = Cairns J, Ruokolainen L, Hultman J, Tamminen M, Virta M, Hiltunen T | title = Ecology determines how low antibiotic concentration impacts community composition and horizontal transfer of resistance genes | journal = Communications Biology | volume = 1 | issue = 1 | page = 35 | date = 2018-04-19 | pmid = 30271921 | pmc = 6123812 | doi = 10.1038/s42003-018-0041-7 }}</ref> | |||
| Horizontal gene transfer is recognized as a pervasive evolutionary process that distributes genes between divergent prokaryotic lineages<ref name="Zhou_2021">{{cite journal | vauthors = Zhou H, Beltrán JF, Brito IL | title = Functions predict horizontal gene transfer and the emergence of antibiotic resistance | journal = Science Advances | volume = 7 | issue = 43 | pages = eabj5056 | date = October 2021 | pmid = 34678056 | doi = 10.1126/sciadv.abj5056 | pmc = 8535800 | bibcode = 2021SciA....7.5056Z }}</ref> and can also involve eukaryotes.<ref name="Sieber_2017">{{cite journal | vauthors = Sieber KB, Bromley RE, Dunning Hotopp JC | title = Lateral gene transfer between prokaryotes and eukaryotes | journal = Experimental Cell Research | volume = 358 | issue = 2 | pages = 421–426 | date = September 2017 | pmid = 28189637 | doi = 10.1016/j.yexcr.2017.02.009 | pmc = 5550378 }}</ref><ref name="Gabaldón_2021">{{cite journal | vauthors = Gabaldón T | title = Origin and Early Evolution of the Eukaryotic Cell | journal = Annual Review of Microbiology | volume = 75 | issue = 1 | pages = 631–647 | date = October 2021 | pmid = 34343017 | doi = 10.1146/annurev-micro-090817-062213 | s2cid = 236916203 }}</ref> HGT events are thought to occur less frequently in eukaryotes than in prokaryotes. However, growing evidence indicates that HGT is relatively common among many eukaryotic species and can have an impact on adaptation to novel environments. Its study, however, is hindered by the complexity of eukaryotic genomes and the abundance of repeat-rich regions, which complicate the accurate identification and characterization of transferred genes.<ref>{{Cite journal |last1=Brockhurst |first1=Michael A. |last2=Harrison |first2=Ellie |last3=Hall |first3=James P.J. |last4=Richards |first4=Thomas |last5=McNally |first5=Alan |last6=MacLean |first6=Craig |date=October 2019 |title=The Ecology and Evolution of Pangenomes |url=http://dx.doi.org/10.1016/j.cub.2019.08.012 |journal=Current Biology |volume=29 |issue=20 |pages=R1094–R1103 |doi=10.1016/j.cub.2019.08.012 |pmid=31639358 |bibcode=2019CBio...29R1094B |issn=0960-9822}}</ref><ref>{{Cite journal |last1=Van Etten |first1=Julia |last2=Bhattacharya |first2=Debashish |date=December 2020 |title=Horizontal Gene Transfer in Eukaryotes: Not if, but How Much? |url=https://linkinghub.elsevier.com/retrieve/pii/S0168952520302067 |journal=Trends in Genetics |language=en |volume=36 |issue=12 |pages=915–925 |doi=10.1016/j.tig.2020.08.006|pmid=33012528 |bibcode=2020TGene..36..915V }}</ref> | |||
| It is postulated that HGT promotes the maintenance of a universal life biochemistry and, subsequently, the universality of the genetic code.<ref>{{cite journal | vauthors = Kubyshkin V, Acevedo-Rocha CG, Budisa N | title = On universal coding events in protein biogenesis | journal = Bio Systems | volume = 164 | pages = 16–25 | date = February 2018 | pmid = 29030023 | doi = 10.1016/j.biosystems.2017.10.004 | doi-access = free | bibcode = 2018BiSys.164...16K }}</ref> | |||
| ==History== | ==History== | ||
| Horizontal gene transfer was first described in Japan in a 1959 publication that demonstrated the transfer of antibiotic resistance between different species of bacteria.<ref>Ochiai, K., Yamanaka, T Kimura K and Sawada, O (1959) Inheritance of drug resistance (and its tranfer) between Shigella strains and Between Shigella and E.coli strains. Hihon Iji Shimpor 1861: 34 (in Japanese)</ref> <ref>Akiba T, Koyama K, Ishiki Y, Kimura S, Fukushima T. On the mechanism of the development of multiple-drug-resistant clones of Shigella. Jpn J Microbiol. 1960 Apr;4:219-27. PMID 13681921.</ref> However, the significance of this research was not appreciated in the west for another ten years. Michael Syvanen was among the earliest western biologists to explore the potential significance of lateral gene transfer. Syvanen published a series of papers on horizontal gene transfer starting in ] <ref>{{cite journal | author = Syvanen, Michael | year = 1985 | title = Cross-species Gene Transfer; Implications for a New Theory of Evolution | journal = J. Theor. Biol. | volume = 112 | pages pp. 333-343 | url = http://www.dcn.davis.ca.us/vme/hgt/JTheoBiolvol112pp333-343yr1985.PDF | accessdate = 2007-09-05}}</ref>, predicting that lateral gene transfer exists, has biological significance, and is a process that shaped evolutionary history from the very beginning of life on earth. Artificial horizontal gene transfer is a form of ]. | |||
| ], reported in 1928 by ],<ref>{{cite journal | vauthors = Griffith F | title = The Significance of Pneumococcal Types | journal = The Journal of Hygiene | volume = 27 | issue = 2 | pages = 113–59 | date = January 1928 | pmid = 20474956 | pmc = 2167760 | doi = 10.1017/S0022172400031879 | publisher = Cambridge University Press | author-link = Frederick Griffith | jstor = 4626734 }}</ref> was the first experiment suggesting that bacteria are capable of transferring genetic information through a process known as ].<ref>{{cite journal | vauthors = Lorenz MG, Wackernagel W | title = Bacterial gene transfer by natural genetic transformation in the environment | journal = Microbiological Reviews | volume = 58 | issue = 3 | pages = 563–602 | date = September 1994 | pmid = 7968924 | pmc = 372978 | doi = 10.1128/MMBR.58.3.563-602.1994 }}</ref><ref>{{cite journal | vauthors = Downie AW | title = Pneumococcal transformation--a backward view. Fourth Griffith Memorial Lecture | journal = Journal of General Microbiology | volume = 73 | issue = 1 | pages = 1–11 | date = November 1972 | pmid = 4143929 | doi = 10.1099/00221287-73-1-1 | url = http://mic.sgmjournals.org/content/73/1/1.full.pdf | doi-access = free | access-date = 2018-05-23 | archive-date = 2012-03-02 | archive-url = https://web.archive.org/web/20120302055327/http://mic.sgmjournals.org/content/73/1/1.full.pdf | url-status = live }}</ref> Griffith's findings ] in the late 1930s and early 1940s that isolated ] as the material that communicated this genetic information. | |||
| As Jain, Rivera and Lake (1999) put it: "Increasingly, studies of genes and genomes are indicating that considerable horizontal transfer has occurred between ]s."<ref>{{cite journal | author = Lake, James A. and Maria C. Rivera | year = 1999 | title = Horizontal gene transfer among genomes: The complexity hypothesis |journal = PNAS (Proceedings of the National Academy of Science) | volume = 96:7 | pages = pp. 3801-3806 | url = http://www.pnas.org/cgi/content/abstract/96/7/3801 | accessdate = 2007-03-18}}</ref> (see also Lake and Rivera, 2007).<ref>{{cite journal | author = Lake, James A. and Maria C. Rivera | year = 2004 | title = The Ring of Life Provides Evidence for a Genome Fusion Origin of Eukaryotes |journal = ] | volume = 431 | accessdate = 2007-03-16}}</ref> The phenomenon appears to have had some significance for ] ]s as well. As Bapteste et al. (2005) observe, "additional evidence suggests that gene transfer might also be an important evolutionary mechanism in ] evolution."<ref>{{cite journal | author = Bapteste et al. | year = 2005 | title = Do Orthologous Gene Phylogenies Really Support Tree-thinking? |journal = BMC Evolutionary Biology | volume = 5:33 | url = http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pubmed&pubmedid=15913459 | accessdate = 2007-03-18}}</ref> | |||
| Horizontal genetic transfer was then described in Seattle in 1951, in a paper demonstrating that the transfer of a viral gene into '']'' created a virulent strain from a non-virulent strain,<ref>{{cite journal | vauthors = Freeman VJ | title = Studies on the virulence of bacteriophage-infected strains of Corynebacterium diphtheriae | journal = Journal of Bacteriology | volume = 61 | issue = 6 | pages = 675–88 | date = June 1951 | pmid = 14850426 | pmc = 386063 | doi = 10.1128/JB.61.6.675-688.1951 }}</ref> simultaneously revealing the mechanism of ] (that patients could be infected with the bacteria but not have any symptoms, and then suddenly convert later or never),<ref>{{cite book | vauthors = Margulies P |title=Diphtheria | series = Epidemics: Deadly diseases throughout history |date=2005 |publisher=Rosen Publishing Group |location=New York |isbn=978-1-4042-0253-5 |edition=1st}}</ref> and giving the first example for the relevance of the ].<ref>{{cite web | vauthors = Lwoff A | date = 1965 | url = http://nobelprize.org/nobel_prizes/medicine/laureates/1965/lwoff-lecture.html | title = Interaction among Virus, Cell, and Organism | work = Nobel Lecture for the Nobel Prize in Physiology or Medicine| archive-url = https://web.archive.org/web/20101016190354/http://nobelprize.org/nobel_prizes/medicine/laureates/1965/lwoff-lecture.html | archive-date=2010-10-16 }}</ref> Inter-bacterial gene transfer was first described in Japan in a 1959 publication that demonstrated the transfer of antibiotic resistance between different species of ].<ref>{{cite journal | vauthors = Ochiai K, Yamanaka T, Kimura K, Sawada O |title=Inheritance of drug resistance (and its transfer) between Shigella strains and Between Shigella and E. coli strains |journal=Hihon Iji Shimpor |volume=1861 |page=34 |year=1959 |language=ja}}</ref><ref>{{cite journal | vauthors = Akiba T, Koyama K, Ishiki Y, Kimura S, Fukushima T | title = On the mechanism of the development of multiple-drug-resistant clones of Shigella | journal = Japanese Journal of Microbiology | volume = 4 | issue = 2 | pages = 219–27 | date = April 1960 | pmid = 13681921 | doi = 10.1111/j.1348-0421.1960.tb00170.x | doi-access = free }}</ref> In the mid-1980s, Syvanen<ref>{{cite journal | vauthors = Syvanen M | title = Cross-species gene transfer; implications for a new theory of evolution | journal = Journal of Theoretical Biology | volume = 112 | issue = 2 | pages = 333–43 | date = January 1985 | pmid = 2984477 | doi = 10.1016/S0022-5193(85)80291-5 | bibcode = 1985JThBi.112..333S | url = http://www.dcn.davis.ca.us/vme/hgt/JTheoBiolvol112pp333-343yr1985.PDF | access-date = 2009-01-13 | archive-date = 2017-07-06 | archive-url = https://web.archive.org/web/20170706072346/http://www.dcn.davis.ca.us/vme/hgt/JTheoBiolvol112pp333-343yr1985.PDF | url-status = live }}</ref> postulated that biologically significant lateral gene transfer has existed since the beginning of life on Earth and has been involved in shaping all of evolutionary history. | |||
| There is some evidence that even higher plants and animals have been affected. Dr. ], a noted scientist and critic of ], writes: "While horizontal gene transfer is well-known among bacteria, it is only within the past 10 years that its occurrence has become recognized among higher plants and animals. The scope for horizontal gene transfer is essentially the entire biosphere, with bacteria and viruses serving both as intermediaries for gene trafficking and as reservoirs for gene multiplication and recombination (the process of making new combinations of genetic material)."<ref name = "Mae-Wan Ho"> Dr ]</ref> But Richardson and Palmer (2007) are more cautious: "Horizontal gene transfer (HGT) has played a major role in bacterial evolution and is fairly common in certain ] eukaryotes. However, the prevalence and importance of HGT in the evolution of ] eukaryotes remain unclear."<ref>{{cite journal | author = Richardson, Aaron O. and Jeffrey D. Palmer | year = January 2007 | title = Horizontal Gene Transfer in Plants |journal = Journal of Experimental Botany | volume = 58 | pages = pp. 1-9 | accessdate = 2007-03-18}}</ref> | |||
| As Jian, Rivera and Lake (1999) put it: "Increasingly, studies of genes and genomes are indicating that considerable horizontal transfer has occurred between ]s"<ref>{{cite journal | vauthors = Jain R, Rivera MC, Lake JA | title = Horizontal gene transfer among genomes: the complexity hypothesis | journal = Proceedings of the National Academy of Sciences of the United States of America | volume = 96 | issue = 7 | pages = 3801–6 | date = March 1999 | pmid = 10097118 | pmc = 22375 | doi = 10.1073/pnas.96.7.3801 | bibcode = 1999PNAS...96.3801J | doi-access = free }}</ref> (see also Lake and Rivera, 2007).<ref>{{cite journal | vauthors = Rivera MC, Lake JA | title = The ring of life provides evidence for a genome fusion origin of eukaryotes | journal = Nature | volume = 431 | issue = 7005 | pages = 152–5 | date = September 2004 | pmid = 15356622 | doi = 10.1038/nature02848 | url = http://www.sdsc.edu/~shindyal/ejc121304.pdf | s2cid = 4349149 | archive-url = https://web.archive.org/web/20070927135534/http://www.sdsc.edu/~shindyal/ejc121304.pdf | bibcode = 2004Natur.431..152R | archive-date = 2007-09-27 }}</ref> The phenomenon appears to have had some significance for unicellular ]s as well. As Bapteste et al. (2005) observe, "additional evidence suggests that gene transfer might also be an important evolutionary mechanism in ] evolution."<ref>{{cite journal | vauthors = Bapteste E, Susko E, Leigh J, MacLeod D, Charlebois RL, Doolittle WF | title = Do orthologous gene phylogenies really support tree-thinking? | journal = BMC Evolutionary Biology | volume = 5 | issue = 1 | page = 33 | date = May 2005 | pmid = 15913459 | pmc = 1156881 | doi = 10.1186/1471-2148-5-33 | bibcode = 2005BMCEE...5...33B | doi-access = free }}</ref> | |||
| Due to the increasing amount of evidence suggesting the importance of these phenomena for evolution (see ]), molecular biologists such as Peter Gogarten have described horizontal gene transfer as "A New Paradigm for Biology".<ref name="gogarten">{{cite journal | author = Gogarten, Peter | year = 2000 | title = Horizontal Gene Transfer: A New Paradigm for Biology | journal = Esalen Center for Theory and Research Conference | url = http://www.esalenctr.org/display/confpage.cfm?confid=10&pageid=105&pgtype=1 | accessdate = 2007-03-18}}</ref> | |||
| Grafting of one plant to another can transfer ]s (]s in plant cells that conduct ]), ], and the entire ] containing the ] to potentially make a new species.<ref>{{cite web|url=https://www.newscientist.com/article/2079813-farmers-may-have-been-accidentally-making-gmos-for-millennia/|title=Farmers may have been accidentally making GMOs for millennia|vauthors=Le Page M|date=2016-03-17|publisher=The New Scientist|language=en|access-date=2016-07-11|archive-date=2018-10-01|archive-url=https://web.archive.org/web/20181001031232/https://www.newscientist.com/article/2079813-farmers-may-have-been-accidentally-making-gmos-for-millennia/|url-status=live}}</ref> Some ] (e.g. ] and ]s) have been genetically modified by horizontal gene transfer from the wasp ].<ref>{{cite journal | vauthors = Gasmi L, Boulain H, Gauthier J, Hua-Van A, Musset K, Jakubowska AK, Aury JM, Volkoff AN, Huguet E, Herrero S, Drezen JM | title = Recurrent Domestication by Lepidoptera of Genes from Their Parasites Mediated by Bracoviruses | journal = PLOS Genetics | volume = 11 | issue = 9 | pages = e1005470 | date = September 2015 | pmid = 26379286 | pmc = 4574769 | doi = 10.1371/journal.pgen.1005470 | doi-access = free }}</ref> Bites from insects in the family ] (assassin bugs) can, via a parasite, infect humans with the ]l ], which can insert its DNA into the human genome.<ref>{{cite web |url=http://phenomena.nationalgeographic.com/2010/02/14/genes-from-chagas-parasite-can-transfer-to-humans-and-be-passed-on-to-children/|archive-url=https://web.archive.org/web/20130106204712/http://phenomena.nationalgeographic.com/2010/02/14/genes-from-chagas-parasite-can-transfer-to-humans-and-be-passed-on-to-children/|archive-date=January 6, 2013|title=Genes from Chagas parasite can transfer to humans and be passed on to children| vauthors = Yong E |date=2010-02-14|publisher=National Geographic|language=en|access-date=2016-07-13}}<br>{{cite journal |last1=Hecht |first1=Mariana M. |last2=Nitz |first2=Nadjar |last3=Araujo |first3=Perla F. |last4=Sousa |first4=Alessandro O. |last5=Rosa |first5=Ana de Cássia |last6=Gomes |first6=Dawidson A. |last7=Leonardecz |first7=Eduardo |last8=Teixeira |first8=Antonio R. L. |title=Inheritance of DNA Transferred from American Trypanosomes to Human Hosts |journal=PLOS ONE |date=12 February 2010 |volume=5 |issue=2 |pages=e9181 |doi=10.1371/journal.pone.0009181 |doi-access=free |pmid=20169193|pmc=2820539 |bibcode=2010PLoSO...5.9181H }}</ref> It has been suggested that lateral gene transfer to humans from bacteria may play a role in cancer.<ref>{{cite journal | vauthors = Riley DR, Sieber KB, Robinson KM, White JR, Ganesan A, Nourbakhsh S, Dunning Hotopp JC | title = Bacteria-human somatic cell lateral gene transfer is enriched in cancer samples | journal = PLOS Computational Biology | volume = 9 | issue = 6 | pages = e1003107 | year = 2013 | pmid = 23840181 | pmc = 3688693 | doi = 10.1371/journal.pcbi.1003107 | bibcode = 2013PLSCB...9E3107R | doi-access = free }}</ref> | |||
| It should also be noted that the process is emphasised by Dr. Mae-Wan Ho as an important factor in "The Hidden Hazards of Genetic Engineering", as it may allow dangerous ] ] (which is optimised for transfer) to spread from species to species.<ref name = "Mae-Wan Ho"/> | |||
| Aaron Richardson and ] state: "Horizontal gene transfer (HGT) has played a major role in bacterial evolution and is fairly common in certain unicellular eukaryotes. However, the prevalence and importance of HGT in the evolution of ] eukaryotes remain unclear."<ref>{{cite journal | vauthors = Richardson AO, Palmer JD | title = Horizontal gene transfer in plants | journal = Journal of Experimental Botany | volume = 58 | issue = 1 | pages = 1–9 | year = 2007 | pmid = 17030541 | doi = 10.1093/jxb/erl148 | url = http://www.sdsc.edu/~shindyal/ejc121304.pdf | archive-url = https://web.archive.org/web/20070927135534/http://www.sdsc.edu/~shindyal/ejc121304.pdf | archive-date = 2007-09-27 | author-link2 = Jeffrey D. Palmer | doi-access = free }}</ref> | |||
| Due to the increasing amount of evidence suggesting the importance of these phenomena for evolution (see ]) molecular biologists such as Peter Gogarten have described horizontal gene transfer as "A New Paradigm for Biology".<ref name="Gogarten_2000">{{cite journal | vauthors = Gogarten P |year= 2000 |title= Horizontal Gene Transfer: A New Paradigm for Biology |journal= Esalen Center for Theory and Research Conference |url= http://www.esalenctr.org/display/confpage.cfm?confid=10&pageid=105&pgtype=1 |access-date= 2007-03-18 |archive-date= 2012-07-21 |archive-url= https://web.archive.org/web/20120721232310/http://www.esalenctr.org/display/confpage.cfm?confid=10&pageid=105&pgtype=1 }}</ref> | |||
| ==Mechanisms== | |||
| There are several mechanisms for horizontal gene transfer:<ref name="Gyles_2014"/><ref name="Todar_2012">{{cite book | chapter-url = http://textbookofbacteriology.net/themicrobialworld/bactresanti.html| chapter = Bacterial Resistance to Antibiotics| vauthors = Todar K |title = The Microbial World: Lectures in Microbiology | publisher = Department of Bacteriology, University of Wisconsin-Madison|access-date=January 6, 2012|archive-url=https://web.archive.org/web/20120115211044/http://textbookofbacteriology.net/themicrobialworld/bactresanti.html|archive-date=January 15, 2012}}</ref><ref name="Maloy_2002">{{cite web|url=http://www.sci.sdsu.edu/~smaloy/MicrobialGenetics/topics/genetic-exchange/exchange/exchange.html|title=Horizontal Gene Transfer| vauthors = Maloy S |date=July 15, 2002|publisher=San Diego State University|access-date=January 6, 2012|archive-date=February 14, 2019|archive-url=https://web.archive.org/web/20190214211829/http://www.sci.sdsu.edu/~smaloy/MicrobialGenetics/topics/genetic-exchange/exchange/exchange.html|url-status=live}}</ref> | |||
| *], the genetic alteration of a ] resulting from the introduction, uptake and ] of foreign genetic material (] or ]).<ref name="Stearns_2005">{{cite book | vauthors = Stearns SC, Hoekstra RF | date = 2005 | title = Evolution: An introduction | edition = 2nd | location = Oxford, New York | publisher = Oxford Univ. Press | pages = 38–40 | isbn = 978-0-19-925563-4 }}</ref> This process is relatively common in bacteria, but less so in eukaryotes.<ref>{{cite book |doi=10.1007/978-94-007-2920-9_10 |publisher=Springer Science+Business Media B.V. |chapter=Horizontal Gene Transfer in Eukaryotes: Fungi-to-Plant and Plant-to-Plant Transfers of Organellar DNA |title=Genomics of Chloroplasts and Mitochondria |series=Advances in Photosynthesis and Respiration |year=2012 | vauthors = Renner SS, Bellot S |volume=35 |pages=223–235 |isbn=978-94-007-2919-3 }}</ref> Transformation is often used in laboratories to insert novel genes into bacteria for experiments or for industrial or medical applications. See also ] and ].{{citation needed|date=February 2023}} | |||
| *], the process in which bacterial DNA is moved from one bacterium to another by a virus (a bacteriophage, or ]).<ref name="Stearns_2005"/> | |||
| *], a process that involves the transfer of DNA via a plasmid from a donor cell to a recombinant recipient cell during cell-to-cell contact.<ref name="Stearns_2005"/> | |||
| *]s, virus-like elements encoded by the host that are found in the ] order ].<ref name="McDaniel_2010">{{cite journal | vauthors = Maxmen A |title= Virus-like particles speed bacterial evolution |journal= Nature |year= 2010 |doi= 10.1038/news.2010.507}}</ref> | |||
| ===Horizontal transposon transfer=== | |||
| A ] (TE) (also called a transposon or jumping gene) is a mobile segment of DNA that can sometimes pick up a resistance gene and insert it into a plasmid or chromosome, thereby inducing horizontal gene transfer of antibiotic resistance.<ref name="Stearns_2005"/> | |||
| Horizontal transposon transfer (HTT) refers to the passage of pieces of DNA that are characterized by their ability to move from one ] to another between genomes by means other than parent-to-offspring inheritance. Horizontal gene transfer has long been thought to be crucial to prokaryotic evolution, but there is a growing amount of data showing that HTT is a common and widespread phenomenon in ] evolution as well.<ref name="Schaack_2010">{{cite journal | vauthors = Schaack S, Gilbert C, Feschotte C | title = Promiscuous DNA: horizontal transfer of transposable elements and why it matters for eukaryotic evolution | journal = Trends in Ecology & Evolution | volume = 25 | issue = 9 | pages = 537–46 | date = September 2010 | pmid = 20591532 | pmc = 2940939 | doi = 10.1016/j.tree.2010.06.001 | bibcode = 2010TEcoE..25..537S }}</ref> On the transposable element side, spreading between genomes via horizontal transfer may be viewed as a strategy to escape purging due to purifying selection, mutational decay and/or host defense mechanisms.<ref name="Dupeyron_2014">{{cite journal | vauthors = Dupeyron M, Leclercq S, Cerveau N, Bouchon D, Gilbert C | title = Horizontal transfer of transposons between and within crustaceans and insects | journal = Mobile DNA | volume = 5 | issue = 1 | page = 4 | date = January 2014 | pmid = 24472097 | pmc = 3922705 | doi = 10.1186/1759-8753-5-4 | doi-access = free }}</ref> | |||
| HTT can occur with any type of transposable elements, but ]s and ] are more likely to be capable of HTT because both have a stable, double-stranded DNA intermediate that is thought to be sturdier than the single-stranded RNA intermediate of ], which can be highly degradable.<ref name="Schaack_2010"/> ] may be less likely to transfer horizontally compared to ] because they do not encode the proteins required for their own mobilization. The structure of these non-autonomous elements generally consists of an intronless gene encoding a ] protein, and may or may not have a promoter sequence. Those that do not have promoter sequences encoded within the mobile region rely on adjacent host promoters for expression.<ref name="Schaack_2010"/> Horizontal transfer is thought to play an important role in the TE life cycle.<ref name="Schaack_2010"/> In plants, it appears that ] of the Copia superfamilies, especially those with low copy numbers from the Ale and Ivana lineages, are more likely to undergo horizontal transfer between different plant species.<ref name="Aubin_2023">{{cite journal | vauthors = Aubin E, Llauro C, Garrigue J, Mirouze M, Panaud O, El Baidouri M | title = Genome-wide analysis of horizontal transfer in non-model wild species from a natural ecosystem reveals new insights into genetic exchange in plants | journal = PLOS Genetics | volume = 19 | issue = 10 | pages = e1010964 | date = October 2023 | pmid = 37856455 | pmc = 10586619 | doi = 10.1371/journal.pgen.1010964 | doi-access = free }}</ref> | |||
| HTT has been shown to occur between species and across continents in both plants<ref name="Baidouri_2014">{{cite journal | vauthors = El Baidouri M, Carpentier MC, Cooke R, Gao D, Lasserre E, Llauro C, Mirouze M, Picault N, Jackson SA, Panaud O | title = Widespread and frequent horizontal transfers of transposable elements in plants | journal = Genome Research | volume = 24 | issue = 5 | pages = 831–8 | date = May 2014 | pmid = 24518071 | pmc = 4009612 | doi = 10.1101/gr.164400.113 }}</ref> and animals (Ivancevic et al. 2013), though some TEs have been shown to more successfully colonize the genomes of certain species over others.<ref name="Ivancevic_2013">{{cite journal | vauthors = Ivancevic AM, Walsh AM, Kortschak RD, Adelson DL | title = Jumping the fine LINE between species: horizontal transfer of transposable elements in animals catalyses genome evolution | journal = BioEssays | volume = 35 | issue = 12 | pages = 1071–82 | date = December 2013 | pmid = 24003001 | doi = 10.1002/bies.201300072 | s2cid = 6968210 }}</ref> Both spatial and taxonomic proximity of species has been proposed to favor HTTs in plants and animals.<ref name="Baidouri_2014"/> It is unknown how the density of a population may affect the rate of HTT events within a population, but close proximity due to ] and cross contamination due to crowding have been proposed to favor HTT in both plants and animals.<ref name="Baidouri_2014"/> In plants, the interaction between lianas and trees has been shown to facilitate HTT in natural ecosystems.<ref name="Aubin_2023"/> Successful transfer of a transposable element requires delivery of DNA from donor to host cell (and to the germ line for multi-cellular organisms), followed by integration into the recipient host genome.<ref name="Schaack_2010"/> Though the actual mechanism for the transportation of TEs from donor cells to host cells is unknown, it is established that ] and RNA can circulate in bodily fluid.<ref name="Schaack_2010"/> Many proposed vectors include arthropods, viruses, freshwater snails (Ivancevic et al. 2013), endosymbiotic bacteria,<ref name="Dupeyron_2014"/> and intracellular parasitic bacteria.<ref name="Schaack_2010"/> In some cases, even TEs facilitate transport for other TEs.<ref name="Ivancevic_2013"/> | |||
| The arrival of a new TE in a host genome can have detrimental consequences because TE mobility may induce mutation. However, HTT can also be beneficial by introducing new genetic material into a genome and promoting the shuffling of genes and TE domains among hosts, which can be co-opted by the host genome to perform new functions.<ref name="Ivancevic_2013"/> Moreover, transposition activity increases the TE copy number and generates ] hotspots.<ref name="Wallau_2012">{{cite journal | vauthors = Wallau GL, Ortiz MF, Loreto EL | title = Horizontal transposon transfer in eukarya: detection, bias, and perspectives | journal = Genome Biology and Evolution | volume = 4 | issue = 8 | pages = 689–99 | year = 2012 | pmid = 22798449 | pmc = 3516303 | doi = 10.1093/gbe/evs055 }}</ref> HTT detection is a difficult task because it is an ongoing phenomenon that is constantly changing in frequency of occurrence and composition of TEs inside host genomes. Furthermore, few species have been analyzed for HTT, making it difficult to establish patterns of HTT events between species. These issues can lead to the underestimation or overestimation of HTT events between ancestral and current eukaryotic species.<ref name="Wallau_2012"/> | |||
| ==Methods of detection== | |||
| ]s of a gene in the two daughter species. A horizontal gene transfer event from one species to another adds a ] of the gene to the receiving genome.]] | |||
| {{Main|Inferring horizontal gene transfer}} | |||
| Horizontal gene transfer is typically inferred using ] methods, either by identifying atypical sequence signatures ("parametric" methods) or by identifying strong discrepancies between the evolutionary history of particular sequences compared to that of their hosts. The transferred gene (]) found in the receiving species is more closely related to the genes of the donor species than would be expected.{{citation needed|date=February 2023}} | |||
| ==Viruses== | |||
| The ] called '']'' infects ]. Another virus, called '']'', also infects amoebae, but it cannot reproduce unless mimivirus has already infected the same cell.<ref name="La_Scola_2008">{{cite journal | vauthors = La Scola B, Desnues C, Pagnier I, Robert C, Barrassi L, Fournous G, Merchat M, Suzan-Monti M, Forterre P, Koonin E, Raoult D | title = The virophage as a unique parasite of the giant mimivirus | journal = Nature | volume = 455 | issue = 7209 | pages = 100–4 | date = September 2008 | pmid = 18690211 | doi = 10.1038/nature07218 | s2cid = 4422249 | bibcode = 2008Natur.455..100L }}</ref> <blockquote>Sputnik's ] reveals further insight into its biology. Although 13 of its genes show little similarity to any other known genes, three are closely related to mimivirus and ] genes, perhaps cannibalized by the tiny virus as it packaged up particles sometime in its history. This suggests that the ] could perform horizontal gene transfer between viruses, paralleling the way that bacteriophages ferry genes between bacteria.<ref>{{cite journal | vauthors = Pearson H | title = 'Virophage' suggests viruses are alive | journal = Nature | volume = 454 | issue = 7205 | page = 677 | date = August 2008 | pmid = 18685665 | doi = 10.1038/454677a | doi-access = | bibcode = 2008Natur.454..677P | s2cid = 205040157 }}</ref></blockquote> Horizontal transfer is also seen between geminiviruses and tobacco plants. | |||
| ==Prokaryotes== | ==Prokaryotes== | ||
| Horizontal gene transfer is common among bacteria, even among very distantly related ones. This process is thought to be a significant cause of increased ]<ref name="Gyles_2014"/><ref name="Barlow_2009">{{cite book | vauthors = Barlow M |chapter=What Antimicrobial Resistance Has Taught Us About Horizontal Gene Transfer |title=Horizontal Gene Transfer |volume=532 |pages=397–411 |year=2009 |pmid=19271198 |doi=10.1007/978-1-60327-853-9_23 |series=Methods in Molecular Biology |publisher=Humana Press |location=Totowa, NJ |isbn=978-1-60327-852-2}}</ref> when one bacterial cell acquires resistance, and the resistance genes are transferred to the other species.<ref name="Hawkey_2009">{{cite journal | vauthors = Hawkey PM, Jones AM | title = The changing epidemiology of resistance | journal = The Journal of Antimicrobial Chemotherapy | volume = 64 | issue = Suppl 1 | pages = i3-10 | date = September 2009 | pmid = 19675017 | doi = 10.1093/jac/dkp256 | doi-access = free }}</ref><ref name="Francino_2012">{{cite book |veditors = Francino MP |year=2012 |title=Horizontal Gene Transfer in Microorganisms |publisher=] |isbn= 978-1-908230-10-2}}</ref> Transposition and horizontal gene transfer, along with strong natural selective forces have led to multi-drug resistant strains of '']'' and many other pathogenic bacteria.<ref name="Stearns_2005"/> Horizontal gene transfer also plays a role in the spread of virulence factors, such as ]s and ]s, amongst bacteria.<ref name="Gyles_2014"/> A prime example concerning the spread of exotoxins is the adaptive evolution of ]s in ''E. coli'' through horizontal gene transfer via transduction with '']'' species of bacteria.<ref>{{cite journal | vauthors = Strauch E, Lurz R, Beutin L | title = Characterization of a Shiga toxin-encoding temperate bacteriophage of Shigella sonnei | journal = Infection and Immunity | volume = 69 | issue = 12 | pages = 7588–95 | date = December 2001 | pmid = 11705937 | pmc = 98851 | doi = 10.1128/IAI.69.12.7588-7595.2001 }}</ref> Strategies to combat certain bacterial infections by targeting these specific virulence factors and mobile genetic elements have been proposed.<ref name="Keen_2012"/> For example, horizontally transferred genetic elements play important roles in the virulence of '']'', '']'', '']'' and '']''.<ref name="Gyles_2014"/> | |||
| Horizontal gene transfer is common among ], even very distantly-related ones. This process is thought to be a significant cause of increased ]; when one bacterial cell acquires resistance, it can quickly transfer the resistance genes to many species. Enteric bacteria appear to exchange genetic material with each other within the ] in which they live. There are three common mechanisms for horizontal gene transfer: | |||
| In prokaryotes, restriction-modification systems are known to provide immunity against horizontal gene transfer and in stabilizing mobile genetic elements. Genes encoding restriction modification systems have been reported to move between prokaryotic genomes within ] (MGE) such as ]s, ]s, insertion sequences/transposons, integrative conjugative elements (ICE),<ref>{{cite journal | vauthors = Johnson CM, Grossman AD | title = Integrative and Conjugative Elements (ICEs): What They Do and How They Work | journal = Annual Review of Genetics | volume = 42 | issue = 1 | pages = 577–601 | date = November 2015 | pmid = 26473380 | pmc = 5180612 | doi = 10.1146/annurev-genet-112414-055018 }}</ref> and ]s. Still, they are more frequently a chromosomal-encoded barrier to MGE than an MGE-encoded tool for cell infection.<ref name="Oliveira_2014">{{cite journal | vauthors = Oliveira PH, Touchon M, Rocha EP | title = The interplay of restriction-modification systems with mobile genetic elements and their prokaryotic hosts | journal = Nucleic Acids Research | volume = 49 | issue = 16 | pages = 10618–10631 | date = September 2014 | pmid = 25120263 | pmc = 4176335 | doi = 10.1093/nar/gku734 | url = }}</ref> | |||
| Lateral gene transfer via a mobile genetic element, namely the integrated conjugative element (ICE) ''Bs1'' has been reported for its role in the global DNA damage SOS response of the gram positive ''Bacillus subtilis''.<ref>{{cite journal |pmid = 17511812|pmc=3320793| doi=10.1111/j.1365-2958.2007.05748.x|volume=64 |issue = 6|pages=1515–1528|title=Identification and characterization of the immunity repressor (ImmR) that controls the mobile genetic element ICE ''Bs1'' of ''Bacillus subtilis''|journal= PLOS Genet | vauthors=Auchtung JM, Lee CA, Garrison KL, Grossman AD|date=June 2007| url=}}</ref> Furthermore, it has been linked with the radiation and desiccation resistance of ''Bacillus pumilus'' SAFR-032 spores,<ref>{{cite journal| pmid=23812891| doi=10.1007/s00792-013-0559-z| volume=17| issue=5| pages=767–774| title=An ICE''Bs1''-like element may be associated with the extreme radiation and desiccation resistance of ''Bacillus pumilus'' SAFR-032 spores.| journal=Extremophiles| vauthors=Tirumalai MR, Fox GE| date=September 2013| s2cid=8675124| url=https://link.springer.com/article/10.1007%2Fs00792-013-0559-z| access-date=2020-09-16| archive-date=2021-11-28| archive-url=https://web.archive.org/web/20211128101257/https://link.springer.com/article/10.1007/s00792-013-0559-z| url-status=live}}</ref> isolated from spacecraft cleanroom facilities.<ref>{{cite journal | pmid = 14502417|doi=10.1007/s00248-003-1029-4 | volume=47 | issue = 2 |pages=159–163| title=Extreme spore UV resistance of ''Bacillus pumilus'' isolates obtained from an ultraclean Spacecraft Assembly Facility.|journal= Microb Ecol|vauthors=Link L, Sawyer J, Venkateswaran K, Nicholson W| date=February 2004|bibcode=2004MicEc..47..159L |s2cid=13416635 }}</ref><ref>{{cite journal | pmid = 16332797| pmc=1317311| doi=10.1128/AEM.71.12.8147-8156.2005|volume=71|issue = 12 |pages=8147–8156| title=Survival of spacecraft-associated microorganisms under simulated martian UV irradiation.|journal=Appl Environ Microbiol| vauthors=Newcombe DA, Schuerger AC, Benardini JN, Dickinson D, Tanner R, Venkateswaran K| date=December 2005| bibcode=2005ApEnM..71.8147N|url=}}</ref><ref>{{cite journal| pmid=15941382| doi=10.1089/ast.2005.5.391| volume=5| issue=3| pages=391–405| title=Recurrent isolation of hydrogen peroxide-resistant spores of ''Bacillus pumilus'' from a spacecraft assembly facility.| journal=Astrobiology| vauthors=Kempf MJ, Chen F, Kern R, Venkateswaran K| date=June 2005| bibcode=2005AsBio...5..391K| url=https://www.liebertpub.com/doi/10.1089/ast.2005.5.391| access-date=2020-09-16| archive-date=2022-03-07| archive-url=https://web.archive.org/web/20220307083042/https://www.liebertpub.com/doi/10.1089/ast.2005.5.391| url-status=live}}</ref> | |||
| Transposon insertion elements have been reported to increase the fitness of gram-negative '']'' strains through either major transpositions or genome rearrangements, and increasing mutation rates.<ref>{{cite journal|pmid=6303898|pmc=1202041|volume=103|issue=4|pages=581–592|title=Evolution of transposons: natural selection for Tn5 in ''Escherichia coli'' K12|journal=Genetics|vauthors=Biel SW, Hartl DL|date=June 1983|doi=10.1093/genetics/103.4.581|url=https://www.genetics.org/content/103/4/581.long|access-date=2020-09-16|archive-date=2021-08-19|archive-url=https://web.archive.org/web/20210819032655/https://www.genetics.org/content/103/4/581.long|url-status=live}}</ref><ref>{{cite journal|pmid= 6303898|pmc=1202041| volume=303 |issue =5918|pages=633–635|title=Transposable elements as mutator genes in evolution|journal= Nature | |||
| |vauthors=Chao L, Vargas C, Spear BB, Cox EC|year= 1983| doi=10.1038/303633a0 |bibcode=1983Natur.303..633C| url=}}</ref> In a study on the effects of long-term exposure of simulated microgravity on non-pathogenic ''E. coli'', the results showed transposon insertions occur at loci, linked to SOS stress response.<ref>{{cite journal | vauthors = Tirumalai MR, Karouia F, Tran Q, Stepanov VG, Bruce RJ, Ott M, Pierson DL, Fox GE| title = The adaptation of ''Escherichia coli'' cells grown in simulated microgravity for an extended period is both phenotypic and genomic.| journal = npj Microgravity | volume =3 |issue= 15| date = May 2017 | page = 15| pmid = 28649637 | pmc = 5460176 | doi = 10.1038/s41526-017-0020-1}}</ref> When the same ''E. coli'' strain was exposed to a combination of simulated microgravity and trace (background) levels of (the broad spectrum) antibiotic (]), the results showed transposon-mediated rearrangements (TMRs), disrupting genes involved in bacterial adhesion, and deleting an entire segment of several genes involved with motility and ].<ref>{{cite journal | vauthors = Tirumalai MR, Karouia F, Tran Q, Stepanov VG, Bruce RJ, Ott M, Pierson DL, Fox GE| title = Evaluation of acquired antibiotic resistance in ''Escherichia coli'' exposed to long-term low-shear modeled microgravity and background antibiotic exposure| journal = mBio | volume =10 |issue= e02637-18| date = January 2019 | pmid = 30647159 | pmc = 6336426 | doi = 10.1128/mBio.02637-18}}</ref> Both these studies have implications for microbial growth, adaptation to and antibiotic resistance in real time space conditions. | |||
| Horizontal gene transfer is particularly active in bacterial genomes around the production of secondary or specialized metabolites.<ref name="Ginolhac_2005">{{cite journal | vauthors = Ginolhac A, Jarrin C, Robe P, Perrière G, Vogel TM, Simonet P, Nalin R | title = Type I polyketide synthases may have evolved through horizontal gene transfer | journal = Journal of Molecular Evolution | volume = 60 | issue = 6 | pages = 716–25 | date = June 2005 | pmid = 15909225 | doi = 10.1007/s00239-004-0161-1 | bibcode = 2005JMolE..60..716G }}</ref> This is clearly exhibited within certain groups of bacteria including ''P. aeruginosa'' and ''actinomycetales'', an order of ''Actinomycetota.''<ref>{{Cite journal |last1=Jagannathan |first1=Sveta V. |last2=Manemann |first2=Erika M. |last3=Rowe |first3=Sarah E. |last4=Callender |first4=Maiya C. |last5=Soto |first5=William |date=July 2021 |title=Marine Actinomycetes, New Sources of Biotechnological Products |journal=Marine Drugs |language=en |volume=19 |issue=7 |pages=365 |doi=10.3390/md19070365 |doi-access=free |issn=1660-3397 |pmc=8304352 |pmid=34201951}}</ref> {{citation needed span |text=]s (PKSs) and ] provide modular organizations of associated genes making these bacteria well-adapted to acquire and discard helpful modular modifications via HGT. |date=February 2023}} Certain areas of genes known as ] further increase the likelihood of horizontally transferred secondary metabolite-producing genes.<ref name="Gross_2009">{{cite journal | vauthors = Gross H, Loper JE | title = Genomics of secondary metabolite production by Pseudomonas spp | journal = Natural Product Reports | volume = 26 | issue = 11 | pages = 1408–46 | date = November 2009 | pmid = 19844639 | doi = 10.1039/b817075b }}</ref> {{citation needed span |text=The promiscuity of enzymes is a reoccurring theme in this particular theatre. |date=February 2023}} | |||
| ===Bacterial transformation=== | |||
| ] | |||
| ] is a bacterial adaptation for DNA transfer (HGT) that depends on the expression of numerous bacterial genes whose products are responsible for this process.<ref name="Chen_2004">{{cite journal | vauthors = Chen I, Dubnau D | title = DNA uptake during bacterial transformation | journal = Nature Reviews. Microbiology | volume = 2 | issue = 3 | pages = 241–9 | date = March 2004 | pmid = 15083159 | doi = 10.1038/nrmicro844 | s2cid = 205499369 }}</ref><ref name="Johnsborg_2007">{{cite journal | vauthors = Johnsborg O, Eldholm V, Håvarstein LS | title = Natural genetic transformation: prevalence, mechanisms and function | journal = Research in Microbiology | volume = 158 | issue = 10 | pages = 767–78 | date = December 2007 | pmid = 17997281 | doi = 10.1016/j.resmic.2007.09.004 | doi-access = free }}</ref> In general, transformation is a complex, energy-requiring developmental process. In order for a bacterium to bind, take up and recombine exogenous DNA into its chromosome, it must become ], that is, enter a special physiological state. Competence development in '']'' requires expression of about 40 genes.<ref name="Solomon_1996">{{cite journal | vauthors = Solomon JM, Grossman AD | title = Who's competent and when: regulation of natural genetic competence in bacteria | journal = Trends in Genetics | volume = 12 | issue = 4 | pages = 150–5 | date = April 1996 | pmid = 8901420 | doi = 10.1016/0168-9525(96)10014-7 }}</ref> The DNA integrated into the host chromosome is usually (but with infrequent exceptions) derived from another bacterium of the same ], and is thus homologous to the resident chromosome. The capacity for natural transformation occurs in at least 67 prokaryotic species.<ref name="Johnsborg_2007"/> | |||
| ] for transformation is typically induced by high cell density and/or nutritional limitation, conditions associated with the ] of bacterial growth. Competence appears to be an adaptation for DNA repair.<ref>{{cite journal | vauthors = Michod RE, Bernstein H, Nedelcu AM | title = Adaptive value of sex in microbial pathogens | journal = Infection, Genetics and Evolution | volume = 8 | issue = 3 | pages = 267–85 | date = May 2008 | pmid = 18295550 | doi = 10.1016/j.meegid.2008.01.002 | bibcode = 2008InfGE...8..267M | url = http://www.hummingbirds.arizona.edu/Faculty/Michod/Downloads/IGE%20review%20sex.pdf | access-date = 2016-10-04 | archive-date = 2020-05-11 | archive-url = https://web.archive.org/web/20200511153411/http://www.hummingbirds.arizona.edu/Faculty/Michod/Downloads/IGE%20review%20sex.pdf | url-status = live }}</ref> Transformation in bacteria can be viewed as a primitive sexual process, since it involves interaction of homologous DNA from two individuals to form recombinant DNA that is passed on to succeeding generations. Although transduction is the form of HGT most commonly associated with ]s, certain phages may also be able to promote transformation.<ref name="Keen_2017">{{cite journal | vauthors = Keen EC, Bliskovsky VV, Malagon F, Baker JD, Prince JS, Klaus JS, Adhya SL | title = Novel "Superspreader" Bacteriophages Promote Horizontal Gene Transfer by Transformation | journal = mBio | volume = 8 | issue = 1 | pages = e02115-16 | date = January 2017 | pmid = 28096488 | pmc = 5241400 | doi = 10.1128/mBio.02115-16 }}</ref> | |||
| ===Bacterial conjugation=== | |||
| ] | |||
| ] | |||
| As mentioned before, ] is a method of horizontal gene transfer through cell to cell contact.<ref name="Stearns_2005" /> Through the process of conjugation, ] (T4SS) are used to passage on DNA from the donor cell to the recipient cell.<ref>{{cite journal | url=https://doi.org/10.1128/jb.00438-22 | doi=10.1128/jb.00438-22 | title=Conjugation's Toolkit: The Roles of Nonstructural Proteins in Bacterial Sex | date=2023 | journal=Journal of Bacteriology | volume=205 | issue=3 | pages=e0043822 | pmid=36847532 | pmc=10029717 | vauthors = Cooke MB, Herman C }}</ref> These T4SS encoded within the plasmid carry other proteins and genes that help supplement the cell in conjugation. Research has shown that there are Single Binding DNA Binding proteins (SSBs) also encoded within the conjugative plasmid may help with conjugation and cell viability.<ref>{{Cite journal |last1=Porter |first1=R D |last2=Black |first2=S |date=April 1991 |title=The single-stranded-DNA-binding protein encoded by the Escherichia coli F factor can complement a deletion of the chromosomal ssb gene |journal=Journal of Bacteriology |language=en |volume=173 |issue=8 |pages=2720–2723 |doi=10.1128/jb.173.8.2720-2723.1991 |issn=0021-9193 |pmc=207845 |pmid=2013585}}</ref> This is thought to be the case because SSBs naturally are expressed to help with stabilizing single-stranded DNA (ssDNA).<ref>{{Cite journal |last1=Maffeo |first1=Christopher |last2=Aksimentiev |first2=Aleksei |date=2017-12-01 |title=Molecular mechanism of DNA association with single-stranded DNA binding protein |url=http://academic.oup.com/nar/article/45/21/12125/4559114 |journal=Nucleic Acids Research |language=en |volume=45 |issue=21 |pages=12125–12139 |doi=10.1093/nar/gkx917 |pmid=29059392 |pmc=5716091 |issn=0305-1048}}</ref> SSBs will also recruit other proteins like RadD or RecA expressed in events of DNA recombination, repair, and replication.<ref>{{Cite journal |last1=Gupta |first1=Sankalp |last2=Yeeles |first2=Joseph T.P. |last3=Marians |first3=Kenneth J. |date=September 2014 |title=Regression of Replication Forks Stalled by Leading-strand Template Damage |journal=Journal of Biological Chemistry |language=en |volume=289 |issue=41 |pages=28388–28398 |doi=10.1074/jbc.M114.587907|doi-access=free |pmid=25138217 |pmc=4192491 }}</ref><ref>{{Cite journal |last1=Chen |first1=Stefanie H. |last2=Byrne-Nash |first2=Rose T. |last3=Cox |first3=Michael M. |date=September 2016 |title=Escherichia coli RadD Protein Functionally Interacts with the Single-stranded DNA-binding Protein |journal=Journal of Biological Chemistry |language=en |volume=291 |issue=39 |pages=20779–20786 |doi=10.1074/jbc.M116.736223 |doi-access=free |pmc=5034066 |pmid=27519413}}</ref> Further showcasing their possible role in conjugation. Although it may help, studies have also shown for proteins like SSB to not be essential in conjugation. For example, the plasmid pCF10 from ''Enterococcus faecalis'', a gram-positive bacterium, has a SSB like-protein called PrgE and was classified for not being required for conjugation.<ref>{{Cite journal |last1=Breidenstein |first1=Annika |last2=Lamy |first2=Anaïs |last3=Bader |first3=Cyrielle PJ |last4=Sun |first4=Wei-Sheng |last5=Wanrooij |first5=Paulina H |last6=Berntsson |first6=Ronnie P-A |date=August 2024 |title=PrgE: an OB-fold protein from plasmid pCF10 with striking differences to prototypical bacterial SSBs |journal=Life Science Alliance |language=en |volume=7 |issue=8 |pages=e202402693 |doi=10.26508/lsa.202402693 |issn=2575-1077 |pmc=11137577 |pmid=38811160}}</ref> More work needs to be done on why proteins that bind to ssDNA are encoded into conjugative plasmids. | |||
| Conjugation in the case of microbiomes and symbioses is very important. From this process new genes are acquired that lead to increasing genetic diversity and evolution such as the acquisition of antibiotic resistance genes. ''Mycobacterium tuberculosis'' is a species that has evolved through methods like conjugation while gaining antibiotic resistance.<ref>{{Cite journal |last1=Parsons |first1=Linda M. |last2=Jankowski |first2=Craig S. |last3=Derbyshire |first3=Keith M. |date=April 1998 |title=Conjugal transfer of chromosomal DNA in Mycobacterium smegmatis |url=https://onlinelibrary.wiley.com/doi/10.1046/j.1365-2958.1998.00818.x |journal=Molecular Microbiology |language=en |volume=28 |issue=3 |pages=571–582 |doi=10.1046/j.1365-2958.1998.00818.x |pmid=9632259 |issn=0950-382X}}</ref><ref>{{Cite journal |last1=Supply |first1=Philip |last2=Marceau |first2=Michael |last3=Mangenot |first3=Sophie |last4=Roche |first4=David |last5=Rouanet |first5=Carine |last6=Khanna |first6=Varun |last7=Majlessi |first7=Laleh |last8=Criscuolo |first8=Alexis |last9=Tap |first9=Julien |last10=Pawlik |first10=Alexandre |last11=Fiette |first11=Laurence |last12=Orgeur |first12=Mickael |last13=Fabre |first13=Michel |last14=Parmentier |first14=Cécile |last15=Frigui |first15=Wafa |date=February 2013 |title=Genomic analysis of smooth tubercle bacilli provides insights into ancestry and pathoadaptation of Mycobacterium tuberculosis |journal=Nature Genetics |language=en |volume=45 |issue=2 |pages=172–179 |doi=10.1038/ng.2517 |pmid=23291586 |pmc=3856870 |issn=1061-4036}}</ref> This evolution or increase in genetic diversity is also seen in many other species.<ref>{{Cite journal |last1=Palmer |first1=Kelli L |last2=Kos |first2=Veronica N |last3=Gilmore |first3=Michael S |date=2010-10-01 |title=Horizontal gene transfer and the genomics of enterococcal antibiotic resistance |journal=Current Opinion in Microbiology |series=Antimicrobials/Genomics |volume=13 |issue=5 |pages=632–639 |doi=10.1016/j.mib.2010.08.004 |issn=1369-5274 |pmc=2955785 |pmid=20837397}}</ref> Due to this, there is a huge concern on how impactful conjugation or horizontal gene transfer can be on human health and your microbiome as pathogenic microbes can become more pathogenic. Studies have shown that even our own microbiome has a plethora of antimicrobial genes which if transferred to pathogenic microbes could be detrimental.<ref>{{Cite journal |last1=Sommer |first1=Morten O. A. |last2=Dantas |first2=Gautam |last3=Church |first3=George M. |date=2009-08-28 |title=Functional Characterization of the Antibiotic Resistance Reservoir in the Human Microflora |journal=Science |language=en |volume=325 |issue=5944 |pages=1128–1131 |doi=10.1126/science.1176950 |issn=0036-8075 |pmc=4720503 |pmid=19713526|bibcode=2009Sci...325.1128S }}</ref> | |||
| ] in '']'', like conjugation in '']'', requires stable and extended contact between a donor and a recipient strain, is ], and the transferred DNA is incorporated into the recipient chromosome by ]. However, unlike ''E. coli'' ] (Hfr), mycobacterial conjugation is a type of HGT that is chromosome rather than plasmid based.<ref name="Gray_2013">{{cite journal | vauthors = Gray TA, Krywy JA, Harold J, Palumbo MJ, Derbyshire KM | title = Distributive conjugal transfer in mycobacteria generates progeny with meiotic-like genome-wide mosaicism, allowing mapping of a mating identity locus | journal = PLOS Biology | volume = 11 | issue = 7 | pages = e1001602 | date = July 2013 | pmid = 23874149 | pmc = 3706393 | doi = 10.1371/journal.pbio.1001602 | doi-access = free }}</ref> Furthermore, in contrast to ''E. coli'' (Hfr) conjugation, in ''M. smegmatis'' all regions of the chromosome are transferred with comparable efficiencies. Substantial blending of the parental genomes was found as a result of conjugation, and this blending was regarded as reminiscent of that seen in the meiotic products of sexual reproduction.<ref name="Gray_2013" /><ref name="Derbyshire_2014">{{cite journal | vauthors = Derbyshire KM, Gray TA | title = Distributive Conjugal Transfer: New Insights into Horizontal Gene Transfer and Genetic Exchange in Mycobacteria | journal = Microbiology Spectrum | volume = 2 | issue = 1 | pages = 61–79 | year = 2014 | pmid = 25505644 | pmc = 4259119 | doi = 10.1128/microbiolspec.MGM2-0022-2013 }}</ref> | |||
| ===Archaeal DNA transfer=== | |||
| ] are aerobic ]s thought to have evolved from anaerobic ]s. A large amount of their genome, 126 composite gene families, are derived from genetic material from bacterial genomes. This has allowed them to adapt to extremely salty environments.<ref>{{cite journal | vauthors = Méheust R, Watson AK, Lapointe FJ, Papke RT, Lopez P, Bapteste E | title = Hundreds of novel composite genes and chimeric genes with bacterial origins contributed to haloarchaeal evolution | journal = Genome Biology | volume = 19 | issue = 1 | page = 75 | date = June 2018 | pmid = 29880023 | pmc = 5992828 | doi = 10.1186/s13059-018-1454-9 | doi-access = free | bibcode = 2018GenBi..19...75M }}</ref><ref>{{cite journal | vauthors = Martijn J, Schön ME, Lind AE, Vosseberg J, Williams TA, Spang A, Ettema TJ | title = Hikarchaeia demonstrate an intermediate stage in the methanogen-to-halophile transition | journal = Nature Communications | volume = 11 | issue = 1 | page = 5490 | date = October 2020 | pmid = 33127909 | doi = 10.1038/s41467-020-19200-2 | pmc = 7599335 | bibcode = 2020NatCo..11.5490M }}</ref> | |||
| The ] '']'', when ] irradiated, strongly induces the formation of ] which then facilitates cellular aggregation.<ref name="Fröls_2008">{{cite journal | vauthors = Fröls S, Ajon M, Wagner M, Teichmann D, Zolghadr B, Folea M, Boekema EJ, Driessen AJ, Schleper C, Albers SV | title = UV-inducible cellular aggregation of the hyperthermophilic archaeon Sulfolobus solfataricus is mediated by pili formation | journal = Molecular Microbiology | volume = 70 | issue = 4 | pages = 938–52 | date = November 2008 | pmid = 18990182 | doi = 10.1111/j.1365-2958.2008.06459.x | url = https://pure.rug.nl/ws/files/56956856/UV_inducible_cellular_aggregation_of_the_hyperthermophilic_archaeon_Sulfolobus_solfataricus_is_mediated_by_pili_formation.pdf | doi-access = free | access-date = 2020-06-23 | archive-date = 2023-04-15 | archive-url = https://web.archive.org/web/20230415071308/https://pure.rug.nl/ws/files/56956856/UV_inducible_cellular_aggregation_of_the_hyperthermophilic_archaeon_Sulfolobus_solfataricus_is_mediated_by_pili_formation.pdf | url-status = live }}</ref><ref name="Allers_2011">{{cite journal | vauthors = Allers T | title = Swapping genes to survive - a new role for archaeal type IV pili | journal = Molecular Microbiology | volume = 82 | issue = 4 | pages = 789–91 | date = November 2011 | pmid = 21992544 | doi = 10.1111/j.1365-2958.2011.07860.x | s2cid = 45490029 | doi-access = }}</ref> Exposure to chemical agents that cause DNA damage also induces cellular aggregation.<ref name="Fröls_2008"/> Other physical stressors, such as temperature shift or pH, do not induce aggregation, suggesting that DNA damage is a specific inducer of cellular aggregation.{{citation needed|date=February 2023}} | |||
| UV-induced cellular aggregation mediates intercellular chromosomal HGT marker exchange with high frequency,<ref name="Ajon_2011">{{cite journal | vauthors = Ajon M, Fröls S, van Wolferen M, Stoecker K, Teichmann D, Driessen AJ, Grogan DW, Albers SV, Schleper C | title = UV-inducible DNA exchange in hyperthermophilic archaea mediated by type IV pili | journal = Molecular Microbiology | volume = 82 | issue = 4 | pages = 807–17 | date = November 2011 | pmid = 21999488 | doi = 10.1111/j.1365-2958.2011.07861.x | url = https://pure.rug.nl/ws/files/6771142/2011MolMicrobiolAjon.pdf | doi-access = free | access-date = 2020-06-23 | archive-date = 2021-10-10 | archive-url = https://web.archive.org/web/20211010101112/https://pure.rug.nl/ws/files/6771142/2011MolMicrobiolAjon.pdf | url-status = live }}</ref> and UV-induced cultures display recombination rates that exceed those of uninduced cultures by as much as three orders of magnitude. ''S. solfataricus'' cells aggregate preferentially with other cells of their own species.<ref name="Ajon_2011"/> Frols et al.<ref name="Fröls_2008"/><ref name="pmid19143598">{{cite journal | vauthors = Fröls S, White MF, Schleper C | title = Reactions to UV damage in the model archaeon Sulfolobus solfataricus | journal = Biochemical Society Transactions | volume = 37 | issue = Pt 1 | pages = 36–41 | date = February 2009 | pmid = 19143598 | doi = 10.1042/BST0370036 }}</ref> and Ajon et al.<ref name="Ajon_2011"/> suggested that UV-inducible DNA transfer is likely an important mechanism for providing increased repair of damaged DNA via homologous recombination. This process can be regarded as a simple form of sexual interaction. | |||
| Another ], '']'', is able to undergo HGT. ''S. acidocaldarius'' can exchange and recombine chromosomal markers at temperatures up to 84 °C.<ref name="Grogan_1996">{{cite journal | vauthors = Grogan DW | title = Exchange of genetic markers at extremely high temperatures in the archaeon Sulfolobus acidocaldarius | journal = Journal of Bacteriology | volume = 178 | issue = 11 | pages = 3207–11 | date = June 1996 | pmid = 8655500 | pmc = 178072 | doi = 10.1128/jb.178.11.3207-3211.1996 }}</ref> UV exposure induces pili formation and cellular aggregation.<ref name="Ajon_2011"/> Cells with the ability to aggregate have greater survival than mutants lacking pili that are unable to aggregate. The frequency of recombination is increased by DNA damage induced by UV-irradiation<ref name="Wood_1997">{{cite journal | vauthors = Wood ER, Ghané F, Grogan DW | title = Genetic responses of the thermophilic archaeon Sulfolobus acidocaldarius to short-wavelength UV light | journal = Journal of Bacteriology | volume = 179 | issue = 18 | pages = 5693–8 | date = September 1997 | pmid = 9294423 | pmc = 179455 | doi = 10.1128/jb.179.18.5693-5698.1997 }}</ref> and by DNA damaging chemicals.<ref name="Reilly_2002">{{cite journal | vauthors = Reilly MS, Grogan DW | title = Biological effects of DNA damage in the hyperthermophilic archaeon Sulfolobus acidocaldarius | journal = FEMS Microbiology Letters | volume = 208 | issue = 1 | pages = 29–34 | date = February 2002 | pmid = 11934490 | doi = 10.1016/s0378-1097(01)00575-4 | doi-access = free }}</ref> | |||
| The ], containing five genes, is highly induced by UV irradiation. The proteins encoded by the ''ups'' operon are employed in UV-induced pili assembly and cellular aggregation leading to intercellular DNA exchange and ].<ref name="Wolferen_2013">{{cite journal | vauthors = van Wolferen M, Ajon M, Driessen AJ, Albers SV | title = Molecular analysis of the UV-inducible pili operon from Sulfolobus acidocaldarius | journal = MicrobiologyOpen | volume = 2 | issue = 6 | pages = 928–37 | date = December 2013 | pmid = 24106028 | pmc = 3892339 | doi = 10.1002/mbo3.128 }}</ref> Since this system increases the fitness of ''S. acidocaldarius'' cells after UV exposure, Wolferen et al.<ref name="Wolferen_2013"/><ref name="van_Wolferen_2015">{{cite journal | vauthors = van Wolferen M, Ma X, Albers SV | title = DNA Processing Proteins Involved in the UV-Induced Stress Response of Sulfolobales | journal = Journal of Bacteriology | volume = 197 | issue = 18 | pages = 2941–51 | date = September 2015 | pmid = 26148716 | pmc = 4542170 | doi = 10.1128/JB.00344-15 }}</ref> considered that transfer of DNA likely takes place in order to repair UV-induced DNA damages by homologous recombination. | |||
| * ''']''', the genetic alteration of a ] resulting from the introduction, uptake and ] of foreign genetic material (] or ]). This process is relatively common in bacteria, but less common in ]s. Transformation is often used to insert novel genes into bacteria for experiments, or for industrial or medical applications. See also ] and ]. | |||
| * ''']''', the process in which bacterial DNA is moved from one bacterium to another by a bacterial virus (a bacteriophage, commonly called a ]). | |||
| * ''']''', a process in which a living bacterial cell transfers genetic material through cell-to-cell contact. | |||
| ==Eukaryotes== | ==Eukaryotes== | ||
| "Sequence comparisons suggest recent horizontal transfer of many genes among diverse species including across the boundaries of ] 'domains'. Thus determining the phylogenetic history of a species can not be done conclusively by determining evolutionary trees for single genes."<ref>{{cite web |url=http://bioinfosu.okstate.edu/MG/MGW3/MG334.html | vauthors = Melcher U | date = 2001 | title = Molecular genetics: Horizontal gene transfer | publisher = Oklahoma State University | location = Stillwater, Oklahoma USA |access-date=2015-08-20 |archive-url=https://web.archive.org/web/20160304071146/http://bioinfosu.okstate.edu/MG/MGW3/MG334.html |archive-date=2016-03-04 }}</ref> | |||
| Analysis of ]s suggests that horizontal gene transfer has also occurred within ]s, from their chloroplast and mitochondrial genome to their nuclear genome. As stated in the ], chloroplasts and mitochondria probably originated as bacterial ]s of a progenitor to the eukaryotic cell.<ref> Jeffrey L. Blanchard and Michael Lynch (2000), "Organellar genes: why do they end up in the nucleus?", ''Trends in Genetics'', '''16''' (7), pp. 315-320. (Discusses theories on how mitochondria and chloroplast genes are transferred into the nucleus, and also what steps a gene needs to go through in order to complete this process.) </ref> | |||
| === Organelle to nuclear genome === | |||
| Horizontal transfer of genes from bacteria to some ], especially the yeast '']'', has been well documented.<ref> ''Hall C, Brachat S, Dietrich FS. "Contribution of Horizontal Gene Transfer to the Evolution of Saccharomyces cerevisiae." Eukaryot Cell 2005 Jun 4(6):1102-15. '' The article argues that horizontal transfer of bacterial DNA to ''Saccharomyces cerevisiae'' has occurred.</ref> | |||
| *Analysis of ]s suggests that horizontal gene transfer has occurred within eukaryotes from the chloroplast and ]s to the ]. As stated in the ], ]s and ] probably originated as bacterial ]s of a progenitor to the eukaryotic cell.<ref>{{cite journal | vauthors = Blanchard JL, Lynch M | title = Organellar genes: why do they end up in the nucleus? | journal = Trends in Genetics | volume = 16 | issue = 7 | pages = 315–20 | date = July 2000 | pmid = 10858662 | doi = 10.1016/S0168-9525(00)02053-9 }} Discusses theories on how mitochondria and chloroplast genes are transferred into the nucleus, and also what steps a gene needs to go through in order to complete this process.</ref> | |||
| === Organelle to organelle === | |||
| There is also recent evidence that the ] has somehow acquired genetic material from its (non-beneficial) endosymbiont '']''. <ref> Natsuko Kondo, Naruo Nikoh, Nobuyuki Ijichi, Masakazu Shimada and Takema Fukatsu (2002) "Genome fragment of Wolbachia endosymbiont transferred to X chromosome of host insect", Proceedings of the National Academy of Sciences of the USA, 99 (22): 14280-14285". '' (Free full article) This article argues that '']'' DNA is in the ] genome (a species of ].</ref> New examples have recently been reported, demonstrating that Wolbachia bacteria represent an important potential source of genetic material in arthropods and ] ]s. <ref> {{ cite journal | author = Hotopp JC, Clark ME, Oliveira DC, Foster JM, Fischer P, Torres MC, Giebel JD, Kumar N, Ishmael N, Wang S, Ingram J, Nene RV, Shepard J, Tomkins J, Richards S, Spiro DJ, Ghedin E, Slatko BE, Tettelin H, Werren JH | title = Widespread Lateral Gene Transfer from Intracellular Bacteria to Multicellular Eukaryotes | journal = Science | date = 30 Aug 2007 | pmid = 17761848 | doi = 10.1126/science.1142490 }}</ref> | |||
| *]s moved to parasites of the ] plant family from their hosts<ref>{{cite journal | vauthors = Davis CC, Wurdack KJ | s2cid = 16180594 | title = Host-to-parasite gene transfer in flowering plants: phylogenetic evidence from Malpighiales | journal = Science | volume = 305 | issue = 5684 | pages = 676–8 | date = July 2004 | pmid = 15256617 | doi = 10.1126/science.1100671 | bibcode = 2004Sci...305..676D | doi-access = free }}</ref><ref>{{cite journal | vauthors = Nickrent DL, Blarer A, Qiu YL, Vidal-Russell R, Anderson FE | title = Phylogenetic inference in Rafflesiales: the influence of rate heterogeneity and horizontal gene transfer | journal = BMC Evolutionary Biology | volume = 4 | issue = 1 | page = 40 | date = October 2004 | pmid = 15496229 | pmc = 528834 | doi = 10.1186/1471-2148-4-40 | author-link1 = Daniel Lee Nickrent | doi-access = free }}</ref> and from chloroplasts of a still-unidentified plant to the mitochondria of the bean '']''.<ref>{{cite journal | vauthors = Woloszynska M, Bocer T, Mackiewicz P, Janska H | title = A fragment of chloroplast DNA was transferred horizontally, probably from non-eudicots, to mitochondrial genome of Phaseolus | journal = Plant Molecular Biology | volume = 56 | issue = 5 | pages = 811–20 | date = November 2004 | pmid = 15803417 | doi = 10.1007/s11103-004-5183-y | s2cid = 14198321 }}</ref> | |||
| === Bacteria to fungi === | |||
| There is also evidence for horizontal transfer of ]s to parasites of the ] plant family from their hosts (also plants),<ref>{{cite journal | journal = Science | date = 30 July 2004 | volume = 305 | issue = 5684 | pages = 676–678 | doi = 10.1126/science.1100671 | title = Host-to-Parasite Gene Transfer in Flowering Plants: Phylogenetic Evidence from Malpighiales | url = http://www.sciencemag.org/cgi/content/abstract/305/5684/676 | author = Charles C. Davis and Kenneth J. Wurdack}}</ref><ref>{{cite journal | title = Phylogenetic inference in Rafflesiales: the influence of rate heterogeneity and horizontal gene transfer | author = Daniel L Nickrent, Albert Blarer, Yin-Long Qiu, Romina Vidal-Russell and Frank E Anderson | journal = BMC Evolutionary Biology | year = 2004 | volume = 4 | issue = 40 | doi = 10.1186/1471-2148-4-40 | url = http://www.biomedcentral.com/1471-2148/4/40 }}</ref> and from ]s of a not-yet-identified plant to the mitochondria of the bean '']''.<ref>{{cite journal | author = Magdalena Woloszynska, Tomasz Bocer, Pawel Mackiewicz and Hanna Janska | title = A fragment of chloroplast DNA was transferred horizontally, probably from non-eudicots, to mitochondrial genome of Phaseolus | journal = Plant Molecular Biology | volume = 56 | issue = 5 | date = November, 2004 | doi = 10.1007/s11103-004-5183-y | pages = 811-820}}</ref> | |||
| *Horizontal transfer occurs from bacteria to some ], such as the yeast '']''.<ref>{{cite journal | vauthors = Hall C, Brachat S, Dietrich FS | title = Contribution of horizontal gene transfer to the evolution of Saccharomyces cerevisiae | journal = Eukaryotic Cell | volume = 4 | issue = 6 | pages = 1102–15 | date = June 2005 | pmid = 15947202 | pmc = 1151995 | doi = 10.1128/EC.4.6.1102-1115.2005 }}</ref> | |||
| === Bacteria to plants=== | |||
| "Sequence comparisons suggest recent horizontal transfer of many ]s among diverse ] including across the boundaries of ] "domains". Thus determining the phylogenetic history of a species can not be done conclusively by determining evolutionary trees for single genes."<ref> </ref> | |||
| * Agrobacterium, a pathogenic bacterium that causes cells to proliferate as crown galls and proliferating roots is an example of a bacterium that can transfer genes to plants and this plays an important role in plant evolution.<ref>{{cite journal | vauthors = Quispe-Huamanquispe DG, Gheysen G, Kreuze JF | title = Agrobacterium T-DNAs | journal = Frontiers in Plant Science | volume = 8 | page = 2015 | year = 2017 | pmid = 29225610 | pmc = 5705623 | doi = 10.3389/fpls.2017.02015 | doi-access = free }}</ref> | |||
| * Land plants and their close relatives, the charophycean green algae, share a set of glycosyl hydrolases. These enzymes were likely transferred from bacteria and fungi to the last common ancestor of these organisms before the origin of land plants.<ref>{{cite journal | vauthors = Kfoury B, Rodrigues WF, Kim SJ, Brandizzi F, Del-Bem LE | title = Multiple horizontal gene transfer events have shaped plant glycosyl hydrolase diversity and function | journal = New Phytologist | volume = 242 | pages = 809–824 | year = 2024 | issue = 2 | doi = 10.1111/nph.19595 | doi-access = free | pmid = 38417454 | bibcode = 2024NewPh.242..809K }}</ref> | |||
| === Bacteria to animals === | |||
| ==Evolutionary theory==<!-- This section is linked from ] --> | |||
| *] is a gene in the genome of the coffee berry borer ('']'') that resembles bacterial genes, and is thought to be transferred from bacteria in the beetle's gut.<ref>{{cite journal |journal=Nature|year=2012|doi=10.1038/nature.2012.10116|title=Bacterial gene helps coffee beetle get its fix| vauthors = Lee Phillips M |s2cid=211729274}}</ref><ref>{{cite journal | vauthors = Acuña R, Padilla BE, Flórez-Ramos CP, Rubio JD, Herrera JC, Benavides P, Lee SJ, Yeats TH, Egan AN, Doyle JJ, Rose JK | title = Adaptive horizontal transfer of a bacterial gene to an invasive insect pest of coffee | journal = Proceedings of the National Academy of Sciences of the United States of America | volume = 109 | issue = 11 | pages = 4197–202 | date = March 2012 | pmid = 22371593 | pmc = 3306691 | doi = 10.1073/pnas.1121190109 | bibcode = 2012PNAS..109.4197A | doi-access = free }}</ref> | |||
| Horizontal gene transfer is a potential ] in inferring ]s based on the ] of one ]. For example, given two distantly related bacteria that have exchanged a gene, a ] including those species will show them to be closely related because that gene is the same, even though most other genes have substantially diverged. For this reason, it is often ideal to use other information to infer robust phylogenies, such as the presence or absence of genes, or, more commonly, to include as wide a range of genes for phylogenetic analysis as possible. | |||
| *] is an essential gene for the specification of the germline in ] and its origin is through to be due to a HGT event followed by a fusion with a LOTUS domain.<ref>{{cite journal | vauthors = Blondel L, Jones ET, Extavour GC | title = Bacterial contribution to genesis of the novel germ line determinant oskar | journal = eLife | volume = 24 | issue = 9 | pages = e45539 | date = Feb 2020 | pmid = 32091394 | doi = 10.7554/eLife.45539 | pmc = 7250577 | doi-access = free }}</ref> | |||
| *]s currently hold the 'record' for HGT in animals with ~8% of their genes from bacterial origins.<ref>{{cite web|url=https://www.science.org/content/article/bdelloids-surviving-borrowed-dna|title=Bdelloids Surviving on Borrowed DNA| vauthors = Watson T |date=15 November 2012|publisher=Science/AAAS News|access-date=30 June 2022|archive-date=6 May 2023|archive-url=https://web.archive.org/web/20230506013639/https://www.science.org/content/article/bdelloids-surviving-borrowed-dna|url-status=live}}</ref> ]s were thought to break the record with 17.5% HGT, but that finding was an artifact of bacterial contamination.<ref name="Koutsovoulos_2016">{{cite journal | vauthors = Koutsovoulos G, Kumar S, Laetsch DR, Stevens L, Daub J, Conlon C, Maroon H, Thomas F, Aboobaker AA, Blaxter M | title = No evidence for extensive horizontal gene transfer in the genome of the tardigrade Hypsibius dujardini | journal = Proceedings of the National Academy of Sciences of the United States of America | volume = 113 | issue = 18 | pages = 5053–8 | date = May 2016 | pmid = 27035985 | pmc = 4983863 | doi = 10.1073/pnas.1600338113 | bibcode = 2016PNAS..113.5053K | doi-access = free }}</ref> | |||
| *A study found the genomes of 40 animals (including 10 primates, four '']'' worms, and 12 '']'' insects) contained genes which the researchers concluded had been transferred from bacteria and fungi by horizontal gene transfer.<ref>{{cite journal | vauthors = Crisp A, Boschetti C, Perry M, Tunnacliffe A, Micklem G | title = Expression of multiple horizontally acquired genes is a hallmark of both vertebrate and invertebrate genomes | journal = Genome Biology | volume = 16 | page = 50 | date = March 2015 | issue = 1 | pmid = 25785303 | pmc = 4358723 | doi = 10.1186/s13059-015-0607-3 | doi-access = free }}</ref> The researchers estimated that for some nematodes and Drosophila insects these genes had been acquired relatively recently.<ref>{{cite web|url=http://www.the-scientist.com/?articles.view/articleNo/42420/title/Horizontal-Gene-Transfer-a-Hallmark-of-Animal-Genomes-/|title=Horizontal Gene Transfer a Hallmark of Animal Genomes?|vauthors=Madhusoodanan J|date=2015-03-12|website=The Scientist|access-date=2016-07-14|archive-date=2016-07-09|archive-url=https://web.archive.org/web/20160709144907/http://www.the-scientist.com//?articles.view/articleNo/42420/title/Horizontal-Gene-Transfer-a-Hallmark-of-Animal-Genomes-/|url-status=live}}</ref> | |||
| *A bacteriophage-mediated mechanism transfers genes between prokaryotes and eukaryotes.<ref>{{cite journal | vauthors = Daugavet MA, Shabelnikov S, Shumeev A, Shaposhnikova T, Adonin LS, Podgornaya O | title = Features of a novel protein, rusticalin, from the ascidian ''Styela rustica'' reveal ancestral horizontal gene transfer event | journal = Mobile DNA | volume = 10 | issue = 1 | page = 4 | date = 2019-01-19 | pmid = 30675192 | pmc = 6339383 | doi = 10.1186/s13100-019-0146-7 | doi-access = free }}</ref> Nuclear localization signals in bacteriophage terminal proteins (TP) prime DNA replication and become covalently linked to the viral genome. The role of virus and bacteriophages in HGT in bacteria, suggests that TP-containing genomes could be a vehicle of inter-kingdom genetic information transference all throughout evolution.<ref>{{cite journal | vauthors = Redrejo-Rodríguez M, Muñoz-Espín D, Holguera I, Mencía M, Salas M | title = Functional eukaryotic nuclear localization signals are widespread in terminal proteins of bacteriophages | journal = Proceedings of the National Academy of Sciences of the United States of America | volume = 109 | issue = 45 | pages = 18482–7 | date = November 2012 | pmid = 23091024 | pmc = 3494942 | doi = 10.1073/pnas.1216635109 | bibcode = 2012PNAS..10918482R | doi-access = free }}</ref> | |||
| *The ] has acquired genetic material from its (non-beneficial) endosymbiont '']''.<ref>{{cite journal | vauthors = Kondo N, Nikoh N, Ijichi N, Shimada M, Fukatsu T | title = Genome fragment of Wolbachia endosymbiont transferred to X chromosome of host insect | journal = Proceedings of the National Academy of Sciences of the United States of America | volume = 99 | issue = 22 | pages = 14280–5 | date = October 2002 | pmid = 12386340 | pmc = 137875 | doi = 10.1073/pnas.222228199 | bibcode = 2002PNAS...9914280K | doi-access = free }}</ref> New examples have recently been reported demonstrating that Wolbachia bacteria represent an important potential source of genetic material in arthropods and ] ]s.<ref>{{cite journal | vauthors = Dunning Hotopp JC, Clark ME, Oliveira DC, Foster JM, Fischer P, Muñoz Torres MC, Giebel JD, Kumar N, Ishmael N, Wang S, Ingram J, Nene RV, Shepard J, Tomkins J, Richards S, Spiro DJ, Ghedin E, Slatko BE, Tettelin H, Werren JH | s2cid = 10787254 | title = Widespread lateral gene transfer from intracellular bacteria to multicellular eukaryotes | journal = Science | volume = 317 | issue = 5845 | pages = 1753–6 | date = September 2007 | pmid = 17761848 | doi = 10.1126/science.1142490 | bibcode = 2007Sci...317.1753H }}</ref> | |||
| *The psyllid ''Pachypsylla venusta'' has acquired genes from its current endosymbiont ''Carsonella'', and from many of its historical endosymbionts, too.<ref>Sloan, D. B., Nakabachi, A., Richards, S., Qu, J., Murali, S. C., Gibbs, R. A., & Moran, N. A. (2014). Parallel histories of horizontal gene transfer facilitated extreme reduction of endosymbiont genomes in sap-feeding insects. Molecular biology and evolution, 31(4), 857-871.</ref> | |||
| === Plant to plant === | |||
| For example, the most common gene to be used for constructing phylogenetic relationships in ]s is the ] gene, since its sequences tend to be conserved among members with close phylogenetic distances, but variable enough that differences can be measured. However, in recent years it has also been argued that 16s rRNA genes can also be horizontally transferred. Although this may be infrequent, validity of 16s rRNA-constructed phylogenetic trees must be reevaluated. | |||
| *'']'', a ] ], has received a gene from ] ('']'') to its nuclear genome.<ref name="Yoshida_2010">{{cite journal | vauthors = Yoshida S, Maruyama S, Nozaki H, Shirasu K | s2cid = 39376164 | title = Horizontal gene transfer by the parasitic plant Striga hermonthica | journal = Science | volume = 328 | issue = 5982 | page = 1128 | date = May 2010 | pmid = 20508124 | doi = 10.1126/science.1187145 | bibcode = 2010Sci...328.1128Y }}</ref> The gene's functionality is unknown. | |||
| *A gene that allowed ferns to survive in dark forests came from the ], which grows in mats on streambanks or trees. The neochrome gene arrived about 180 million years ago.<ref>{{cite news|title=Plants That Practice Genetic Engineering|date=April 17, 2014| vauthors = Zimmer C |work=New York Times|url=https://www.nytimes.com/2014/04/17/science/plants-that-practice-genetic-engineering.html|access-date=February 27, 2017|archive-date=December 26, 2022|archive-url=https://web.archive.org/web/20221226055131/https://www.nytimes.com/2014/04/17/science/plants-that-practice-genetic-engineering.html|url-status=live}}</ref> | |||
| *Transfer of mRNA between host plants and heterotrophs plants in the ] have been directly observed. mRNA transcripts can therefore be a factor involved in the transfer and integration of foreign DNA in heterotrophs.<ref>David‐Schwartz R, Runo S, Townsley B, Machuka J, Sinha N. 2008. Long‐distance transport | |||
| of mRNA via parenchyma cells and phloem across the host–parasite junction in | |||
| Cuscuta. New Phytologist 179: 1133– 1141.</ref> | |||
| === Plants to animals === | |||
| Biologist Gogarten suggests "the original metaphor of a tree no longer fits the data from recent genome research" therefore "biologists should use the metaphor of a mosaic to describe the different histories combined in individual genomes and use the metaphor of a net to visualize the rich exchange and cooperative effects of HGT among microbes."<ref name="gogarten" /> | |||
| *The eastern emerald sea slug '']'' has been suggested by ] (FISH) analysis to contain photosynthesis-supporting genes obtained from an algae (''])'' in their diet.<ref>{{cite journal | vauthors = Schwartz JA, Curtis NE, Pierce SK | title = FISH labeling reveals a horizontally transferred algal (Vaucheria litorea) nuclear gene on a sea slug (Elysia chlorotica) chromosome | journal = The Biological Bulletin | volume = 227 | issue = 3 | pages = 300–12 | date = December 2014 | pmid = 25572217 | doi = 10.1086/BBLv227n3p300 | s2cid = 21742354 }}</ref> LGT in Sacoglossa is now thought to be an artifact<ref>{{cite journal | vauthors = Rauch C, Vries J, Rommel S, Rose LE, Woehle C, Christa G, Laetz EM, Wägele H, Tielens AG, Nickelsen J, Schumann T, Jahns P, Gould SB | title = Why It Is Time to Look Beyond Algal Genes in Photosynthetic Slugs | journal = Genome Biology and Evolution | volume = 7 | issue = 9 | pages = 2602–7 | date = August 2015 | pmid = 26319575 | pmc = 4607529 | doi = 10.1093/gbe/evv173 }}</ref> and no trace of LGT was found upon sequencing the genome of '']''.<ref>{{cite journal | vauthors = Bhattacharya D, Pelletreau KN, Price DC, Sarver KE, Rumpho ME | title = Genome analysis of Elysia chlorotica Egg DNA provides no evidence for horizontal gene transfer into the germ line of this Kleptoplastic Mollusc | journal = Molecular Biology and Evolution | volume = 30 | issue = 8 | pages = 1843–52 | date = August 2013 | pmid = 23645554 | pmc = 3708498 | doi = 10.1093/molbev/mst084 }}</ref> | |||
| *The whitefly '']'' acquired a plant detoxification gene that neutralizes plant toxins.<ref>{{cite journal | vauthors = Xia J, Guo Z, Yang Z, Han H, Wang S, Xu H, Yang X, Yang F, Wu Q, Xie W, Zhou X, Dermauw W, Turlings TC, Zhang Y | title = Whitefly hijacks a plant detoxification gene that neutralizes plant toxins | journal = Cell | volume = 184 | issue = 7 | pages = 1693–1705.e17 | date = April 2021 | pmid = 33770502 | doi = 10.1016/j.cell.2021.02.014 | s2cid = 232359463 | doi-access = free }}</ref> | |||
| === Plant to fungus === | |||
| Using single ]s as ]s, it is difficult to trace organismal ] in the presence of horizontal gene transfer. Combining the simple ] model of ] with rare HGT horizontal gene transfer events suggest there was no single ] that contained all of the genes ancestral to those shared among the three domains of ]. Each contemporary ] has its own history and traces back to an individual molecule ]. However, these molecular ancestors were likely to be present in different organisms at different times."<ref> </ref> | |||
| {{Main|Plant-fungus horizontal gene transfer}} | |||
| * Gene transfer between plants and fungi has been posited for a number of cases, including rice ('']'').{{citation needed|date=February 2023}} | |||
| * Evidence of gene transfer from plants was documented in the fungus ''Colletotrichum.''<ref name="Armijos_Jaramillo_2013">{{cite journal | vauthors = Armijos Jaramillo VD, Vargas WA, Sukno SA, Thon MR | title = New insights into the evolution and structure of Colletotrichum plant-like subtilisins (CPLSs) | journal = Communicative & Integrative Biology | volume = 6 | issue = 6 | pages = e25727 | date = November 2013 | pmid = 24563701| pmc = 3917961 | doi = 10.4161/cib.25727 }}</ref> | |||
| * Plant expansin genes were transferred to fungi further enabling the fungi to infect plants.<ref name="Nikolaidis_2014">{{cite journal | vauthors = Nikolaidis N, Doran N, Cosgrove DJ | title = Plant expansins in bacteria and fungi: evolution by horizontal gene transfer and independent domain fusion | journal = Molecular Biology and Evolution | volume = 31 | issue = 2 | pages = 376–86 | date = February 2014 | pmid = 24150040 | doi = 10.1093/molbev/mst206 }}</ref> | |||
| === Plant to bacteria === | |||
| ''Uprooting the Tree of Life'' by W. ] ('']'', February 2000, pp 72-77)<ref>{{cite journal | author = ] | year = February 2000 | title = Uprooting the Tree of Life | journal = ] | pages = pp. 72-77 }}</ref> contains a discussion of the Last Universal Common Ancestor, and the problems that arose with respect to that concept when one considers horizontal gene transfer. The article covers a wide area - the ] hypothesis for ]s, the use of small subunit ribosomal ] (SSU rRNA) as a measure of evolutionary distances (this was the field ] worked in when formulating the first modern "tree of life", and his research results with SSU rRNA led him to propose the ] as a third domain of ]) and other relevant topics. Indeed, it was while examining the new three-domain view of life that horizontal gene transfer arose as a complicating issue: ''Archaeoglobus fulgidus'' is cited in the article (p.76) as being an anomaly with respect to a ] tree based upon the encoding for the ] ] - the organism in question is a definite Archaean, with all the cell lipids and transcription machinery that are expected of an Archaean, but whose HMGCoA genes are actually of bacterial origin.<ref name = "Scientific American">''Uprooting the Tree of Life'' by W. ] ('']'', February 2000, pp 72-77)</ref> | |||
| * Plant expansin genes were transferred to bacteria further enabling the bacteria to infect plants.<ref name="Nikolaidis_2014"/> | |||
| === Fungi to insects === | |||
| Again on p.76, the article continues with: | |||
| *Pea aphids ('']'') contain multiple genes from ].<ref name="Moran_2010" /><ref>{{cite journal | vauthors = Fukatsu T | s2cid = 23686682 | title = Evolution. A fungal past to insect color | journal = Science | volume = 328 | issue = 5978 | pages = 574–5 | date = April 2010 | pmid = 20431000 | doi = 10.1126/science.1190417 | bibcode = 2010Sci...328..574F }}</ref> Plants, fungi, and microorganisms can synthesize ]s, but ] made by pea ]s is the only carotenoid known to be synthesized by an organism in the animal kingdom.<ref name="Moran_2010">{{cite journal | vauthors = Moran NA, Jarvik T | s2cid = 14785276 | title = Lateral transfer of genes from fungi underlies carotenoid production in aphids | journal = Science | volume = 328 | issue = 5978 | pages = 624–7 | date = April 2010 | pmid = 20431015 | doi = 10.1126/science.1187113 | bibcode = 2010Sci...328..624M }}</ref> | |||
| === Fungi to fungi === | |||
| : "The weight of evidence still supports the likelihood that ] in ]s derived from ]l cells and that ]s came from ingested ], but it is no longer safe to assume that those were the only lateral gene transfers that occurred after the first eukaryotes arose. Only in later, multicellular eukaryotes do we know of definite restrictions on horizontal gene exchange, such as the advent of separated (and protected) ]s."<ref name = "Scientific American"/> | |||
| * The toxin ] is found in numerous, seemingly unrelated genera fungi such as '']'', '']'', and '']''. Two biosynthetic genes involved in the production of α-amanitin are P450-29 and FMO1. Phylogenetic and genetic analyses of these genes strongly indicate that they were transferred between the genera via horizontal gene transfer.<ref>{{cite journal | vauthors = Luo H, Hallen-Adams HE, Lüli Y, Sgambelluri RM, Li X, Smith M, Yang ZL, Martin FM | title = Genes and evolutionary fates of the amanitin biosynthesis pathway in poisonous mushrooms | journal = Proceedings of the National Academy of Sciences of the United States of America | volume = 119 | issue = 20 | pages = e2201113119 | date = May 2022 | pmid = 35533275 | doi = 10.1073/pnas.2201113119 | doi-access = free | pmc = 9171917 | bibcode = 2022PNAS..11901113L | s2cid = 248668772 }}</ref> | |||
| The article continues with: | |||
| * The ToxA protein (wheat virulence protein) included in a ∼14 kb element, containing both coding and non-coding regions was transferred between different fungal wheat patogens: ''Parastagonospora nodorum'', ''Pyrenophora tritici-repentis'', and ''Bipolaris sorokiniana''.<ref>{{Cite journal |last1=McDonald |first1=Megan C. |last2=Taranto |first2=Adam P. |last3=Hill |first3=Erin |last4=Schwessinger |first4=Benjamin |last5=Liu |first5=Zhaohui |last6=Simpfendorfer |first6=Steven |last7=Milgate |first7=Andrew |last8=Solomon |first8=Peter S. |date=2019-10-29 |editor-last=Di Pietro |editor-first=Antonio |title=Transposon-Mediated Horizontal Transfer of the Host-Specific Virulence Protein ToxA between Three Fungal Wheat Pathogens |journal=mBio |language=en |volume=10 |issue=5 |doi=10.1128/mBio.01515-19 |pmid=31506307 |pmc=6737239 |issn=2161-2129}}</ref> | |||
| * A large genomic element named "Wallaby," approximately 500 kb in length, was recently transferred between two Penicillium species used in cheesemaking: ''P. camemberti'' and ''P. roqueforti''. Wallaby contains around 250 genes, including several that are thought to play a role in microbial competition.<ref>{{Cite journal |last1=Cheeseman |first1=Kevin |last2=Ropars |first2=Jeanne |last3=Renault |first3=Pierre |last4=Dupont |first4=Joëlle |last5=Gouzy |first5=Jérôme |last6=Branca |first6=Antoine |last7=Abraham |first7=Anne-Laure |last8=Ceppi |first8=Maurizio |last9=Conseiller |first9=Emmanuel |last10=Debuchy |first10=Robert |last11=Malagnac |first11=Fabienne |last12=Goarin |first12=Anne |last13=Silar |first13=Philippe |last14=Lacoste |first14=Sandrine |last15=Sallet |first15=Erika |date=2014-01-10 |title=Multiple recent horizontal transfers of a large genomic region in cheese making fungi |journal=Nature Communications |language=en |volume=5 |issue=1 |page=2876 |doi=10.1038/ncomms3876 |issn=2041-1723 |pmc=3896755 |pmid=24407037|bibcode=2014NatCo...5.2876C }}</ref> | |||
| === Fungi to oomycetes === | |||
| :"If there had never been any lateral gene transfer, all these individual gene trees would have the same topology (the same branching order), and the ancestral genes at the root of each tree would have all been present in the last universal common ancestor, a single ancient cell. But extensive transfer means that neither is the case: gene trees will differ (although many will have regions of similar topology) ''and'' there would never have been a single cell that could be called the last universal common ancestor.<ref name = "Scientific American"/> | |||
| * 4 genes from ''Magnaporthe grisea'', the rice blast fungus, were suspected to be horizontally transferred from the genus ''Phytophthora'', and hypothesized to play a role in the fungus evolution into a plant pathogen.<ref>{{cite journal | vauthors = Richards TA, Dacks JB, Jenkinson JM, Thornton CR, Talbot NJ | title = Evolution of filamentous plant pathogens: gene exchange across eukaryotic kingdoms | journal = Current Biology | volume = 16 | issue = 18 | pages = 1857–1864 | date = September 2006 | pmid = 16979565 | doi = 10.1016/j.cub.2006.07.052 | bibcode = 2006CBio...16.1857R }}</ref> | |||
| :"As Woese has written, 'the ancestor cannot have been a particular organism, a single organismal lineage. It was communal, a loosely knit, diverse conglomeration of primitive cells that evolved as a unit, and it eventually developed to a stage where it broke into several distinct communities, which in their turn became the three primary lines of descent (], ] and ]s)' In other words, early cells, each having relatively few genes, differed in many ways. By swapping ]s freely, they shared various of their talents with their contemporaries. Eventually this collection of eclectic and changeable cells coalesced into the three basic domains known today. These domains become recognisable because much (though by no means all) of the gene transfer that occurs these days goes on within domains."<ref name = "Scientific American"/> | |||
| === Oomycetes to fungi === | |||
| With regard to how horizontal gene transfer affects evolutionary theory (common descent, universal phylogenetic tree) ] says: | |||
| * The oomycete species ''Phytophthora ramorum'', ''Phytophthora sojae'', ''Phytophthora infestans'', and ''Hyaloperonospora parasitica'' were estimated to have 33 horizontal gene transfers from fungi. The transferred genes were hypothesized to be involved in functions that facilitate plant tissues colonization, such as secreted proteins to evade plant immune response and breaking down plant cell walls.<ref>{{cite journal | vauthors = Richards TA, Soanes DM, Jones MD, Vasieva O, Leonard G, Paszkiewicz K, Foster PG, Hall N, Talbot NJ | title = Horizontal gene transfer facilitated the evolution of plant parasitic mechanisms in the oomycetes | journal = Proceedings of the National Academy of Sciences of the United States of America | volume = 108 | issue = 37 | pages = 15258–15263 | date = September 2011 | pmid = 21878562 | pmc = 3174590 | doi = 10.1073/pnas.1105100108 | doi-access = free | bibcode = 2011PNAS..10815258R }}</ref> | |||
| :"What elevated common descent to doctrinal status almost certainly was the much later discovery of the universality of biochemistry, which was seemingly impossible to explain otherwise. But that was before horizontal gene transfer (HGT), which could offer an alternative explanation for the universality of biochemistry, was recognized as a major part of the evolutionary dynamic. In questioning the doctrine of common descent, one necessarily questions the universal phylogenetic tree. That compelling tree image resides deep in our representation of biology. But the tree is no more than a graphical device; it is not some a priori form that nature imposes upon the evolutionary process. It is not a matter of whether your data are consistent with a tree, but whether tree topology is a useful way to represent your data. Ordinarily it is, of course, but the universal tree is no ordinary tree, and its root no ordinary root. Under conditions of extreme HGT, there is no (organismal) "tree." Evolution is basically reticulate."<ref> article ''A New Biology for a New Century'' by ]</ref> | |||
| === |
=== Animals to animals === | ||
| * ] fish received ] (AFP) gene from ] through a direct horizontal transfer.<ref>{{cite web|vauthors=Wilcox C|date=2021-06-09|title=DNA Jumps Between Animal Species. No One Knows How Often.|url=https://www.quantamagazine.org/dna-jumps-between-animal-species-no-one-knows-how-often-20210609/|access-date=2021-06-15|website=Quanta Magazine|language=en|archive-date=2021-06-14|archive-url=https://web.archive.org/web/20210614232456/https://www.quantamagazine.org/dna-jumps-between-animal-species-no-one-knows-how-often-20210609/|url-status=live}}</ref> | |||
| {{incomplete list}} | |||
| === Animals to bacteria === | |||
| There is evidence for historical horizontal transfer of the following genes: | |||
| *The strikingly fish-like copper/zinc superoxide dismutase of '']''<ref>{{cite journal | vauthors = Martin JP, Fridovich I | title = Evidence for a natural gene transfer from the ponyfish to its bioluminescent bacterial symbiont Photobacter leiognathi. The close relationship between bacteriocuprein and the copper-zinc superoxide dismutase of teleost fishes | journal = The Journal of Biological Chemistry | volume = 256 | issue = 12 | pages = 6080–6089 | date = June 1981 | doi = 10.1016/S0021-9258(19)69131-3 | pmid = 6787049 | doi-access = free }}</ref> is most easily explained in terms of transfer of a gene from an ancestor of its fish host. | |||
| *] ] for ] ], between ] and ].<ref> | |||
| {{cite journal | |||
| === Human to protozoan === | |||
| |author=D.A. Bryant & N.-U. Frigaard | |||
| *The ] ] '']'' acquired genetic material from humans that might help facilitate its long stay in the body.<ref>{{cite journal | vauthors = Bar D |title=Evidence of Massive Horizontal Gene Transfer Between Humans and ''Plasmodium vivax'' |date=16 February 2011 |journal=Nature Precedings |doi=10.1038/npre.2011.5690.1 |url=http://precedings.nature.com/documents/5690/version/1/ |doi-access=free |access-date=13 May 2011 |archive-date=31 March 2019 |archive-url=https://web.archive.org/web/20190331211353/http://precedings.nature.com/documents/5690/version/1/ |url-status=live }}</ref> | |||
| |month=Nov | |||
| |year=2006 | |||
| === Human genome === | |||
| |title=Prokaryotic photosynthesis and phototrophy illuminated | |||
| *One study identified approximately 100 of humans' approximately 20,000 total genes which likely resulted from horizontal gene transfer,<ref>{{cite news|date=14 March 2015|title=Human beings' ancestors have routinely stolen genes from other species|url=https://www.economist.com/news/science-and-technology/21646197-human-beings-ancestors-have-routinely-stolen-genes-other-species-genetically?fsrc=scn/gp/wl/pe/geneticallymodifiedpeople|newspaper=]|access-date=17 March 2015|archive-date=16 March 2015|archive-url=https://web.archive.org/web/20150316224918/http://www.economist.com/news/science-and-technology/21646197-human-beings-ancestors-have-routinely-stolen-genes-other-species-genetically?fsrc=scn/gp/wl/pe/geneticallymodifiedpeople|url-status=live}}</ref> but this number has been challenged by several researchers arguing these candidate genes for HGT are more likely the result of gene loss combined with differences in the rate of evolution.{{Citation needed|date=August 2024|reason=See talk page}} | |||
| |journal=Trends Microbiol. | |||
| |volume=14 | |||
| == Compounds found to promote horizontal gene transfer == | |||
| |issue=11 | |||
| Through research into the growing issue of ]<ref>{{cite journal | vauthors = Andersson DI, Hughes D | title = Microbiological effects of sublethal levels of antibiotics | journal = Nature Reviews. Microbiology | volume = 12 | issue = 7 | pages = 465–478 | date = July 2014 | pmid = 24861036 | doi = 10.1038/nrmicro3270 | s2cid = 3351736 }}</ref> certain compounds have been observed to promote horizontal gene transfer.<ref name="Wang_2021">{{cite journal | vauthors = Wang Y, Lu J, Zhang S, Li J, Mao L, Yuan Z, Bond PL, Guo J | title = Non-antibiotic pharmaceuticals promote the transmission of multidrug resistance plasmids through intra- and intergenera conjugation | journal = The ISME Journal | volume = 15 | issue = 9 | pages = 2493–2508 | date = September 2021 | pmid = 33692486 | pmc = 8397710 | doi = 10.1038/s41396-021-00945-7 | bibcode = 2021ISMEJ..15.2493W }}</ref><ref name="Jiao_2017">{{cite journal | vauthors = Jiao YN, Chen H, Gao RX, Zhu YG, Rensing C | title = Organic compounds stimulate horizontal transfer of antibiotic resistance genes in mixed wastewater treatment systems | journal = Chemosphere | volume = 184 | pages = 53–61 | date = October 2017 | pmid = 28578196 | doi = 10.1016/j.chemosphere.2017.05.149 | bibcode = 2017Chmsp.184...53J }}</ref><ref name="Ma_2021">{{cite journal | vauthors = Ma X, Zhang X, Xia J, Sun H, Zhang X, Ye L | title = Phenolic compounds promote the horizontal transfer of antibiotic resistance genes in activated sludge | journal = The Science of the Total Environment | volume = 800 | page = 149549 | date = December 2021 | pmid = 34392203 | doi = 10.1016/j.scitotenv.2021.149549 | bibcode = 2021ScTEn.80049549M }}</ref><ref name="Zhang_2018">{{cite journal | vauthors = Zhang Y, Gu AZ, Cen T, Li X, He M, Li D, Chen J | title = Sub-inhibitory concentrations of heavy metals facilitate the horizontal transfer of plasmid-mediated antibiotic resistance genes in water environment | journal = Environmental Pollution | volume = 237 | pages = 74–82 | date = June 2018 | pmid = 29477117 | doi = 10.1016/j.envpol.2018.01.032 | s2cid = 4911120 | doi-access = free | bibcode = 2018EPoll.237...74Z }}</ref> Antibiotics given to bacteria at non-lethal levels have been known to be a cause of antibiotic resistance<ref name="Zhang_2018" /> but emerging research is now showing that certain non-antibiotic pharmaceuticals (], ], ], ], ], etc.) also have a role in promoting antibiotic resistance through their ability to promote horizontal gene transfer (HGT) of genes responsible for antibiotic resistance. The transfer of antibiotic resistance genes (ARGs) through ] is significantly accelerated when donor cells with ]s and recipient cells are introduced to each other in the presence of one of the pharmaceuticals.<ref name="Wang_2021" /> Non-antibiotic pharmaceuticals were also found to cause some responses in bacteria similar to those responses to antibiotics, such as increasing expression of the genes lexA, umuC, umuD and soxR involved in the bacteria's SOS response as well as other genes also expressed during exposure to antibiotics.<ref name="Wang_2021" /> These findings are from 2021 and due to the widespread use of non-antibiotic pharmaceuticals, more research needs to be done in order to further understanding on the issue.<ref name="Wang_2021" /> | |||
| |pages=488 | |||
| |doi=10.1016/j.tim.2006.09.001 | |||
| Alongside non-antibiotic pharmaceuticals, other compounds relevant to antibiotic resistance have been tested such as ], ], ], ], ], ] solutions, ] (PNP), ] (PAP), and ] (PhOH).<ref name="Jiao_2017" /><ref name="Ma_2021" /> It is a global concern that ARGs have been found in ]<ref name="Jiao_2017" /> ] wastewater has been found to contain 3- to 13-fold higher abundance of ] than other samples of wastewater.<ref name="Jiao_2017" /> The cause of this is the organic compounds used for textile dying (''o''-xylene, ethylbenzene, trioxymethylene, styrene, 2,4-dichloroaniline, and malachite green)<ref name="Jiao_2017" /> raising the frequency of ] when bacteria and plasmid (with donor) are introduced in the presence of these molecules.<ref name="Jiao_2017" /> When textile wastewater combines with wastewater from ], the ARGs present in wastewater are transferred at a higher rate due to the addition of textile dyeing compounds increasing the occurrence of HGT.{{citation needed|date=February 2023}} | |||
| }}</ref> | |||
| Other organic pollutants commonly found in wastewater have been the subject of similar experiments.<ref name="Ma_2021" /> A 2021 study used similar methods of using plasmid in a donor and mixing that with a receptor in the presence of compound in order to test horizontal gene transfer of antibiotic resistance genes but this time in the presence of ].<ref name="Ma_2021" /> Phenolic compounds are commonly found in wastewater and have been found to change functions and structures of the ] during the wastewater treatment process.<ref name="Ma_2021" /> Additionally, HGT increases in frequency in the presence of the compounds p-nitrophenol (PNP), p-aminophenol (PAP), and phenol. These compounds result in a 2- to 9-fold increase in HGT (p-nitrophenol being on the lower side of 2-fold increases and p-aminophenol and phenol having a maximum increase of 9-fold).<ref name="Ma_2021" /> This increase in HGT is on average less than the compounds ibuprofen, naproxen, gemfibrozil, diclofenac, propranolol, o-xylene, ethylbenzene, trioxymethylene, styrene, 2,4-dichloroaniline, and malachite green<ref name="Wang_2021" /><ref name="Jiao_2017" /> but their increases is still significant.<ref name="Ma_2021" /> The study that came to this conclusion is similar to the study on horizontal gene transfer and non-antibiotic pharmaceuticals in that it was done in 2021 and leaves room for more research, specifically in the focus of the study which is ].<ref name="Ma_2021" /> | |||
| ] have also been found to promote conjugative transfer of antibiotic resistance genes.<ref name="Zhang_2018" /> The paper that led to the discovery of this was done in 2017 during the emerging field of horizontal gene transfer assisting compound research.<ref name="Zhang_2018" /> Metals assist in the spread of antibiotic resistance through both co-resistance as well as ] mechanisms.<ref name="Zhang_2018" /> In quantities relevant to the environment, ], ], ], and ] promote HGT from donor and receptor strains of ].<ref name="Zhang_2018" /> The presence of these metals triggered SOS response from bacterial cells and made the cells more permeable. These are the mechanisms that make even low levels of heavy metal pollution in the environment impact HGT and therefore the spread of ARGs. | |||
| == Promiscuous DNA == | |||
| Promiscuous DNA is a form of horizontal gene transfer that transmits genetic information across organellar barriers.<ref>{{Cite journal |last1=Schaack |first1=Sarah |last2=Gilbert |first2=Clément |last3=Feschotte |first3=Cédric |date=2010-09-01 |title=Promiscuous DNA: horizontal transfer of transposable elements and why it matters for eukaryotic evolution |journal=Trends in Ecology & Evolution |volume=25 |issue=9 |pages=537–546 |doi=10.1016/j.tree.2010.06.001 |pmid=20591532 |pmc=2940939 |bibcode=2010TEcoE..25..537S |issn=0169-5347}}</ref> Promiscuous DNA transfer has substantial evidence in its movement across the genome of numerous organisms, from movements in chloroplast to the nucleus,<ref>{{Cite journal |last1=Stegemann |first1=Sandra |last2=Hartmann |first2=Stefanie |last3=Ruf |first3=Stephanie |last4=Bock |first4=Ralph |date=2003-07-22 |title=High-frequency gene transfer from the chloroplast genome to the nucleus |journal=Proceedings of the National Academy of Sciences |volume=100 |issue=15 |pages=8828–8833 |doi=10.1073/pnas.1430924100 |doi-access=free |pmc=166398 |pmid=12817081|bibcode=2003PNAS..100.8828S }}</ref> chloroplast to the mitochondria,<ref>{{Cite journal |last1=Cerutti |first1=Heriberto |last2=Jagendorf |first2=André |date=1995-11-01 |title=Movement of DNA across the chloroplast envelope: Implications for the transfer of promiscuous DNA |url=https://link.springer.com/article/10.1007/BF00020448 |journal=Photosynthesis Research |language=en |volume=46 |issue=1 |pages=329–337 |doi=10.1007/BF00020448 |pmid=24301600 |bibcode=1995PhoRe..46..329C |issn=1573-5079}}</ref> and mitochondria to the nucleus.<ref name=":0">{{Cite web |last1=Sacerdot |first1=Christine |last2=Casaregola |first2=Serge |last3=Lafontaine |first3=Ingrid |last4=Tekaia |first4=Fredj |last5=Dujon |first5=Bernard |last6=Ozier-Kalogeropoulos |first6=Odile |date=1 September 2008 |title=Promiscuous DNA in the nuclear genomes of hemiascomycetous yeasts |url=https://academic.oup.com/femsyr/article/8/6/846/568124 |website=academic.oup.com}}</ref> | |||
| === History === | |||
| In 1982, ] defined this type of transpositional transfer mutation as “].”<ref>{{Cite journal |last=Ellis |first=John |date=October 1982 |title=Promiscuous DNA—chloroplast genes inside plant mitochondria |url=https://www.nature.com/articles/299678a0 |journal=Nature |language=en |volume=299 |issue=5885 |pages=678–679 |doi=10.1038/299678a0 |pmid=7121600 |bibcode=1982Natur.299..678E |issn=1476-4687}}</ref> Ellis further explored the phenomenon of “intracellular promiscuity” through the experiments of David Stern and David Lonsdale,<ref>{{Cite journal |last1=Stern |first1=David B. |last2=Lonsdale |first2=David M. |date=October 1982 |title=Mitochondrial and chloroplast genomes of maize have a 12-kilobase DNA sequence in common |url=https://www.nature.com/articles/299698a0 |journal=Nature |language=en |volume=299 |issue=5885 |pages=698–702 |doi=10.1038/299698a0 |pmid=6889685 |bibcode=1982Natur.299..698S |issn=1476-4687}}</ref> in which genetic transfer between chloroplasts to mitochondria was discovered, aiding in the definition and discovery of promiscuous DNA. | |||
| === Mechanism === | |||
| While much remains to be understood about how promiscuous DNA undergoes movement and transfer, numerous experiments have pointed to plastid sequences, ptDNA, as a key player.<ref>{{Cite journal |last1=Zeltz |first1=Patric |last2=Kadowaki |first2=Koh-ichi |last3=Kubo |first3=Nakao |last4=Maier |first4=Rainer M. |last5=Hirai |first5=Atsushi |last6=Kössel |first6=Hans |date=1996-06-01 |title=A promiscuous chloroplast DNA fragment is transcribed in plant mitochondria but the encoded RNA is not edited |url=https://link.springer.com/article/10.1007/BF00042236 |journal=Plant Molecular Biology |language=en |volume=31 |issue=3 |pages=647–656 |doi=10.1007/BF00042236 |pmid=8790296 |issn=1573-5028}}</ref><ref>{{Cite journal |last1=Park |first1=Hyun-Seung |last2=Jayakodi |first2=Murukarthick |last3=Lee |first3=Sae Hyun |last4=Jeon |first4=Jae-Hyeon |last5=Lee |first5=Hyun-Oh |last6=Park |first6=Jee Young |last7=Moon |first7=Byeong Cheol |last8=Kim |first8=Chang-Kug |last9=Wing |first9=Rod A. |last10=Newmaster |first10=Steven G. |last11=Kim |first11=Ji Yeon |last12=Yang |first12=Tae-Jin |date=2020-04-09 |title=Mitochondrial plastid DNA can cause DNA barcoding paradox in plants |journal=Scientific Reports |language=en |volume=10 |issue=1 |pages=6112 |doi=10.1038/s41598-020-63233-y |pmid=32273595 |pmc=7145815 |bibcode=2020NatSR..10.6112P |issn=2045-2322}}</ref><ref>{{Cite journal |last1=Ayliffe |first1=M A |last2=Scott |first2=N S |last3=Timmis |first3=J N |date=1 June 1998 |title="Analysis of plastid DNA-like sequences within the nuclear genomes of higher plants." |url=https://academic.oup.com/mbe/article/15/6/738/1026281 |journal=Molecular Biology and Evolution |volume=15 |issue=6 |pages=738–745 |doi=10.1093/oxfordjournals.molbev.a025977 |pmid=9615455 |via=Oxford Academic}}</ref> Plasmids, with their mobile nature and crucial role in horizontal gene transfer, are seen as a significant element in DNA that exchanges genetic information.<ref>{{Cite journal |last1=Suzuki |first1=Haruo |last2=Yano |first2=Hirokazu |last3=Brown |first3=Celeste J |last4=Top |first4=Eva M |date=27 September 2010 |title=Predicting Plasmid Promiscuity Based on Genomic Signature |journal=Journal of Bacteriology |volume=192 |issue=22 |pages=6045–6055 |doi=10.1128/JB.00277-10 |pmid=20851899 |pmc=2976448 }}</ref> This mobility makes ptDNA a potential donor for promiscuous DNA to traverse organellar barriers.<ref>{{Cite journal |last1=Cerutti |first1=Heriberto |last2=Jagendorf |first2=André |date=1995-11-01 |title=Movement of DNA across the chloroplast envelope: Implications for the transfer of promiscuous DNA |url=https://link.springer.com/article/10.1007/BF00020448 |journal=Photosynthesis Research |language=en |volume=46 |issue=1 |pages=329–337 |doi=10.1007/BF00020448 |pmid=24301600 |bibcode=1995PhoRe..46..329C |issn=1573-5079}}</ref> | |||
| === Types === | |||
| ==== NUMTs ==== | |||
| ] (nuclear sequences of mitochondrial) are a type of promiscuous DNA that arises from the natural transfer of ] (mtDNA) to the ] (nDNA).<ref>{{Cite web |last=Harutyunyan |first=Tigran |date=7 October 2023 |title=The known unknowns of mitochondrial carcinogenesis: de novo NUMTs and intercellular mitochondrial transfer |url=https://academic.oup.com/mutage/article-abstract/39/1/1/7296469 |website=Oxford Academic}}</ref> These NUMTs, with their varying frequencies, sizes, and features, contribute to the genetic diversity across all eukaryotes and, in some cases, to diseases among humans.<ref name=":0" /> | |||
| ==== NUPTs ==== | |||
| NUPTs (nuclear plastid DNA sequences) are a type of promiscuous DNA that arises from the natural transfer of ptDNA (]) into ].<ref>{{Cite journal |last1=Namasivayam |first1=Sivaranjani |last2=Sun |first2=Cheng |last3=Bah |first3=Assiatu B. |last4=Oberstaller |first4=Jenna |last5=Pierre-Louis |first5=Edwin |last6=Etheridge |first6=Ronald Drew |last7=Feschotte |first7=Cedric |last8=Pritham |first8=Ellen J. |last9=Kissinger |first9=Jessica C. |date=2023-11-07 |title=Massive invasion of organellar DNA drives nuclear genome evolution in Toxoplasma |journal=Proceedings of the National Academy of Sciences |volume=120 |issue=45 |pages=e2308569120 |doi=10.1073/pnas.2308569120 |pmc=10636329 |pmid=37917792|bibcode=2023PNAS..12008569N }}</ref> These fragments of ptDNA, similar to NUMTs in frequency, size, and features, also exhibit variability across species.<ref>{{Cite journal |last1=Michalovova |first1=M. |last2=Vyskot |first2=B. |last3=Kejnovsky |first3=E. |date=October 2013 |title=Analysis of plastid and mitochondrial DNA insertions in the nucleus (NUPTs and NUMTs) of six plant species: size, relative age and chromosomal localization |journal=Heredity |language=en |volume=111 |issue=4 |pages=314–320 |doi=10.1038/hdy.2013.51 |pmid=23715017 |issn=1365-2540|pmc=3807264 }}</ref> | |||
| ==Artificial horizontal gene transfer== | |||
| {{see also|Gene therapy}} | |||
| ], in which electric currents create the holes in the membrane. After conditions return to normal the holes in the membrane close and the ] are trapped inside the bacteria where they become part of the genetic material and their genes are expressed by the bacteria.]] | |||
| ] is essentially horizontal gene transfer, albeit with synthetic expression cassettes. The ]<ref>{{cite journal |vauthors=Ivics Z, Hackett PB, Plasterk RH, Izsvák Z |title=Molecular reconstruction of Sleeping Beauty, a Tc1-like transposon from fish, and its transposition in human cells |journal=Cell |volume=91 |issue=4 |pages=501–510 |date=November 1997 |pmid=9390559 |doi=10.1016/S0092-8674(00)80436-5 |s2cid=17908472 |doi-access=free}}</ref> (SB) was developed as a synthetic gene transfer agent that was based on the known abilities of ] transposons to invade genomes of extremely diverse species.<ref>{{cite book |vauthors=Plasterk RH |chapter=The Tc1/Mariner Transposon Family |veditors=Saedler H, Gierl A |title=Transposable Elements |pages=125–143 |date=1996 |pmid=8556864 |doi=10.1007/978-3-642-79795-8_6 |isbn=978-3-642-79797-2 |series=Current Topics in Microbiology and Immunology |volume=204|publisher=Springer |location=Berlin, Heidelberg }}</ref> The SB system has been used to introduce genetic sequences into a wide variety of animal genomes.<ref>{{cite journal |vauthors=Izsvák Z, Ivics Z, Plasterk RH |title=Sleeping Beauty, a wide host-range transposon vector for genetic transformation in vertebrates |journal=Journal of Molecular Biology |volume=302 |issue=1 |pages=93–102 |date=September 2000 |pmid=10964563 |doi=10.1006/jmbi.2000.4047}}</ref><ref>{{cite journal |vauthors=Kurtti TJ, Mattila JT, Herron MJ, Felsheim RF, Baldridge GD, Burkhardt NY, Blazar BR, Hackett PB, Meyer JM, Munderloh UG |title=Transgene expression and silencing in a tick cell line: A model system for functional tick genomics |journal=Insect Biochemistry and Molecular Biology |volume=38 |issue=10 |pages=963–968 |date=October 2008 |pmid=18722527 |pmc=2581827 |doi=10.1016/j.ibmb.2008.07.008|bibcode=2008IBMB...38..963K }}</ref> | |||
| == In evolution ==<!-- This section is linked from ] --> | |||
| {{Main articles|Horizontal gene transfer in evolution}} | |||
| Horizontal gene transfer is a potential ] in inferring ]s based on the ] of one gene.<ref name="Lawton_2009">{{cite web | vauthors = Lawton G | url = https://www.newscientist.com/article/mg20126921.600-why-darwin-was-wrong-about-the-tree-of-life.html?page=1 | title = Why Darwin was wrong about the tree of life | archive-url = https://web.archive.org/web/20150414011040/http://www.newscientist.com/article/mg20126921.600-why-darwin-was-wrong-about-the-tree-of-life.html?page=1 |archive-date=2015-04-14 | work = ] Magazine | issue = 2692 | date = 21 January 2009 }}</ref> For example, given two distantly related bacteria that have exchanged a gene a phylogenetic tree including those species will show them to be closely related because that gene is the same even though most other genes are dissimilar. For this reason, it is often ideal to use other information to infer robust phylogenies such as the presence or absence of genes or, more commonly, to include as wide a range of genes for phylogenetic analysis as possible. | |||
| For example, the most common gene to be used for constructing phylogenetic relationships in ]s is the ] gene since its sequences tend to be conserved among members with close phylogenetic distances, but variable enough that differences can be measured. However, in recent years it has also been argued that 16s rRNA genes can also be horizontally transferred. Although this may be infrequent, the validity of 16s rRNA-constructed phylogenetic trees must be reevaluated.<ref>{{cite journal | vauthors = Badger JH, Eisen JA, Ward NL | title = Genomic analysis of Hyphomonas neptunium contradicts 16S rRNA gene-based phylogenetic analysis: implications for the taxonomy of the orders 'Rhodobacterales' and Caulobacterales | journal = International Journal of Systematic and Evolutionary Microbiology | volume = 55 | issue = Pt 3 | pages = 1021–1026 | date = May 2005 | pmid = 15879228 | doi = 10.1099/ijs.0.63510-0 | doi-access = free }}</ref> | |||
| Biologist ] suggests "the original metaphor of a tree no longer fits the data from recent genome research" therefore "biologists should use the metaphor of a mosaic to describe the different histories combined in individual genomes and use the metaphor of a net to visualize the rich exchange and cooperative effects of HGT among microbes".<ref name="Gogarten_2000" /> There exist several methods to infer such ]s. | |||
| Using single genes as ]s, it is difficult to trace organismal ] in the presence of horizontal gene transfer. Combining the simple ] model of ] with rare HGT horizontal gene transfer events suggest there was no single ] that contained all of the genes ancestral to those shared among the three domains of ]. Each contemporary ] has its own history and traces back to an individual molecule ]. However, these molecular ancestors were likely to be present in different organisms at different times."<ref>{{cite journal | vauthors = Zhaxybayeva O, Gogarten JP | title = Cladogenesis, coalescence and the evolution of the three domains of life | journal = Trends in Genetics | volume = 20 | issue = 4 | pages = 182–7 | date = April 2004 | pmid = 15041172 | doi = 10.1016/j.tig.2004.02.004 }}</ref> | |||
| ===Challenge to the tree of life=== | |||
| {{further|Last universal common ancestor|tree of life (science)}} | |||
| Horizontal gene transfer poses a possible challenge to the concept of the ] (LUCA) at the root of the ] first formulated by ], which led him to propose the ] as a third domain of life.<ref name="Doolittle_2000">{{cite journal | vauthors = Doolittle WF | title = Uprooting the tree of life | journal = Scientific American | volume = 282 | issue = 2 | pages = 90–5 | date = February 2000 | pmid = 10710791 | doi = 10.1038/scientificamerican0200-90 | author-link = Ford Doolittle | bibcode = 2000SciAm.282b..90D }}</ref> Indeed, it was while examining the new three-domain view of life that horizontal gene transfer arose as a complicating issue: '']'' was seen as an anomaly with respect to a phylogenetic tree based upon the encoding for the ] ]—the organism in question is a definite Archaean, with all the cell lipids and transcription machinery that are expected of an Archaean, but whose HMGCoA genes are of bacterial origin.<ref name="Doolittle_2000"/> Scientists are broadly agreed on ], that ] in eukaryotes derived from ]l cells and that ]s came from ingested ], and other gene transfers may have affected early eukaryotes. (In contrast, multicellular eukaryotes have mechanisms to prevent horizontal gene transfer, including separated ]s.) If there had been continued and extensive gene transfer, there would be a complex network with many ancestors, instead of a tree of life with sharply delineated lineages leading back to a LUCA.<ref name="Doolittle_2000"/><ref>{{cite journal | vauthors = Woese CR | title = A new biology for a new century | journal = Microbiology and Molecular Biology Reviews | volume = 68 | issue = 2 | pages = 173–86 | date = June 2004 | pmid = 15187180 | pmc = 419918 | doi = 10.1128/MMBR.68.2.173-186.2004 }}</ref> However, a LUCA can be identified, so horizontal transfers must have been relatively limited.<ref name="Theobald_2010">{{cite journal | vauthors = Theobald DL | title = A formal test of the theory of universal common ancestry | journal = Nature | volume = 465 | issue = 7295 | pages = 219–22 | date = May 2010 | pmid = 20463738 | doi = 10.1038/nature09014 | s2cid = 4422345 | bibcode = 2010Natur.465..219T }}</ref> | |||
| Other early HGTs are thought to have happened. The ] (FUCA), earliest ancestor of LUCA, had other descendants that had their own lineages.<ref name="Harris_2021">{{cite journal | vauthors = Harris HM, Hill C | title = A Place for Viruses on the Tree of Life | journal = Frontiers in Microbiology | volume = 11 | pages = 604048 | date = 2021 | pmid = 33519747 | pmc = 7840587 | doi = 10.3389/fmicb.2020.604048 | doi-access = free }}</ref> These now-extinct sister lineages of LUCA descending from FUCA are thought to have horizontally transferred some of their genes into the genome of early descendants of LUCA.<ref name="Harris_2021" /> | |||
| === Phylogenetic information in HGT === | |||
| It has been remarked that, despite the complications, the detection of horizontal gene transfers brings valuable phylogenetic and dating information.<ref>{{cite book | vauthors = Huang J, Gogarten JP | chapter = Ancient Gene Transfer as a Tool in Phylogenetic Reconstruction | title = Horizontal Gene Transfer | volume = 532 | pages = 127–39 | date = 2009 | pmid = 19271182 | doi = 10.1007/978-1-60327-853-9_7 | publisher = Humana Press | isbn = 978-1-60327-852-2 | series = Methods in Molecular Biology }}</ref> | |||
| The potential of HGT to be used for dating phylogenies has recently been confirmed.<ref>{{cite journal | vauthors = Davín AA, Tannier E, Williams TA, Boussau B, Daubin V, Szöllősi GJ | title = Gene transfers can date the tree of life | language = En | journal = Nature Ecology & Evolution | volume = 2 | issue = 5 | pages = 904–909 | date = May 2018 | pmid = 29610471 | pmc = 5912509 | doi = 10.1038/s41559-018-0525-3 | bibcode = 2018NatEE...2..904D }}</ref><ref>{{cite journal | vauthors = Wolfe JM, Fournier GP | title = Horizontal gene transfer constrains the timing of methanogen evolution | language = En | journal = Nature Ecology & Evolution | volume = 2 | issue = 5 | pages = 897–903 | date = May 2018 | pmid = 29610466 | doi = 10.1038/s41559-018-0513-7 | bibcode = 2018NatEE...2..897W | hdl-access = free | s2cid = 4968981 | hdl = 1721.1/118329 }}</ref> | |||
| ===The chromosomal organization of horizontal gene transfer=== | |||
| The acquisition of new genes has the potential to disorganize the other genetic elements and hinder the function of the bacterial cell, thus affecting the competitiveness of bacteria. Consequently, bacterial adaptation lies in a conflict between the advantages of acquiring beneficial genes, and the need to maintain the organization of the rest of its genome. Horizontally transferred genes are typically concentrated in only ~1% of the chromosome (in regions called hotspots). This concentration increases with genome size and with the rate of transfer. Hotspots diversify by rapid gene turnover; their chromosomal distribution depends on local contexts (neighboring core genes), and content in mobile genetic elements. Hotspots concentrate most changes in gene repertoires, reduce the trade-off between genome diversification and organization, and should be treasure troves of strain-specific adaptive genes. Most mobile genetic elements and antibiotic resistance genes are in hotspots, but many hotspots lack recognizable mobile genetic elements and exhibit frequent homologous recombination at flanking core genes. Overrepresentation of hotspots with fewer mobile genetic elements in naturally transformable bacteria suggests that homologous recombination and horizontal gene transfer are tightly linked in genome evolution.<ref>{{cite journal | vauthors = Oliveira PH, Touchon M, Cury J, Rocha EP | title = The chromosomal organization of horizontal gene transfer in bacteria | journal = Nature Communications | volume = 8 | issue = 1 | page = 841 | date = October 2017 | pmid = 29018197 | pmc = 5635113 | doi = 10.1038/s41467-017-00808-w | bibcode = 2017NatCo...8..841O }}</ref> | |||
| ===Genes=== | |||
| {{Incomplete list|date=August 2008}} | |||
| There is evidence for historical horizontal transfer of the following genes: | |||
| *] ] for ] ], between ] and "]".<ref> | |||
| {{cite journal | vauthors = Bryant DA, Frigaard NU | title = Prokaryotic photosynthesis and phototrophy illuminated | journal = Trends in Microbiology | volume = 14 | issue = 11 | pages = 488–96 | date = November 2006 | pmid = 16997562 | doi = 10.1016/j.tim.2006.09.001 }}</ref> | |||
| *''TetO'' gene conferring resistance to ], between '']''.<ref>{{cite journal | vauthors = Avrain L, Vernozy-Rozand C, Kempf I | title = Evidence for natural horizontal transfer of tetO gene between Campylobacter jejuni strains in chickens | journal = Journal of Applied Microbiology | volume = 97 | issue = 1 | pages = 134–40 | year = 2004 | pmid = 15186450 | doi = 10.1111/j.1365-2672.2004.02306.x | s2cid = 19184139 }}</ref> | |||
| *Neochrome, a gene in some ferns that enhances their ability to survive in dim light. Believed to have been acquired from algae sometime during the Cretaceous.<ref> {{Webarchive|url=https://web.archive.org/web/20160307202422/http://blogs.scientificamerican.com/artful-amoeba/2014/05/06/in-darkened-forests-ferns-stole-gene-from-an-unlikely-source-and-then-each-other/In/ |date=2016-03-07 }} by Jennifer Frazer (May 6, 2014). ''Scientific American''.</ref><ref>{{cite journal | vauthors = Li FW, Rothfels CJ, Melkonian M, Villarreal JC, Stevenson DW, Graham SW, Wong GK, Mathews S, Pryer KM | title = The origin and evolution of phototropins | journal = Frontiers in Plant Science | volume = 6 | page = 637 | date = 2015 | pmid = 26322073 | pmc = 4532919 | doi = 10.3389/fpls.2015.00637 | doi-access = free }}</ref> | |||
| *Transfer of a ] from a bacterium into ] ]s and ] allowing the detoxification of ] produced by host plants.<ref>{{cite journal | vauthors = Wybouw N, Dermauw W, Tirry L, Stevens C, Grbić M, Feyereisen R, Van Leeuwen T | title = A gene horizontally transferred from bacteria protects arthropods from host plant cyanide poisoning | journal = eLife | volume = 3 | pages = e02365 | date = April 2014 | pmid = 24843024 | pmc = 4011162 | doi = 10.7554/eLife.02365 | doi-access = free }}</ref> | |||
| *The ] sequence has transferred from humans to the ] bacteria.<ref>{{cite web |url=http://phenomena.nationalgeographic.com/2011/02/16/gonorrhea-has-picked-up-human-dna-and-thats-just-the-beginning/|archive-url=https://web.archive.org/web/20130106204723/http://phenomena.nationalgeographic.com/2011/02/16/gonorrhea-has-picked-up-human-dna-and-thats-just-the-beginning/|archive-date=January 6, 2013|title=Gonorrhea has picked up human DNA (and that's just the beginning)| vauthors = Yong E |date=2011-02-16|publisher=National Geographic|language=en|access-date=2016-07-14}}</ref> | |||
| ==See also== | == See also == | ||
| {{Portal|Evolutionary biology}} | |||
| *] is a bacteria that is well known for its ability to transfer DNA between itself and plants. | |||
| {{div col|colwidth=18em}} | |||
| *], a bacterium well known for its ability to transfer DNA between itself and plants. | |||
| *] | *] | ||
| *] | |||
| *] | |||
| *] | |||
| *] | |||
| *] | *] | ||
| *] | |||
| *] | |||
| *] | |||
| *] | *] | ||
| *] | |||
| *] | *] | ||
| *] | |||
| *] | |||
| *] | |||
| *] | |||
| *] | |||
| {{div col end}} | |||
| == |
== References ==<!-- GenomeRes9:689 --> | ||
| {{ |
{{Reflist|30em}} | ||
| ==Further reading== | == Further reading == | ||
| {{refbegin|30em}} | |||
| * - article in ''Citizendium'' | |||
| * {{cite book |title=The Tangled Tree: A Radical New History of Life |year=2018 | vauthors = Quammen D |publisher=Simon & Schuster |isbn=978-1-4767-7662-0}} | |||
| * - article in ''Citizendium'' | |||
| * {{cite journal | vauthors = Gyles C, Boerlin P | title = Horizontally transferred genetic elements and their role in pathogenesis of bacterial disease | journal = Veterinary Pathology | volume = 51 | issue = 2 | pages = 328–40 | date = March 2014 | pmid = 24318976 | doi = 10.1177/0300985813511131 | s2cid = 206510894 | doi-access = free }} | |||
| * - article in ''Citizendium'' | |||
| * {{Webarchive|url=https://web.archive.org/web/20220227031337/http://vme.net/hgt/ |date=2022-02-27 }} | |||
| * - article in ''Citizendium'' | |||
| * {{cite journal | vauthors = Salzberg SL, White O, Peterson J, Eisen JA | s2cid = 17016011 | title = Microbial genes in the human genome: lateral transfer or gene loss? | journal = Science | volume = 292 | issue = 5523 | pages = 1903–6 | date = June 2001 | pmid = 11358996 | doi = 10.1126/science.1061036 | url = http://www.cbcb.umd.edu/~salzberg/docs/ScienceLateralTransfer.pdf | quote = About 40 genes were found to be exclusively shared by humans and bacteria and are candidate examples of horizontal transfer from bacteria to vertebrates. Gene loss combined with sample size effects and evolutionary rate variation provide an alternative, more biologically plausible explanation | bibcode = 2001Sci...292.1903S | access-date = 2005-12-29 | archive-date = 2006-09-01 | archive-url = https://web.archive.org/web/20060901131814/http://www.cbcb.umd.edu/~salzberg/docs/ScienceLateralTransfer.pdf }} | |||
| * | |||
| * {{cite journal | vauthors = Qi Z, Cui Y, Fang W, Ling L, Chen R | title = Autosomal similarity revealed by eukaryotic genomic comparison | journal = Journal of Biological Physics | volume = 30 | issue = 4 | pages = 305–12 | date = January 2004 | pmid = 23345874 | pmc = 3456315 | doi = 10.1007/s10867-004-0996-0 }} | |||
| * ''Steven L. Salzberg, Owen White, Jeremy Peterson, and Jonathan A. Eisen (2001) "Microbial Genes in the Human Genome: Lateral Transfer or Gene Loss?" Science 292, 1903-1906. (Free full article)'' This article points out that one dramatic claim of horizontal gene transfer - in which a distinguished group of scientists claimed that bacteria transferred their DNA directly into the human lineage - was simply wrong. | |||
| * |
* {{cite journal | vauthors = Woese CR | title = On the evolution of cells | journal = Proceedings of the National Academy of Sciences of the United States of America | volume = 99 | issue = 13 | pages = 8742–7 | date = June 2002 | pmid = 12077305 | pmc = 124369 | doi = 10.1073/pnas.132266999 | bibcode = 2002PNAS...99.8742W | doi-access = free }} This article seeks to shift the emphasis in early ] from vertical to horizontal gene transfer. He uses the term "Darwinian Threshold" for the time of major transition of evolutionary mechanisms from mostly horizontal to mostly vertical transfer, and the "origin of speciation". | ||
| * |
* {{cite journal | vauthors = Snel B, Bork P, Huynen MA | title = Genome phylogeny based on gene content | journal = Nature Genetics | volume = 21 | issue = 1 | pages = 108–10 | date = January 1999 | pmid = 9916801 | doi = 10.1038/5052 | s2cid = 10296406 }} This article proposes using the presence or absence of a set of genes to infer phylogenies, in order to avoid confounding factors such as horizontal gene transfer. | ||
| * |
* {{cite web |url=http://www.nature.com/nrmicro/focus/genetransfer/index.html |title=Webfocus in Nature with free review articles |archive-url=https://web.archive.org/web/20051102081911/http://www.nature.com/nrmicro/focus/genetransfer/index.html |archive-date=2005-11-02 }} | ||
| * |
* {{cite journal | vauthors = Patil PB, Sonti RV | title = Variation suggestive of horizontal gene transfer at a lipopolysaccharide (lps) biosynthetic locus in Xanthomonas oryzae pv. oryzae, the bacterial leaf blight pathogen of rice | journal = BMC Microbiology | volume = 4 | issue = 1 | page = 40 | date = October 2004 | pmid = 15473911 | pmc = 524487 | doi = 10.1186/1471-2180-4-40 | doi-access = free }} | ||
| * {{cite journal | vauthors = Jin G, Nakhleh L, Snir S, Tuller T | title = Maximum likelihood of phylogenetic networks | journal = Bioinformatics | volume = 22 | issue = 21 | pages = 2604–11 | date = November 2006 | pmid = 16928736 | doi = 10.1093/bioinformatics/btl452 | doi-access = free }} | |||
| *Bioinformatics Vol. 22 no. 21 2006, pages 2604–2611 for a technique to decrease the impact of HGT events on maximum likelihood cladistical analyses. | |||
| * {{cite journal | vauthors = Jain R, Rivera MC, Lake JA | title = Horizontal gene transfer among genomes: the complexity hypothesis | journal = Proceedings of the National Academy of Sciences of the United States of America | volume = 96 | issue = 7 | pages = 3801–6 | date = March 1999 | pmid = 10097118 | pmc = 22375 | doi = 10.1073/pnas.96.7.3801 | bibcode = 1999PNAS...96.3801J | doi-access = free }} | |||
| * | |||
| * {{cite journal | vauthors = Ochman H, Lawrence JG, Groisman EA | title = Lateral gene transfer and the nature of bacterial innovation | journal = Nature | volume = 405 | issue = 6784 | pages = 299–304 | date = May 2000 | pmid = 10830951 | doi = 10.1038/35012500 | bibcode = 2000Natur.405..299O | s2cid = 85739173 }} | |||
| * | |||
| * {{cite magazine |title=The Demon in the Freezer |url=https://www.newyorker.com/magazine/1999/07/12/the-demon-in-the-freezer |vauthors=Preston R |magazine=The New Yorker |date=July 12, 1999 |pages=44–61 |quote=Smallpox knows how to make a mouse protein. How did smallpox learn that? 'The poxviruses are promiscuous at capturing genes from their hosts,' Esposito said. 'It tells you that smallpox was once inside a mouse or some other small rodent.' |access-date=February 20, 2020 |archive-date=March 18, 2023 |archive-url=https://web.archive.org/web/20230318013950/https://www.newyorker.com/magazine/1999/07/12/the-demon-in-the-freezer |url-status=live }} | |||
| * | |||
| * {{cite journal | vauthors = Szpirer C, Top E, Couturier M, Mergeay M | title = Retrotransfer or gene capture: a feature of conjugative plasmids, with ecological and evolutionary significance | journal = Microbiology | volume = 145 | issue = Pt 12 | pages = 3321–3329 | date = December 1999 | pmid = 10627031 | doi = 10.1099/00221287-145-12-3321 | doi-access = free | url = http://mic.sgmjournals.org/cgi/content/full/145/12/3321 | access-date = 2006-05-12 | archive-date = 2007-02-23 | archive-url = https://web.archive.org/web/20070223201609/http://mic.sgmjournals.org/cgi/content/full/145/12/3321 | url-status = live }} | |||
| * | |||
| * {{cite web |url=http://www.gmo-safety.eu/topics/3?mode=prj |work=GMO Safety: Results of research into horizontal gene transfer |title=Can transgenes from genetically modified plants be absorbed by micro-organisms and spread in this way? |archive-url=https://web.archive.org/web/20110721173953/http://www.gmo-safety.eu/topics/3?mode=prj |archive-date=2011-07-21 }} | |||
| * | |||
| * {{cite journal | vauthors = Whitaker JW, McConkey GA, Westhead DR | title = The transferome of metabolic genes explored: analysis of the horizontal transfer of enzyme encoding genes in unicellular eukaryotes | journal = Genome Biology | volume = 10 | issue = 4 | pages = R36 | year = 2009 | pmid = 19368726 | pmc = 2688927 | doi = 10.1186/gb-2009-10-4-r36 | doi-access = free }} | |||
| * | |||
| {{refend}} | |||
| * | |||
| * "Smallpox knows how to make a mouse protein. How did smallpox learn that? 'The poxviruses are promiscuous at capturing genes from their hosts,' Esposito said. 'It tells you that smallpox was once inside a mouse or some other small rodent.'" | |||
| * | |||
| * Can transgenes from genetically modified plants be absorbed by micro-organisms and spread in this way? | |||
| * | |||
| * {{cite book |chapterurl=http://www.horizonpress.com/pla|author= Sota M; Top EM|year=2008|chapter=Horizontal Gene Transfer Mediated by Plasmids|title=Plasmids: Current Research and Future Trends|publisher=Caister Academic Press|id=}} | |||
| == External links == | |||
| {{Scholia|topic}} | |||
| *] | |||
| *] | |||
| *] | |||
| *] | |||
| {{Genetic recombination}} | {{Genetic recombination}} | ||
| ] | |||
| ] | |||
| ] | |||
| {{DEFAULTSORT:Horizontal Gene Transfer}} | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
| ] | |||
Latest revision as of 01:38, 26 December 2024
Transfer of genes from unrelated organisms"HGT" redirects here. For other uses, see HGT (disambiguation). This article is about the natural process. For artificial gene transfer, see Gene delivery.
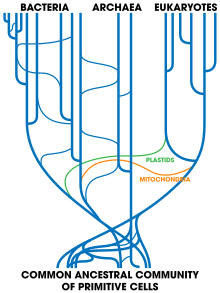
Horizontal gene transfer (HGT) or lateral gene transfer (LGT) is the movement of genetic material between organisms other than by the ("vertical") transmission of DNA from parent to offspring (reproduction). HGT is an important factor in the evolution of many organisms. HGT is influencing scientific understanding of higher-order evolution while more significantly shifting perspectives on bacterial evolution.
Horizontal gene transfer is the primary mechanism for the spread of antibiotic resistance in bacteria, and plays an important role in the evolution of bacteria that can degrade novel compounds such as human-created pesticides and in the evolution, maintenance, and transmission of virulence. It often involves temperate bacteriophages and plasmids. Genes responsible for antibiotic resistance in one species of bacteria can be transferred to another species of bacteria through various mechanisms of HGT such as transformation, transduction and conjugation, subsequently arming the antibiotic resistant genes' recipient against antibiotics. The rapid spread of antibiotic resistance genes in this manner is becoming a challenge to manage in the field of medicine. Ecological factors may also play a role in the HGT of antibiotic resistant genes.
Horizontal gene transfer is recognized as a pervasive evolutionary process that distributes genes between divergent prokaryotic lineages and can also involve eukaryotes. HGT events are thought to occur less frequently in eukaryotes than in prokaryotes. However, growing evidence indicates that HGT is relatively common among many eukaryotic species and can have an impact on adaptation to novel environments. Its study, however, is hindered by the complexity of eukaryotic genomes and the abundance of repeat-rich regions, which complicate the accurate identification and characterization of transferred genes.
It is postulated that HGT promotes the maintenance of a universal life biochemistry and, subsequently, the universality of the genetic code.
History
Griffith's experiment, reported in 1928 by Frederick Griffith, was the first experiment suggesting that bacteria are capable of transferring genetic information through a process known as transformation. Griffith's findings were followed by research in the late 1930s and early 1940s that isolated DNA as the material that communicated this genetic information.
Horizontal genetic transfer was then described in Seattle in 1951, in a paper demonstrating that the transfer of a viral gene into Corynebacterium diphtheriae created a virulent strain from a non-virulent strain, simultaneously revealing the mechanism of diphtheria (that patients could be infected with the bacteria but not have any symptoms, and then suddenly convert later or never), and giving the first example for the relevance of the lysogenic cycle. Inter-bacterial gene transfer was first described in Japan in a 1959 publication that demonstrated the transfer of antibiotic resistance between different species of bacteria. In the mid-1980s, Syvanen postulated that biologically significant lateral gene transfer has existed since the beginning of life on Earth and has been involved in shaping all of evolutionary history.
As Jian, Rivera and Lake (1999) put it: "Increasingly, studies of genes and genomes are indicating that considerable horizontal transfer has occurred between prokaryotes" (see also Lake and Rivera, 2007). The phenomenon appears to have had some significance for unicellular eukaryotes as well. As Bapteste et al. (2005) observe, "additional evidence suggests that gene transfer might also be an important evolutionary mechanism in protist evolution."
Grafting of one plant to another can transfer chloroplasts (organelles in plant cells that conduct photosynthesis), mitochondrial DNA, and the entire cell nucleus containing the genome to potentially make a new species. Some Lepidoptera (e.g. monarch butterflies and silkworms) have been genetically modified by horizontal gene transfer from the wasp bracovirus. Bites from insects in the family Reduviidae (assassin bugs) can, via a parasite, infect humans with the trypanosomal Chagas disease, which can insert its DNA into the human genome. It has been suggested that lateral gene transfer to humans from bacteria may play a role in cancer.
Aaron Richardson and Jeffrey D. Palmer state: "Horizontal gene transfer (HGT) has played a major role in bacterial evolution and is fairly common in certain unicellular eukaryotes. However, the prevalence and importance of HGT in the evolution of multicellular eukaryotes remain unclear."
Due to the increasing amount of evidence suggesting the importance of these phenomena for evolution (see below) molecular biologists such as Peter Gogarten have described horizontal gene transfer as "A New Paradigm for Biology".
Mechanisms
There are several mechanisms for horizontal gene transfer:
- Transformation, the genetic alteration of a cell resulting from the introduction, uptake and expression of foreign genetic material (DNA or RNA). This process is relatively common in bacteria, but less so in eukaryotes. Transformation is often used in laboratories to insert novel genes into bacteria for experiments or for industrial or medical applications. See also molecular biology and biotechnology.
- Transduction, the process in which bacterial DNA is moved from one bacterium to another by a virus (a bacteriophage, or phage).
- Bacterial conjugation, a process that involves the transfer of DNA via a plasmid from a donor cell to a recombinant recipient cell during cell-to-cell contact.
- Gene transfer agents, virus-like elements encoded by the host that are found in the alphaproteobacteria order Rhodobacterales.
Horizontal transposon transfer
A transposable element (TE) (also called a transposon or jumping gene) is a mobile segment of DNA that can sometimes pick up a resistance gene and insert it into a plasmid or chromosome, thereby inducing horizontal gene transfer of antibiotic resistance.
Horizontal transposon transfer (HTT) refers to the passage of pieces of DNA that are characterized by their ability to move from one locus to another between genomes by means other than parent-to-offspring inheritance. Horizontal gene transfer has long been thought to be crucial to prokaryotic evolution, but there is a growing amount of data showing that HTT is a common and widespread phenomenon in eukaryote evolution as well. On the transposable element side, spreading between genomes via horizontal transfer may be viewed as a strategy to escape purging due to purifying selection, mutational decay and/or host defense mechanisms.
HTT can occur with any type of transposable elements, but DNA transposons and LTR retroelements are more likely to be capable of HTT because both have a stable, double-stranded DNA intermediate that is thought to be sturdier than the single-stranded RNA intermediate of non-LTR retroelements, which can be highly degradable. Non-autonomous elements may be less likely to transfer horizontally compared to autonomous elements because they do not encode the proteins required for their own mobilization. The structure of these non-autonomous elements generally consists of an intronless gene encoding a transposase protein, and may or may not have a promoter sequence. Those that do not have promoter sequences encoded within the mobile region rely on adjacent host promoters for expression. Horizontal transfer is thought to play an important role in the TE life cycle. In plants, it appears that LTR retrotransposons of the Copia superfamilies, especially those with low copy numbers from the Ale and Ivana lineages, are more likely to undergo horizontal transfer between different plant species.
HTT has been shown to occur between species and across continents in both plants and animals (Ivancevic et al. 2013), though some TEs have been shown to more successfully colonize the genomes of certain species over others. Both spatial and taxonomic proximity of species has been proposed to favor HTTs in plants and animals. It is unknown how the density of a population may affect the rate of HTT events within a population, but close proximity due to parasitism and cross contamination due to crowding have been proposed to favor HTT in both plants and animals. In plants, the interaction between lianas and trees has been shown to facilitate HTT in natural ecosystems. Successful transfer of a transposable element requires delivery of DNA from donor to host cell (and to the germ line for multi-cellular organisms), followed by integration into the recipient host genome. Though the actual mechanism for the transportation of TEs from donor cells to host cells is unknown, it is established that naked DNA and RNA can circulate in bodily fluid. Many proposed vectors include arthropods, viruses, freshwater snails (Ivancevic et al. 2013), endosymbiotic bacteria, and intracellular parasitic bacteria. In some cases, even TEs facilitate transport for other TEs.
The arrival of a new TE in a host genome can have detrimental consequences because TE mobility may induce mutation. However, HTT can also be beneficial by introducing new genetic material into a genome and promoting the shuffling of genes and TE domains among hosts, which can be co-opted by the host genome to perform new functions. Moreover, transposition activity increases the TE copy number and generates chromosomal rearrangement hotspots. HTT detection is a difficult task because it is an ongoing phenomenon that is constantly changing in frequency of occurrence and composition of TEs inside host genomes. Furthermore, few species have been analyzed for HTT, making it difficult to establish patterns of HTT events between species. These issues can lead to the underestimation or overestimation of HTT events between ancestral and current eukaryotic species.
Methods of detection

Horizontal gene transfer is typically inferred using bioinformatics methods, either by identifying atypical sequence signatures ("parametric" methods) or by identifying strong discrepancies between the evolutionary history of particular sequences compared to that of their hosts. The transferred gene (xenolog) found in the receiving species is more closely related to the genes of the donor species than would be expected.
Viruses
The virus called Mimivirus infects amoebae. Another virus, called Sputnik, also infects amoebae, but it cannot reproduce unless mimivirus has already infected the same cell.
Sputnik's genome reveals further insight into its biology. Although 13 of its genes show little similarity to any other known genes, three are closely related to mimivirus and mamavirus genes, perhaps cannibalized by the tiny virus as it packaged up particles sometime in its history. This suggests that the satellite virus could perform horizontal gene transfer between viruses, paralleling the way that bacteriophages ferry genes between bacteria.
Horizontal transfer is also seen between geminiviruses and tobacco plants.
Prokaryotes
Horizontal gene transfer is common among bacteria, even among very distantly related ones. This process is thought to be a significant cause of increased drug resistance when one bacterial cell acquires resistance, and the resistance genes are transferred to the other species. Transposition and horizontal gene transfer, along with strong natural selective forces have led to multi-drug resistant strains of S. aureus and many other pathogenic bacteria. Horizontal gene transfer also plays a role in the spread of virulence factors, such as exotoxins and exoenzymes, amongst bacteria. A prime example concerning the spread of exotoxins is the adaptive evolution of Shiga toxins in E. coli through horizontal gene transfer via transduction with Shigella species of bacteria. Strategies to combat certain bacterial infections by targeting these specific virulence factors and mobile genetic elements have been proposed. For example, horizontally transferred genetic elements play important roles in the virulence of E. coli, Salmonella, Streptococcus and Clostridium perfringens.
In prokaryotes, restriction-modification systems are known to provide immunity against horizontal gene transfer and in stabilizing mobile genetic elements. Genes encoding restriction modification systems have been reported to move between prokaryotic genomes within mobile genetic elements (MGE) such as plasmids, prophages, insertion sequences/transposons, integrative conjugative elements (ICE), and integrons. Still, they are more frequently a chromosomal-encoded barrier to MGE than an MGE-encoded tool for cell infection.
Lateral gene transfer via a mobile genetic element, namely the integrated conjugative element (ICE) Bs1 has been reported for its role in the global DNA damage SOS response of the gram positive Bacillus subtilis. Furthermore, it has been linked with the radiation and desiccation resistance of Bacillus pumilus SAFR-032 spores, isolated from spacecraft cleanroom facilities.
Transposon insertion elements have been reported to increase the fitness of gram-negative E. coli strains through either major transpositions or genome rearrangements, and increasing mutation rates. In a study on the effects of long-term exposure of simulated microgravity on non-pathogenic E. coli, the results showed transposon insertions occur at loci, linked to SOS stress response. When the same E. coli strain was exposed to a combination of simulated microgravity and trace (background) levels of (the broad spectrum) antibiotic (chloramphenicol), the results showed transposon-mediated rearrangements (TMRs), disrupting genes involved in bacterial adhesion, and deleting an entire segment of several genes involved with motility and chemotaxis. Both these studies have implications for microbial growth, adaptation to and antibiotic resistance in real time space conditions.
Horizontal gene transfer is particularly active in bacterial genomes around the production of secondary or specialized metabolites. This is clearly exhibited within certain groups of bacteria including P. aeruginosa and actinomycetales, an order of Actinomycetota. Polyketide synthases (PKSs) and biosynthetic gene clusters provide modular organizations of associated genes making these bacteria well-adapted to acquire and discard helpful modular modifications via HGT. Certain areas of genes known as hotspots further increase the likelihood of horizontally transferred secondary metabolite-producing genes. The promiscuity of enzymes is a reoccurring theme in this particular theatre.
Bacterial transformation

Natural transformation is a bacterial adaptation for DNA transfer (HGT) that depends on the expression of numerous bacterial genes whose products are responsible for this process. In general, transformation is a complex, energy-requiring developmental process. In order for a bacterium to bind, take up and recombine exogenous DNA into its chromosome, it must become competent, that is, enter a special physiological state. Competence development in Bacillus subtilis requires expression of about 40 genes. The DNA integrated into the host chromosome is usually (but with infrequent exceptions) derived from another bacterium of the same species, and is thus homologous to the resident chromosome. The capacity for natural transformation occurs in at least 67 prokaryotic species. Competence for transformation is typically induced by high cell density and/or nutritional limitation, conditions associated with the stationary phase of bacterial growth. Competence appears to be an adaptation for DNA repair. Transformation in bacteria can be viewed as a primitive sexual process, since it involves interaction of homologous DNA from two individuals to form recombinant DNA that is passed on to succeeding generations. Although transduction is the form of HGT most commonly associated with bacteriophages, certain phages may also be able to promote transformation.
Bacterial conjugation
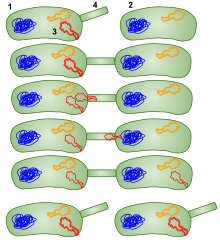
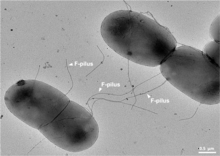
As mentioned before, conjugation is a method of horizontal gene transfer through cell to cell contact. Through the process of conjugation, type IV Secretion Systems (T4SS) are used to passage on DNA from the donor cell to the recipient cell. These T4SS encoded within the plasmid carry other proteins and genes that help supplement the cell in conjugation. Research has shown that there are Single Binding DNA Binding proteins (SSBs) also encoded within the conjugative plasmid may help with conjugation and cell viability. This is thought to be the case because SSBs naturally are expressed to help with stabilizing single-stranded DNA (ssDNA). SSBs will also recruit other proteins like RadD or RecA expressed in events of DNA recombination, repair, and replication. Further showcasing their possible role in conjugation. Although it may help, studies have also shown for proteins like SSB to not be essential in conjugation. For example, the plasmid pCF10 from Enterococcus faecalis, a gram-positive bacterium, has a SSB like-protein called PrgE and was classified for not being required for conjugation. More work needs to be done on why proteins that bind to ssDNA are encoded into conjugative plasmids.
Conjugation in the case of microbiomes and symbioses is very important. From this process new genes are acquired that lead to increasing genetic diversity and evolution such as the acquisition of antibiotic resistance genes. Mycobacterium tuberculosis is a species that has evolved through methods like conjugation while gaining antibiotic resistance. This evolution or increase in genetic diversity is also seen in many other species. Due to this, there is a huge concern on how impactful conjugation or horizontal gene transfer can be on human health and your microbiome as pathogenic microbes can become more pathogenic. Studies have shown that even our own microbiome has a plethora of antimicrobial genes which if transferred to pathogenic microbes could be detrimental.
Conjugation in Mycobacterium smegmatis, like conjugation in E. coli, requires stable and extended contact between a donor and a recipient strain, is DNase resistant, and the transferred DNA is incorporated into the recipient chromosome by homologous recombination. However, unlike E. coli high frequency of recombination conjugation (Hfr), mycobacterial conjugation is a type of HGT that is chromosome rather than plasmid based. Furthermore, in contrast to E. coli (Hfr) conjugation, in M. smegmatis all regions of the chromosome are transferred with comparable efficiencies. Substantial blending of the parental genomes was found as a result of conjugation, and this blending was regarded as reminiscent of that seen in the meiotic products of sexual reproduction.
Archaeal DNA transfer
Haloarchaea are aerobic halophiles thought to have evolved from anaerobic methanogens. A large amount of their genome, 126 composite gene families, are derived from genetic material from bacterial genomes. This has allowed them to adapt to extremely salty environments.
The archaeon Sulfolobus solfataricus, when UV irradiated, strongly induces the formation of type IV pili which then facilitates cellular aggregation. Exposure to chemical agents that cause DNA damage also induces cellular aggregation. Other physical stressors, such as temperature shift or pH, do not induce aggregation, suggesting that DNA damage is a specific inducer of cellular aggregation.
UV-induced cellular aggregation mediates intercellular chromosomal HGT marker exchange with high frequency, and UV-induced cultures display recombination rates that exceed those of uninduced cultures by as much as three orders of magnitude. S. solfataricus cells aggregate preferentially with other cells of their own species. Frols et al. and Ajon et al. suggested that UV-inducible DNA transfer is likely an important mechanism for providing increased repair of damaged DNA via homologous recombination. This process can be regarded as a simple form of sexual interaction.
Another thermophilic species, Sulfolobus acidocaldarius, is able to undergo HGT. S. acidocaldarius can exchange and recombine chromosomal markers at temperatures up to 84 °C. UV exposure induces pili formation and cellular aggregation. Cells with the ability to aggregate have greater survival than mutants lacking pili that are unable to aggregate. The frequency of recombination is increased by DNA damage induced by UV-irradiation and by DNA damaging chemicals.
The ups operon, containing five genes, is highly induced by UV irradiation. The proteins encoded by the ups operon are employed in UV-induced pili assembly and cellular aggregation leading to intercellular DNA exchange and homologous recombination. Since this system increases the fitness of S. acidocaldarius cells after UV exposure, Wolferen et al. considered that transfer of DNA likely takes place in order to repair UV-induced DNA damages by homologous recombination.
Eukaryotes
"Sequence comparisons suggest recent horizontal transfer of many genes among diverse species including across the boundaries of phylogenetic 'domains'. Thus determining the phylogenetic history of a species can not be done conclusively by determining evolutionary trees for single genes."
Organelle to nuclear genome
- Analysis of DNA sequences suggests that horizontal gene transfer has occurred within eukaryotes from the chloroplast and mitochondrial genomes to the nuclear genome. As stated in the endosymbiotic theory, chloroplasts and mitochondria probably originated as bacterial endosymbionts of a progenitor to the eukaryotic cell.
Organelle to organelle
- Mitochondrial genes moved to parasites of the Rafflesiaceae plant family from their hosts and from chloroplasts of a still-unidentified plant to the mitochondria of the bean Phaseolus.
Bacteria to fungi
- Horizontal transfer occurs from bacteria to some fungi, such as the yeast Saccharomyces cerevisiae.
Bacteria to plants
- Agrobacterium, a pathogenic bacterium that causes cells to proliferate as crown galls and proliferating roots is an example of a bacterium that can transfer genes to plants and this plays an important role in plant evolution.
- Land plants and their close relatives, the charophycean green algae, share a set of glycosyl hydrolases. These enzymes were likely transferred from bacteria and fungi to the last common ancestor of these organisms before the origin of land plants.
Bacteria to animals
- HhMAN1 is a gene in the genome of the coffee berry borer (Hypothenemus hampei) that resembles bacterial genes, and is thought to be transferred from bacteria in the beetle's gut.
- oskar is an essential gene for the specification of the germline in Holometabola and its origin is through to be due to a HGT event followed by a fusion with a LOTUS domain.
- Bdelloid rotifers currently hold the 'record' for HGT in animals with ~8% of their genes from bacterial origins. Tardigrades were thought to break the record with 17.5% HGT, but that finding was an artifact of bacterial contamination.
- A study found the genomes of 40 animals (including 10 primates, four Caenorhabditis worms, and 12 Drosophila insects) contained genes which the researchers concluded had been transferred from bacteria and fungi by horizontal gene transfer. The researchers estimated that for some nematodes and Drosophila insects these genes had been acquired relatively recently.
- A bacteriophage-mediated mechanism transfers genes between prokaryotes and eukaryotes. Nuclear localization signals in bacteriophage terminal proteins (TP) prime DNA replication and become covalently linked to the viral genome. The role of virus and bacteriophages in HGT in bacteria, suggests that TP-containing genomes could be a vehicle of inter-kingdom genetic information transference all throughout evolution.
- The adzuki bean beetle has acquired genetic material from its (non-beneficial) endosymbiont Wolbachia. New examples have recently been reported demonstrating that Wolbachia bacteria represent an important potential source of genetic material in arthropods and filarial nematodes.
- The psyllid Pachypsylla venusta has acquired genes from its current endosymbiont Carsonella, and from many of its historical endosymbionts, too.
Plant to plant
- Striga hermonthica, a parasitic eudicot, has received a gene from sorghum (Sorghum bicolor) to its nuclear genome. The gene's functionality is unknown.
- A gene that allowed ferns to survive in dark forests came from the hornwort, which grows in mats on streambanks or trees. The neochrome gene arrived about 180 million years ago.
- Transfer of mRNA between host plants and heterotrophs plants in the Orobanchaceae have been directly observed. mRNA transcripts can therefore be a factor involved in the transfer and integration of foreign DNA in heterotrophs.
Plants to animals
- The eastern emerald sea slug Elysia chlorotica has been suggested by fluorescence in situ hybridization (FISH) analysis to contain photosynthesis-supporting genes obtained from an algae (Vaucheria litorea) in their diet. LGT in Sacoglossa is now thought to be an artifact and no trace of LGT was found upon sequencing the genome of Elysia chlorotica.
- The whitefly Bemisia tabaci acquired a plant detoxification gene that neutralizes plant toxins.
Plant to fungus
Main article: Plant-fungus horizontal gene transfer- Gene transfer between plants and fungi has been posited for a number of cases, including rice (Oryza sativa).
- Evidence of gene transfer from plants was documented in the fungus Colletotrichum.
- Plant expansin genes were transferred to fungi further enabling the fungi to infect plants.
Plant to bacteria
- Plant expansin genes were transferred to bacteria further enabling the bacteria to infect plants.
Fungi to insects
- Pea aphids (Acyrthosiphon pisum) contain multiple genes from fungi. Plants, fungi, and microorganisms can synthesize carotenoids, but torulene made by pea aphids is the only carotenoid known to be synthesized by an organism in the animal kingdom.
Fungi to fungi
- The toxin α-amanitin is found in numerous, seemingly unrelated genera fungi such as Amanita, Lepiota, and Galerina. Two biosynthetic genes involved in the production of α-amanitin are P450-29 and FMO1. Phylogenetic and genetic analyses of these genes strongly indicate that they were transferred between the genera via horizontal gene transfer.
- The ToxA protein (wheat virulence protein) included in a ∼14 kb element, containing both coding and non-coding regions was transferred between different fungal wheat patogens: Parastagonospora nodorum, Pyrenophora tritici-repentis, and Bipolaris sorokiniana.
- A large genomic element named "Wallaby," approximately 500 kb in length, was recently transferred between two Penicillium species used in cheesemaking: P. camemberti and P. roqueforti. Wallaby contains around 250 genes, including several that are thought to play a role in microbial competition.
Fungi to oomycetes
- 4 genes from Magnaporthe grisea, the rice blast fungus, were suspected to be horizontally transferred from the genus Phytophthora, and hypothesized to play a role in the fungus evolution into a plant pathogen.
Oomycetes to fungi
- The oomycete species Phytophthora ramorum, Phytophthora sojae, Phytophthora infestans, and Hyaloperonospora parasitica were estimated to have 33 horizontal gene transfers from fungi. The transferred genes were hypothesized to be involved in functions that facilitate plant tissues colonization, such as secreted proteins to evade plant immune response and breaking down plant cell walls.
Animals to animals
- Smelt fish received antifreeze protein (AFP) gene from herring through a direct horizontal transfer.
Animals to bacteria
- The strikingly fish-like copper/zinc superoxide dismutase of Photobacterium leiognathi is most easily explained in terms of transfer of a gene from an ancestor of its fish host.
Human to protozoan
- The malaria pathogen Plasmodium vivax acquired genetic material from humans that might help facilitate its long stay in the body.
Human genome
- One study identified approximately 100 of humans' approximately 20,000 total genes which likely resulted from horizontal gene transfer, but this number has been challenged by several researchers arguing these candidate genes for HGT are more likely the result of gene loss combined with differences in the rate of evolution.
Compounds found to promote horizontal gene transfer
Through research into the growing issue of antibiotic resistance certain compounds have been observed to promote horizontal gene transfer. Antibiotics given to bacteria at non-lethal levels have been known to be a cause of antibiotic resistance but emerging research is now showing that certain non-antibiotic pharmaceuticals (ibuprofen, naproxen, gemfibrozil, diclofenac, propranolol, etc.) also have a role in promoting antibiotic resistance through their ability to promote horizontal gene transfer (HGT) of genes responsible for antibiotic resistance. The transfer of antibiotic resistance genes (ARGs) through conjugation is significantly accelerated when donor cells with plasmids and recipient cells are introduced to each other in the presence of one of the pharmaceuticals. Non-antibiotic pharmaceuticals were also found to cause some responses in bacteria similar to those responses to antibiotics, such as increasing expression of the genes lexA, umuC, umuD and soxR involved in the bacteria's SOS response as well as other genes also expressed during exposure to antibiotics. These findings are from 2021 and due to the widespread use of non-antibiotic pharmaceuticals, more research needs to be done in order to further understanding on the issue.
Alongside non-antibiotic pharmaceuticals, other compounds relevant to antibiotic resistance have been tested such as malachite green, ethylbenzene, styrene, 2,4-dichloroaniline, trioxymethylene, o-xylene solutions, p-nitrophenol (PNP), p-aminophenol (PAP), and phenol (PhOH). It is a global concern that ARGs have been found in wastewater treatment plants Textile wastewater has been found to contain 3- to 13-fold higher abundance of mobile genetic elements than other samples of wastewater. The cause of this is the organic compounds used for textile dying (o-xylene, ethylbenzene, trioxymethylene, styrene, 2,4-dichloroaniline, and malachite green) raising the frequency of conjugative transfer when bacteria and plasmid (with donor) are introduced in the presence of these molecules. When textile wastewater combines with wastewater from domestic sewage, the ARGs present in wastewater are transferred at a higher rate due to the addition of textile dyeing compounds increasing the occurrence of HGT.
Other organic pollutants commonly found in wastewater have been the subject of similar experiments. A 2021 study used similar methods of using plasmid in a donor and mixing that with a receptor in the presence of compound in order to test horizontal gene transfer of antibiotic resistance genes but this time in the presence of phenolic compounds. Phenolic compounds are commonly found in wastewater and have been found to change functions and structures of the microbial communities during the wastewater treatment process. Additionally, HGT increases in frequency in the presence of the compounds p-nitrophenol (PNP), p-aminophenol (PAP), and phenol. These compounds result in a 2- to 9-fold increase in HGT (p-nitrophenol being on the lower side of 2-fold increases and p-aminophenol and phenol having a maximum increase of 9-fold). This increase in HGT is on average less than the compounds ibuprofen, naproxen, gemfibrozil, diclofenac, propranolol, o-xylene, ethylbenzene, trioxymethylene, styrene, 2,4-dichloroaniline, and malachite green but their increases is still significant. The study that came to this conclusion is similar to the study on horizontal gene transfer and non-antibiotic pharmaceuticals in that it was done in 2021 and leaves room for more research, specifically in the focus of the study which is activated sludge.
Heavy metals have also been found to promote conjugative transfer of antibiotic resistance genes. The paper that led to the discovery of this was done in 2017 during the emerging field of horizontal gene transfer assisting compound research. Metals assist in the spread of antibiotic resistance through both co-resistance as well as cross-resistance mechanisms. In quantities relevant to the environment, Cu(II), Ag(I), Cr(VI), and Zn(II) promote HGT from donor and receptor strains of E. coli. The presence of these metals triggered SOS response from bacterial cells and made the cells more permeable. These are the mechanisms that make even low levels of heavy metal pollution in the environment impact HGT and therefore the spread of ARGs.
Promiscuous DNA
Promiscuous DNA is a form of horizontal gene transfer that transmits genetic information across organellar barriers. Promiscuous DNA transfer has substantial evidence in its movement across the genome of numerous organisms, from movements in chloroplast to the nucleus, chloroplast to the mitochondria, and mitochondria to the nucleus.
History
In 1982, R. John Ellis defined this type of transpositional transfer mutation as “intracellular promiscuity.” Ellis further explored the phenomenon of “intracellular promiscuity” through the experiments of David Stern and David Lonsdale, in which genetic transfer between chloroplasts to mitochondria was discovered, aiding in the definition and discovery of promiscuous DNA.
Mechanism
While much remains to be understood about how promiscuous DNA undergoes movement and transfer, numerous experiments have pointed to plastid sequences, ptDNA, as a key player. Plasmids, with their mobile nature and crucial role in horizontal gene transfer, are seen as a significant element in DNA that exchanges genetic information. This mobility makes ptDNA a potential donor for promiscuous DNA to traverse organellar barriers.
Types
NUMTs
NUMTs (nuclear sequences of mitochondrial) are a type of promiscuous DNA that arises from the natural transfer of mitochondria DNA (mtDNA) to the nuclear genome (nDNA). These NUMTs, with their varying frequencies, sizes, and features, contribute to the genetic diversity across all eukaryotes and, in some cases, to diseases among humans.
NUPTs
NUPTs (nuclear plastid DNA sequences) are a type of promiscuous DNA that arises from the natural transfer of ptDNA (plastid DNA) into nDNA. These fragments of ptDNA, similar to NUMTs in frequency, size, and features, also exhibit variability across species.
Artificial horizontal gene transfer
See also: Gene therapy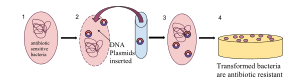
Genetic engineering is essentially horizontal gene transfer, albeit with synthetic expression cassettes. The Sleeping Beauty transposon system (SB) was developed as a synthetic gene transfer agent that was based on the known abilities of Tc1/mariner transposons to invade genomes of extremely diverse species. The SB system has been used to introduce genetic sequences into a wide variety of animal genomes.
In evolution
Main article: Horizontal gene transfer in evolutionHorizontal gene transfer is a potential confounding factor in inferring phylogenetic trees based on the sequence of one gene. For example, given two distantly related bacteria that have exchanged a gene a phylogenetic tree including those species will show them to be closely related because that gene is the same even though most other genes are dissimilar. For this reason, it is often ideal to use other information to infer robust phylogenies such as the presence or absence of genes or, more commonly, to include as wide a range of genes for phylogenetic analysis as possible.
For example, the most common gene to be used for constructing phylogenetic relationships in prokaryotes is the 16S ribosomal RNA gene since its sequences tend to be conserved among members with close phylogenetic distances, but variable enough that differences can be measured. However, in recent years it has also been argued that 16s rRNA genes can also be horizontally transferred. Although this may be infrequent, the validity of 16s rRNA-constructed phylogenetic trees must be reevaluated.
Biologist Johann Peter Gogarten suggests "the original metaphor of a tree no longer fits the data from recent genome research" therefore "biologists should use the metaphor of a mosaic to describe the different histories combined in individual genomes and use the metaphor of a net to visualize the rich exchange and cooperative effects of HGT among microbes". There exist several methods to infer such phylogenetic networks.
Using single genes as phylogenetic markers, it is difficult to trace organismal phylogeny in the presence of horizontal gene transfer. Combining the simple coalescence model of cladogenesis with rare HGT horizontal gene transfer events suggest there was no single most recent common ancestor that contained all of the genes ancestral to those shared among the three domains of life. Each contemporary molecule has its own history and traces back to an individual molecule cenancestor. However, these molecular ancestors were likely to be present in different organisms at different times."
Challenge to the tree of life
Further information: Last universal common ancestor and tree of life (science)Horizontal gene transfer poses a possible challenge to the concept of the last universal common ancestor (LUCA) at the root of the tree of life first formulated by Carl Woese, which led him to propose the Archaea as a third domain of life. Indeed, it was while examining the new three-domain view of life that horizontal gene transfer arose as a complicating issue: Archaeoglobus fulgidus was seen as an anomaly with respect to a phylogenetic tree based upon the encoding for the enzyme HMGCoA reductase—the organism in question is a definite Archaean, with all the cell lipids and transcription machinery that are expected of an Archaean, but whose HMGCoA genes are of bacterial origin. Scientists are broadly agreed on symbiogenesis, that mitochondria in eukaryotes derived from alpha-proteobacterial cells and that chloroplasts came from ingested cyanobacteria, and other gene transfers may have affected early eukaryotes. (In contrast, multicellular eukaryotes have mechanisms to prevent horizontal gene transfer, including separated germ cells.) If there had been continued and extensive gene transfer, there would be a complex network with many ancestors, instead of a tree of life with sharply delineated lineages leading back to a LUCA. However, a LUCA can be identified, so horizontal transfers must have been relatively limited.
Other early HGTs are thought to have happened. The first common ancestor (FUCA), earliest ancestor of LUCA, had other descendants that had their own lineages. These now-extinct sister lineages of LUCA descending from FUCA are thought to have horizontally transferred some of their genes into the genome of early descendants of LUCA.
Phylogenetic information in HGT
It has been remarked that, despite the complications, the detection of horizontal gene transfers brings valuable phylogenetic and dating information.
The potential of HGT to be used for dating phylogenies has recently been confirmed.
The chromosomal organization of horizontal gene transfer
The acquisition of new genes has the potential to disorganize the other genetic elements and hinder the function of the bacterial cell, thus affecting the competitiveness of bacteria. Consequently, bacterial adaptation lies in a conflict between the advantages of acquiring beneficial genes, and the need to maintain the organization of the rest of its genome. Horizontally transferred genes are typically concentrated in only ~1% of the chromosome (in regions called hotspots). This concentration increases with genome size and with the rate of transfer. Hotspots diversify by rapid gene turnover; their chromosomal distribution depends on local contexts (neighboring core genes), and content in mobile genetic elements. Hotspots concentrate most changes in gene repertoires, reduce the trade-off between genome diversification and organization, and should be treasure troves of strain-specific adaptive genes. Most mobile genetic elements and antibiotic resistance genes are in hotspots, but many hotspots lack recognizable mobile genetic elements and exhibit frequent homologous recombination at flanking core genes. Overrepresentation of hotspots with fewer mobile genetic elements in naturally transformable bacteria suggests that homologous recombination and horizontal gene transfer are tightly linked in genome evolution.
Genes
| This list is incomplete; you can help by adding missing items. (August 2008) |
There is evidence for historical horizontal transfer of the following genes:
- Lycopene cyclase for carotenoid biosynthesis, between Chlorobiota and "Cyanobacteria".
- TetO gene conferring resistance to tetracycline, between Campylobacter jejuni.
- Neochrome, a gene in some ferns that enhances their ability to survive in dim light. Believed to have been acquired from algae sometime during the Cretaceous.
- Transfer of a cysteine synthase from a bacterium into phytophagous mites and Lepidoptera allowing the detoxification of cyanogenic glucosides produced by host plants.
- The LINE1 sequence has transferred from humans to the gonorrhea bacteria.
See also
- Agrobacterium, a bacterium well known for its ability to transfer DNA between itself and plants.
- Endogenous retrovirus
- Genetically modified organism
- Inferring horizontal gene transfer
- Integron
- Mobile genetic elements
- Phylogenetic network
- Phylogenetic tree
- Provirus
- Reassortment
- Retrotransposon
- Symbiogenesis
- Tree of life (biology)
- Xenobiology
References
- Ochman H, Lawrence JG, Groisman EA (May 2000). "Lateral gene transfer and the nature of bacterial innovation". Nature. 405 (6784): 299–304. Bibcode:2000Natur.405..299O. doi:10.1038/35012500. PMID 10830951. S2CID 85739173.
- Dunning Hotopp JC (April 2011). "Horizontal gene transfer between bacteria and animals". Trends in Genetics. 27 (4): 157–63. doi:10.1016/j.tig.2011.01.005. PMC 3068243. PMID 21334091.
- Robinson KM, Sieber KB, Dunning Hotopp JC (October 2013). "A review of bacteria-animal lateral gene transfer may inform our understanding of diseases like cancer". PLOS Genetics. 9 (10): e1003877. doi:10.1371/journal.pgen.1003877. PMC 3798261. PMID 24146634.
- Keeling PJ, Palmer JD (August 2008). "Horizontal gene transfer in eukaryotic evolution". Nature Reviews. Genetics. 9 (8): 605–18. doi:10.1038/nrg2386. PMID 18591983. S2CID 213613.
- ^ Gyles C, Boerlin P (March 2014). "Horizontally transferred genetic elements and their role in pathogenesis of bacterial disease". Veterinary Pathology. 51 (2): 328–40. doi:10.1177/0300985813511131. PMID 24318976. S2CID 206510894.
- Vaux F, Trewick SA, Morgan-Richards M (2017). "Speciation through the looking-glass". Biological Journal of the Linnean Society. 120 (2): 480–488. doi:10.1111/bij.12872.
- Ochman H, Lerat E, Daubin V (May 2005). "Examining bacterial species under the specter of gene transfer and exchange". Proceedings of the National Academy of Sciences of the United States of America. 102 (Suppl 1): 6595–6599. Bibcode:2005PNAS..102.6595O. doi:10.1073/pnas.0502035102. PMC 1131874. PMID 15851673.
- Huddleston JR (2014). "Horizontal gene transfer in the human gastrointestinal tract: potential spread of antibiotic resistance genes". Infection and Drug Resistance. 7: 167–176. doi:10.2147/idr.s48820. PMC 4073975. PMID 25018641.
- Koonin EV, Makarova KS, Aravind L (2001). "Horizontal gene transfer in prokaryotes: quantification and classification". Annual Review of Microbiology. 55 (1): 709–42. doi:10.1146/annurev.micro.55.1.709. PMC 4781227. PMID 11544372.
- Nielsen KM (1998). "Barriers to horizontal gene transfer by natural transformation in soil bacteria". APMIS. 84 (S84): 77–84. doi:10.1111/j.1600-0463.1998.tb05653.x. PMID 9850687. S2CID 26490197.
- McGowan C, Fulthorpe R, Wright A, Tiedje JM (October 1998). "Evidence for interspecies gene transfer in the evolution of 2,4-dichlorophenoxyacetic acid degraders". Applied and Environmental Microbiology. 64 (10): 4089–92. Bibcode:1998ApEnM..64.4089M. doi:10.1128/AEM.64.10.4089-4092.1998. PMC 106609. PMID 9758850.
- ^ Keen EC (December 2012). "Paradigms of pathogenesis: targeting the mobile genetic elements of disease". Frontiers in Cellular and Infection Microbiology. 2: 161. doi:10.3389/fcimb.2012.00161. PMC 3522046. PMID 23248780.
- Naik GA, Bhat LN, Chpoade BA, Lynch JM (1994). "Transfer of broad-host-range antibiotic resistance plasmids in soil microcosms". Curr. Microbiol. 28 (4): 209–215. doi:10.1007/BF01575963. S2CID 21015053.
- Varga M, Kuntová L, Pantůček R, Mašlaňová I, Růžičková V, Doškař J (July 2012). "Efficient transfer of antibiotic resistance plasmids by transduction within methicillin-resistant Staphylococcus aureus USA300 clone". FEMS Microbiology Letters. 332 (2): 146–52. doi:10.1111/j.1574-6968.2012.02589.x. PMID 22553940.
- Varga M, Pantu Ček R, Ru Žičková V, Doškař J (January 2016). "Molecular characterization of a new efficiently transducing bacteriophage identified in meticillin-resistant Staphylococcus aureus". The Journal of General Virology. 97 (1): 258–268. doi:10.1099/jgv.0.000329. PMID 26537974.
- Cairns J, Ruokolainen L, Hultman J, Tamminen M, Virta M, Hiltunen T (2018-04-19). "Ecology determines how low antibiotic concentration impacts community composition and horizontal transfer of resistance genes". Communications Biology. 1 (1): 35. doi:10.1038/s42003-018-0041-7. PMC 6123812. PMID 30271921.
- Zhou H, Beltrán JF, Brito IL (October 2021). "Functions predict horizontal gene transfer and the emergence of antibiotic resistance". Science Advances. 7 (43): eabj5056. Bibcode:2021SciA....7.5056Z. doi:10.1126/sciadv.abj5056. PMC 8535800. PMID 34678056.
- Sieber KB, Bromley RE, Dunning Hotopp JC (September 2017). "Lateral gene transfer between prokaryotes and eukaryotes". Experimental Cell Research. 358 (2): 421–426. doi:10.1016/j.yexcr.2017.02.009. PMC 5550378. PMID 28189637.
- Gabaldón T (October 2021). "Origin and Early Evolution of the Eukaryotic Cell". Annual Review of Microbiology. 75 (1): 631–647. doi:10.1146/annurev-micro-090817-062213. PMID 34343017. S2CID 236916203.
- Brockhurst MA, Harrison E, Hall JP, Richards T, McNally A, MacLean C (October 2019). "The Ecology and Evolution of Pangenomes". Current Biology. 29 (20): R1094–R1103. Bibcode:2019CBio...29R1094B. doi:10.1016/j.cub.2019.08.012. ISSN 0960-9822. PMID 31639358.
- Van Etten J, Bhattacharya D (December 2020). "Horizontal Gene Transfer in Eukaryotes: Not if, but How Much?". Trends in Genetics. 36 (12): 915–925. Bibcode:2020TGene..36..915V. doi:10.1016/j.tig.2020.08.006. PMID 33012528.
- Kubyshkin V, Acevedo-Rocha CG, Budisa N (February 2018). "On universal coding events in protein biogenesis". Bio Systems. 164: 16–25. Bibcode:2018BiSys.164...16K. doi:10.1016/j.biosystems.2017.10.004. PMID 29030023.
- Griffith F (January 1928). "The Significance of Pneumococcal Types". The Journal of Hygiene. 27 (2). Cambridge University Press: 113–59. doi:10.1017/S0022172400031879. JSTOR 4626734. PMC 2167760. PMID 20474956.
- Lorenz MG, Wackernagel W (September 1994). "Bacterial gene transfer by natural genetic transformation in the environment". Microbiological Reviews. 58 (3): 563–602. doi:10.1128/MMBR.58.3.563-602.1994. PMC 372978. PMID 7968924.
- Downie AW (November 1972). "Pneumococcal transformation--a backward view. Fourth Griffith Memorial Lecture" (PDF). Journal of General Microbiology. 73 (1): 1–11. doi:10.1099/00221287-73-1-1. PMID 4143929. Archived (PDF) from the original on 2012-03-02. Retrieved 2018-05-23.
- Freeman VJ (June 1951). "Studies on the virulence of bacteriophage-infected strains of Corynebacterium diphtheriae". Journal of Bacteriology. 61 (6): 675–88. doi:10.1128/JB.61.6.675-688.1951. PMC 386063. PMID 14850426.
- Margulies P (2005). Diphtheria. Epidemics: Deadly diseases throughout history (1st ed.). New York: Rosen Publishing Group. ISBN 978-1-4042-0253-5.
- Lwoff A (1965). "Interaction among Virus, Cell, and Organism". Nobel Lecture for the Nobel Prize in Physiology or Medicine. Archived from the original on 2010-10-16.
- Ochiai K, Yamanaka T, Kimura K, Sawada O (1959). "Inheritance of drug resistance (and its transfer) between Shigella strains and Between Shigella and E. coli strains". Hihon Iji Shimpor (in Japanese). 1861: 34.
- Akiba T, Koyama K, Ishiki Y, Kimura S, Fukushima T (April 1960). "On the mechanism of the development of multiple-drug-resistant clones of Shigella". Japanese Journal of Microbiology. 4 (2): 219–27. doi:10.1111/j.1348-0421.1960.tb00170.x. PMID 13681921.
- Syvanen M (January 1985). "Cross-species gene transfer; implications for a new theory of evolution" (PDF). Journal of Theoretical Biology. 112 (2): 333–43. Bibcode:1985JThBi.112..333S. doi:10.1016/S0022-5193(85)80291-5. PMID 2984477. Archived (PDF) from the original on 2017-07-06. Retrieved 2009-01-13.
- Jain R, Rivera MC, Lake JA (March 1999). "Horizontal gene transfer among genomes: the complexity hypothesis". Proceedings of the National Academy of Sciences of the United States of America. 96 (7): 3801–6. Bibcode:1999PNAS...96.3801J. doi:10.1073/pnas.96.7.3801. PMC 22375. PMID 10097118.
- Rivera MC, Lake JA (September 2004). "The ring of life provides evidence for a genome fusion origin of eukaryotes" (PDF). Nature. 431 (7005): 152–5. Bibcode:2004Natur.431..152R. doi:10.1038/nature02848. PMID 15356622. S2CID 4349149. Archived from the original (PDF) on 2007-09-27.
- Bapteste E, Susko E, Leigh J, MacLeod D, Charlebois RL, Doolittle WF (May 2005). "Do orthologous gene phylogenies really support tree-thinking?". BMC Evolutionary Biology. 5 (1): 33. Bibcode:2005BMCEE...5...33B. doi:10.1186/1471-2148-5-33. PMC 1156881. PMID 15913459.
- Le Page M (2016-03-17). "Farmers may have been accidentally making GMOs for millennia". The New Scientist. Archived from the original on 2018-10-01. Retrieved 2016-07-11.
- Gasmi L, Boulain H, Gauthier J, Hua-Van A, Musset K, Jakubowska AK, et al. (September 2015). "Recurrent Domestication by Lepidoptera of Genes from Their Parasites Mediated by Bracoviruses". PLOS Genetics. 11 (9): e1005470. doi:10.1371/journal.pgen.1005470. PMC 4574769. PMID 26379286.
- Yong E (2010-02-14). "Genes from Chagas parasite can transfer to humans and be passed on to children". National Geographic. Archived from the original on January 6, 2013. Retrieved 2016-07-13.
Hecht MM, Nitz N, Araujo PF, Sousa AO, Rosa Ad, Gomes DA, et al. (12 February 2010). "Inheritance of DNA Transferred from American Trypanosomes to Human Hosts". PLOS ONE. 5 (2): e9181. Bibcode:2010PLoSO...5.9181H. doi:10.1371/journal.pone.0009181. PMC 2820539. PMID 20169193. - Riley DR, Sieber KB, Robinson KM, White JR, Ganesan A, Nourbakhsh S, et al. (2013). "Bacteria-human somatic cell lateral gene transfer is enriched in cancer samples". PLOS Computational Biology. 9 (6): e1003107. Bibcode:2013PLSCB...9E3107R. doi:10.1371/journal.pcbi.1003107. PMC 3688693. PMID 23840181.
- Richardson AO, Palmer JD (2007). "Horizontal gene transfer in plants" (PDF). Journal of Experimental Botany. 58 (1): 1–9. doi:10.1093/jxb/erl148. PMID 17030541. Archived from the original (PDF) on 2007-09-27.
- ^ Gogarten P (2000). "Horizontal Gene Transfer: A New Paradigm for Biology". Esalen Center for Theory and Research Conference. Archived from the original on 2012-07-21. Retrieved 2007-03-18.
- Todar K. "Bacterial Resistance to Antibiotics". The Microbial World: Lectures in Microbiology. Department of Bacteriology, University of Wisconsin-Madison. Archived from the original on January 15, 2012. Retrieved January 6, 2012.
- Maloy S (July 15, 2002). "Horizontal Gene Transfer". San Diego State University. Archived from the original on February 14, 2019. Retrieved January 6, 2012.
- ^ Stearns SC, Hoekstra RF (2005). Evolution: An introduction (2nd ed.). Oxford, New York: Oxford Univ. Press. pp. 38–40. ISBN 978-0-19-925563-4.
- Renner SS, Bellot S (2012). "Horizontal Gene Transfer in Eukaryotes: Fungi-to-Plant and Plant-to-Plant Transfers of Organellar DNA". Genomics of Chloroplasts and Mitochondria. Advances in Photosynthesis and Respiration. Vol. 35. Springer Science+Business Media B.V. pp. 223–235. doi:10.1007/978-94-007-2920-9_10. ISBN 978-94-007-2919-3.
- Maxmen A (2010). "Virus-like particles speed bacterial evolution". Nature. doi:10.1038/news.2010.507.
- ^ Schaack S, Gilbert C, Feschotte C (September 2010). "Promiscuous DNA: horizontal transfer of transposable elements and why it matters for eukaryotic evolution". Trends in Ecology & Evolution. 25 (9): 537–46. Bibcode:2010TEcoE..25..537S. doi:10.1016/j.tree.2010.06.001. PMC 2940939. PMID 20591532.
- ^ Dupeyron M, Leclercq S, Cerveau N, Bouchon D, Gilbert C (January 2014). "Horizontal transfer of transposons between and within crustaceans and insects". Mobile DNA. 5 (1): 4. doi:10.1186/1759-8753-5-4. PMC 3922705. PMID 24472097.
- ^ Aubin E, Llauro C, Garrigue J, Mirouze M, Panaud O, El Baidouri M (October 2023). "Genome-wide analysis of horizontal transfer in non-model wild species from a natural ecosystem reveals new insights into genetic exchange in plants". PLOS Genetics. 19 (10): e1010964. doi:10.1371/journal.pgen.1010964. PMC 10586619. PMID 37856455.
- ^ El Baidouri M, Carpentier MC, Cooke R, Gao D, Lasserre E, Llauro C, et al. (May 2014). "Widespread and frequent horizontal transfers of transposable elements in plants". Genome Research. 24 (5): 831–8. doi:10.1101/gr.164400.113. PMC 4009612. PMID 24518071.
- ^ Ivancevic AM, Walsh AM, Kortschak RD, Adelson DL (December 2013). "Jumping the fine LINE between species: horizontal transfer of transposable elements in animals catalyses genome evolution". BioEssays. 35 (12): 1071–82. doi:10.1002/bies.201300072. PMID 24003001. S2CID 6968210.
- ^ Wallau GL, Ortiz MF, Loreto EL (2012). "Horizontal transposon transfer in eukarya: detection, bias, and perspectives". Genome Biology and Evolution. 4 (8): 689–99. doi:10.1093/gbe/evs055. PMC 3516303. PMID 22798449.
- La Scola B, Desnues C, Pagnier I, Robert C, Barrassi L, Fournous G, et al. (September 2008). "The virophage as a unique parasite of the giant mimivirus". Nature. 455 (7209): 100–4. Bibcode:2008Natur.455..100L. doi:10.1038/nature07218. PMID 18690211. S2CID 4422249.
- Pearson H (August 2008). "'Virophage' suggests viruses are alive". Nature. 454 (7205): 677. Bibcode:2008Natur.454..677P. doi:10.1038/454677a. PMID 18685665. S2CID 205040157.
- Barlow M (2009). "What Antimicrobial Resistance Has Taught Us About Horizontal Gene Transfer". Horizontal Gene Transfer. Methods in Molecular Biology. Vol. 532. Totowa, NJ: Humana Press. pp. 397–411. doi:10.1007/978-1-60327-853-9_23. ISBN 978-1-60327-852-2. PMID 19271198.
- Hawkey PM, Jones AM (September 2009). "The changing epidemiology of resistance". The Journal of Antimicrobial Chemotherapy. 64 (Suppl 1): i3-10. doi:10.1093/jac/dkp256. PMID 19675017.
- Francino MP, ed. (2012). Horizontal Gene Transfer in Microorganisms. Caister Academic Press. ISBN 978-1-908230-10-2.
- Strauch E, Lurz R, Beutin L (December 2001). "Characterization of a Shiga toxin-encoding temperate bacteriophage of Shigella sonnei". Infection and Immunity. 69 (12): 7588–95. doi:10.1128/IAI.69.12.7588-7595.2001. PMC 98851. PMID 11705937.
- Johnson CM, Grossman AD (November 2015). "Integrative and Conjugative Elements (ICEs): What They Do and How They Work". Annual Review of Genetics. 42 (1): 577–601. doi:10.1146/annurev-genet-112414-055018. PMC 5180612. PMID 26473380.
- Oliveira PH, Touchon M, Rocha EP (September 2014). "The interplay of restriction-modification systems with mobile genetic elements and their prokaryotic hosts". Nucleic Acids Research. 49 (16): 10618–10631. doi:10.1093/nar/gku734. PMC 4176335. PMID 25120263.
- Auchtung JM, Lee CA, Garrison KL, Grossman AD (June 2007). "Identification and characterization of the immunity repressor (ImmR) that controls the mobile genetic element ICE Bs1 of Bacillus subtilis". PLOS Genet. 64 (6): 1515–1528. doi:10.1111/j.1365-2958.2007.05748.x. PMC 3320793. PMID 17511812.
- Tirumalai MR, Fox GE (September 2013). "An ICEBs1-like element may be associated with the extreme radiation and desiccation resistance of Bacillus pumilus SAFR-032 spores". Extremophiles. 17 (5): 767–774. doi:10.1007/s00792-013-0559-z. PMID 23812891. S2CID 8675124. Archived from the original on 2021-11-28. Retrieved 2020-09-16.
- Link L, Sawyer J, Venkateswaran K, Nicholson W (February 2004). "Extreme spore UV resistance of Bacillus pumilus isolates obtained from an ultraclean Spacecraft Assembly Facility". Microb Ecol. 47 (2): 159–163. Bibcode:2004MicEc..47..159L. doi:10.1007/s00248-003-1029-4. PMID 14502417. S2CID 13416635.
- Newcombe DA, Schuerger AC, Benardini JN, Dickinson D, Tanner R, Venkateswaran K (December 2005). "Survival of spacecraft-associated microorganisms under simulated martian UV irradiation". Appl Environ Microbiol. 71 (12): 8147–8156. Bibcode:2005ApEnM..71.8147N. doi:10.1128/AEM.71.12.8147-8156.2005. PMC 1317311. PMID 16332797.
- Kempf MJ, Chen F, Kern R, Venkateswaran K (June 2005). "Recurrent isolation of hydrogen peroxide-resistant spores of Bacillus pumilus from a spacecraft assembly facility". Astrobiology. 5 (3): 391–405. Bibcode:2005AsBio...5..391K. doi:10.1089/ast.2005.5.391. PMID 15941382. Archived from the original on 2022-03-07. Retrieved 2020-09-16.
- Biel SW, Hartl DL (June 1983). "Evolution of transposons: natural selection for Tn5 in Escherichia coli K12". Genetics. 103 (4): 581–592. doi:10.1093/genetics/103.4.581. PMC 1202041. PMID 6303898. Archived from the original on 2021-08-19. Retrieved 2020-09-16.
- Chao L, Vargas C, Spear BB, Cox EC (1983). "Transposable elements as mutator genes in evolution". Nature. 303 (5918): 633–635. Bibcode:1983Natur.303..633C. doi:10.1038/303633a0. PMC 1202041. PMID 6303898.
- Tirumalai MR, Karouia F, Tran Q, Stepanov VG, Bruce RJ, Ott M, et al. (May 2017). "The adaptation of Escherichia coli cells grown in simulated microgravity for an extended period is both phenotypic and genomic". npj Microgravity. 3 (15): 15. doi:10.1038/s41526-017-0020-1. PMC 5460176. PMID 28649637.
- Tirumalai MR, Karouia F, Tran Q, Stepanov VG, Bruce RJ, Ott M, et al. (January 2019). "Evaluation of acquired antibiotic resistance in Escherichia coli exposed to long-term low-shear modeled microgravity and background antibiotic exposure". mBio. 10 (e02637-18). doi:10.1128/mBio.02637-18. PMC 6336426. PMID 30647159.
- Ginolhac A, Jarrin C, Robe P, Perrière G, Vogel TM, Simonet P, et al. (June 2005). "Type I polyketide synthases may have evolved through horizontal gene transfer". Journal of Molecular Evolution. 60 (6): 716–25. Bibcode:2005JMolE..60..716G. doi:10.1007/s00239-004-0161-1. PMID 15909225.
- Jagannathan SV, Manemann EM, Rowe SE, Callender MC, Soto W (July 2021). "Marine Actinomycetes, New Sources of Biotechnological Products". Marine Drugs. 19 (7): 365. doi:10.3390/md19070365. ISSN 1660-3397. PMC 8304352. PMID 34201951.
- Gross H, Loper JE (November 2009). "Genomics of secondary metabolite production by Pseudomonas spp". Natural Product Reports. 26 (11): 1408–46. doi:10.1039/b817075b. PMID 19844639.
- Chen I, Dubnau D (March 2004). "DNA uptake during bacterial transformation". Nature Reviews. Microbiology. 2 (3): 241–9. doi:10.1038/nrmicro844. PMID 15083159. S2CID 205499369.
- ^ Johnsborg O, Eldholm V, Håvarstein LS (December 2007). "Natural genetic transformation: prevalence, mechanisms and function". Research in Microbiology. 158 (10): 767–78. doi:10.1016/j.resmic.2007.09.004. PMID 17997281.
- Solomon JM, Grossman AD (April 1996). "Who's competent and when: regulation of natural genetic competence in bacteria". Trends in Genetics. 12 (4): 150–5. doi:10.1016/0168-9525(96)10014-7. PMID 8901420.
- Michod RE, Bernstein H, Nedelcu AM (May 2008). "Adaptive value of sex in microbial pathogens" (PDF). Infection, Genetics and Evolution. 8 (3): 267–85. Bibcode:2008InfGE...8..267M. doi:10.1016/j.meegid.2008.01.002. PMID 18295550. Archived (PDF) from the original on 2020-05-11. Retrieved 2016-10-04.
- Keen EC, Bliskovsky VV, Malagon F, Baker JD, Prince JS, Klaus JS, et al. (January 2017). "Novel "Superspreader" Bacteriophages Promote Horizontal Gene Transfer by Transformation". mBio. 8 (1): e02115-16. doi:10.1128/mBio.02115-16. PMC 5241400. PMID 28096488.
- Cooke MB, Herman C (2023). "Conjugation's Toolkit: The Roles of Nonstructural Proteins in Bacterial Sex". Journal of Bacteriology. 205 (3): e0043822. doi:10.1128/jb.00438-22. PMC 10029717. PMID 36847532.
- Porter RD, Black S (April 1991). "The single-stranded-DNA-binding protein encoded by the Escherichia coli F factor can complement a deletion of the chromosomal ssb gene". Journal of Bacteriology. 173 (8): 2720–2723. doi:10.1128/jb.173.8.2720-2723.1991. ISSN 0021-9193. PMC 207845. PMID 2013585.
- Maffeo C, Aksimentiev A (2017-12-01). "Molecular mechanism of DNA association with single-stranded DNA binding protein". Nucleic Acids Research. 45 (21): 12125–12139. doi:10.1093/nar/gkx917. ISSN 0305-1048. PMC 5716091. PMID 29059392.
- Gupta S, Yeeles JT, Marians KJ (September 2014). "Regression of Replication Forks Stalled by Leading-strand Template Damage". Journal of Biological Chemistry. 289 (41): 28388–28398. doi:10.1074/jbc.M114.587907. PMC 4192491. PMID 25138217.
- Chen SH, Byrne-Nash RT, Cox MM (September 2016). "Escherichia coli RadD Protein Functionally Interacts with the Single-stranded DNA-binding Protein". Journal of Biological Chemistry. 291 (39): 20779–20786. doi:10.1074/jbc.M116.736223. PMC 5034066. PMID 27519413.
- Breidenstein A, Lamy A, Bader CP, Sun WS, Wanrooij PH, Berntsson RP (August 2024). "PrgE: an OB-fold protein from plasmid pCF10 with striking differences to prototypical bacterial SSBs". Life Science Alliance. 7 (8): e202402693. doi:10.26508/lsa.202402693. ISSN 2575-1077. PMC 11137577. PMID 38811160.
- Parsons LM, Jankowski CS, Derbyshire KM (April 1998). "Conjugal transfer of chromosomal DNA in Mycobacterium smegmatis". Molecular Microbiology. 28 (3): 571–582. doi:10.1046/j.1365-2958.1998.00818.x. ISSN 0950-382X. PMID 9632259.
- Supply P, Marceau M, Mangenot S, Roche D, Rouanet C, Khanna V, et al. (February 2013). "Genomic analysis of smooth tubercle bacilli provides insights into ancestry and pathoadaptation of Mycobacterium tuberculosis". Nature Genetics. 45 (2): 172–179. doi:10.1038/ng.2517. ISSN 1061-4036. PMC 3856870. PMID 23291586.
- Palmer KL, Kos VN, Gilmore MS (2010-10-01). "Horizontal gene transfer and the genomics of enterococcal antibiotic resistance". Current Opinion in Microbiology. Antimicrobials/Genomics. 13 (5): 632–639. doi:10.1016/j.mib.2010.08.004. ISSN 1369-5274. PMC 2955785. PMID 20837397.
- Sommer MO, Dantas G, Church GM (2009-08-28). "Functional Characterization of the Antibiotic Resistance Reservoir in the Human Microflora". Science. 325 (5944): 1128–1131. Bibcode:2009Sci...325.1128S. doi:10.1126/science.1176950. ISSN 0036-8075. PMC 4720503. PMID 19713526.
- ^ Gray TA, Krywy JA, Harold J, Palumbo MJ, Derbyshire KM (July 2013). "Distributive conjugal transfer in mycobacteria generates progeny with meiotic-like genome-wide mosaicism, allowing mapping of a mating identity locus". PLOS Biology. 11 (7): e1001602. doi:10.1371/journal.pbio.1001602. PMC 3706393. PMID 23874149.
- Derbyshire KM, Gray TA (2014). "Distributive Conjugal Transfer: New Insights into Horizontal Gene Transfer and Genetic Exchange in Mycobacteria". Microbiology Spectrum. 2 (1): 61–79. doi:10.1128/microbiolspec.MGM2-0022-2013. PMC 4259119. PMID 25505644.
- Méheust R, Watson AK, Lapointe FJ, Papke RT, Lopez P, Bapteste E (June 2018). "Hundreds of novel composite genes and chimeric genes with bacterial origins contributed to haloarchaeal evolution". Genome Biology. 19 (1): 75. Bibcode:2018GenBi..19...75M. doi:10.1186/s13059-018-1454-9. PMC 5992828. PMID 29880023.
- Martijn J, Schön ME, Lind AE, Vosseberg J, Williams TA, Spang A, et al. (October 2020). "Hikarchaeia demonstrate an intermediate stage in the methanogen-to-halophile transition". Nature Communications. 11 (1): 5490. Bibcode:2020NatCo..11.5490M. doi:10.1038/s41467-020-19200-2. PMC 7599335. PMID 33127909.
- ^ Fröls S, Ajon M, Wagner M, Teichmann D, Zolghadr B, Folea M, et al. (November 2008). "UV-inducible cellular aggregation of the hyperthermophilic archaeon Sulfolobus solfataricus is mediated by pili formation" (PDF). Molecular Microbiology. 70 (4): 938–52. doi:10.1111/j.1365-2958.2008.06459.x. PMID 18990182. Archived (PDF) from the original on 2023-04-15. Retrieved 2020-06-23.
- Allers T (November 2011). "Swapping genes to survive - a new role for archaeal type IV pili". Molecular Microbiology. 82 (4): 789–91. doi:10.1111/j.1365-2958.2011.07860.x. PMID 21992544. S2CID 45490029.
- ^ Ajon M, Fröls S, van Wolferen M, Stoecker K, Teichmann D, Driessen AJ, et al. (November 2011). "UV-inducible DNA exchange in hyperthermophilic archaea mediated by type IV pili" (PDF). Molecular Microbiology. 82 (4): 807–17. doi:10.1111/j.1365-2958.2011.07861.x. PMID 21999488. Archived (PDF) from the original on 2021-10-10. Retrieved 2020-06-23.
- Fröls S, White MF, Schleper C (February 2009). "Reactions to UV damage in the model archaeon Sulfolobus solfataricus". Biochemical Society Transactions. 37 (Pt 1): 36–41. doi:10.1042/BST0370036. PMID 19143598.
- Grogan DW (June 1996). "Exchange of genetic markers at extremely high temperatures in the archaeon Sulfolobus acidocaldarius". Journal of Bacteriology. 178 (11): 3207–11. doi:10.1128/jb.178.11.3207-3211.1996. PMC 178072. PMID 8655500.
- Wood ER, Ghané F, Grogan DW (September 1997). "Genetic responses of the thermophilic archaeon Sulfolobus acidocaldarius to short-wavelength UV light". Journal of Bacteriology. 179 (18): 5693–8. doi:10.1128/jb.179.18.5693-5698.1997. PMC 179455. PMID 9294423.
- Reilly MS, Grogan DW (February 2002). "Biological effects of DNA damage in the hyperthermophilic archaeon Sulfolobus acidocaldarius". FEMS Microbiology Letters. 208 (1): 29–34. doi:10.1016/s0378-1097(01)00575-4. PMID 11934490.
- ^ van Wolferen M, Ajon M, Driessen AJ, Albers SV (December 2013). "Molecular analysis of the UV-inducible pili operon from Sulfolobus acidocaldarius". MicrobiologyOpen. 2 (6): 928–37. doi:10.1002/mbo3.128. PMC 3892339. PMID 24106028.
- van Wolferen M, Ma X, Albers SV (September 2015). "DNA Processing Proteins Involved in the UV-Induced Stress Response of Sulfolobales". Journal of Bacteriology. 197 (18): 2941–51. doi:10.1128/JB.00344-15. PMC 4542170. PMID 26148716.
- Melcher U (2001). "Molecular genetics: Horizontal gene transfer". Stillwater, Oklahoma USA: Oklahoma State University. Archived from the original on 2016-03-04. Retrieved 2015-08-20.
- Blanchard JL, Lynch M (July 2000). "Organellar genes: why do they end up in the nucleus?". Trends in Genetics. 16 (7): 315–20. doi:10.1016/S0168-9525(00)02053-9. PMID 10858662. Discusses theories on how mitochondria and chloroplast genes are transferred into the nucleus, and also what steps a gene needs to go through in order to complete this process.
- Davis CC, Wurdack KJ (July 2004). "Host-to-parasite gene transfer in flowering plants: phylogenetic evidence from Malpighiales". Science. 305 (5684): 676–8. Bibcode:2004Sci...305..676D. doi:10.1126/science.1100671. PMID 15256617. S2CID 16180594.
- Nickrent DL, Blarer A, Qiu YL, Vidal-Russell R, Anderson FE (October 2004). "Phylogenetic inference in Rafflesiales: the influence of rate heterogeneity and horizontal gene transfer". BMC Evolutionary Biology. 4 (1): 40. doi:10.1186/1471-2148-4-40. PMC 528834. PMID 15496229.
- Woloszynska M, Bocer T, Mackiewicz P, Janska H (November 2004). "A fragment of chloroplast DNA was transferred horizontally, probably from non-eudicots, to mitochondrial genome of Phaseolus". Plant Molecular Biology. 56 (5): 811–20. doi:10.1007/s11103-004-5183-y. PMID 15803417. S2CID 14198321.
- Hall C, Brachat S, Dietrich FS (June 2005). "Contribution of horizontal gene transfer to the evolution of Saccharomyces cerevisiae". Eukaryotic Cell. 4 (6): 1102–15. doi:10.1128/EC.4.6.1102-1115.2005. PMC 1151995. PMID 15947202.
- Quispe-Huamanquispe DG, Gheysen G, Kreuze JF (2017). "Agrobacterium T-DNAs". Frontiers in Plant Science. 8: 2015. doi:10.3389/fpls.2017.02015. PMC 5705623. PMID 29225610.
- Kfoury B, Rodrigues WF, Kim SJ, Brandizzi F, Del-Bem LE (2024). "Multiple horizontal gene transfer events have shaped plant glycosyl hydrolase diversity and function". New Phytologist. 242 (2): 809–824. Bibcode:2024NewPh.242..809K. doi:10.1111/nph.19595. PMID 38417454.
- Lee Phillips M (2012). "Bacterial gene helps coffee beetle get its fix". Nature. doi:10.1038/nature.2012.10116. S2CID 211729274.
- Acuña R, Padilla BE, Flórez-Ramos CP, Rubio JD, Herrera JC, Benavides P, et al. (March 2012). "Adaptive horizontal transfer of a bacterial gene to an invasive insect pest of coffee". Proceedings of the National Academy of Sciences of the United States of America. 109 (11): 4197–202. Bibcode:2012PNAS..109.4197A. doi:10.1073/pnas.1121190109. PMC 3306691. PMID 22371593.
- Blondel L, Jones ET, Extavour GC (Feb 2020). "Bacterial contribution to genesis of the novel germ line determinant oskar". eLife. 24 (9): e45539. doi:10.7554/eLife.45539. PMC 7250577. PMID 32091394.
- Watson T (15 November 2012). "Bdelloids Surviving on Borrowed DNA". Science/AAAS News. Archived from the original on 6 May 2023. Retrieved 30 June 2022.
- Koutsovoulos G, Kumar S, Laetsch DR, Stevens L, Daub J, Conlon C, et al. (May 2016). "No evidence for extensive horizontal gene transfer in the genome of the tardigrade Hypsibius dujardini". Proceedings of the National Academy of Sciences of the United States of America. 113 (18): 5053–8. Bibcode:2016PNAS..113.5053K. doi:10.1073/pnas.1600338113. PMC 4983863. PMID 27035985.
- Crisp A, Boschetti C, Perry M, Tunnacliffe A, Micklem G (March 2015). "Expression of multiple horizontally acquired genes is a hallmark of both vertebrate and invertebrate genomes". Genome Biology. 16 (1): 50. doi:10.1186/s13059-015-0607-3. PMC 4358723. PMID 25785303.
- Madhusoodanan J (2015-03-12). "Horizontal Gene Transfer a Hallmark of Animal Genomes?". The Scientist. Archived from the original on 2016-07-09. Retrieved 2016-07-14.
- Daugavet MA, Shabelnikov S, Shumeev A, Shaposhnikova T, Adonin LS, Podgornaya O (2019-01-19). "Features of a novel protein, rusticalin, from the ascidian Styela rustica reveal ancestral horizontal gene transfer event". Mobile DNA. 10 (1): 4. doi:10.1186/s13100-019-0146-7. PMC 6339383. PMID 30675192.
- Redrejo-Rodríguez M, Muñoz-Espín D, Holguera I, Mencía M, Salas M (November 2012). "Functional eukaryotic nuclear localization signals are widespread in terminal proteins of bacteriophages". Proceedings of the National Academy of Sciences of the United States of America. 109 (45): 18482–7. Bibcode:2012PNAS..10918482R. doi:10.1073/pnas.1216635109. PMC 3494942. PMID 23091024.
- Kondo N, Nikoh N, Ijichi N, Shimada M, Fukatsu T (October 2002). "Genome fragment of Wolbachia endosymbiont transferred to X chromosome of host insect". Proceedings of the National Academy of Sciences of the United States of America. 99 (22): 14280–5. Bibcode:2002PNAS...9914280K. doi:10.1073/pnas.222228199. PMC 137875. PMID 12386340.
- Dunning Hotopp JC, Clark ME, Oliveira DC, Foster JM, Fischer P, Muñoz Torres MC, et al. (September 2007). "Widespread lateral gene transfer from intracellular bacteria to multicellular eukaryotes". Science. 317 (5845): 1753–6. Bibcode:2007Sci...317.1753H. doi:10.1126/science.1142490. PMID 17761848. S2CID 10787254.
- Sloan, D. B., Nakabachi, A., Richards, S., Qu, J., Murali, S. C., Gibbs, R. A., & Moran, N. A. (2014). Parallel histories of horizontal gene transfer facilitated extreme reduction of endosymbiont genomes in sap-feeding insects. Molecular biology and evolution, 31(4), 857-871.
- Yoshida S, Maruyama S, Nozaki H, Shirasu K (May 2010). "Horizontal gene transfer by the parasitic plant Striga hermonthica". Science. 328 (5982): 1128. Bibcode:2010Sci...328.1128Y. doi:10.1126/science.1187145. PMID 20508124. S2CID 39376164.
- Zimmer C (April 17, 2014). "Plants That Practice Genetic Engineering". New York Times. Archived from the original on December 26, 2022. Retrieved February 27, 2017.
- David‐Schwartz R, Runo S, Townsley B, Machuka J, Sinha N. 2008. Long‐distance transport of mRNA via parenchyma cells and phloem across the host–parasite junction in Cuscuta. New Phytologist 179: 1133– 1141.
- Schwartz JA, Curtis NE, Pierce SK (December 2014). "FISH labeling reveals a horizontally transferred algal (Vaucheria litorea) nuclear gene on a sea slug (Elysia chlorotica) chromosome". The Biological Bulletin. 227 (3): 300–12. doi:10.1086/BBLv227n3p300. PMID 25572217. S2CID 21742354.
- Rauch C, Vries J, Rommel S, Rose LE, Woehle C, Christa G, et al. (August 2015). "Why It Is Time to Look Beyond Algal Genes in Photosynthetic Slugs". Genome Biology and Evolution. 7 (9): 2602–7. doi:10.1093/gbe/evv173. PMC 4607529. PMID 26319575.
- Bhattacharya D, Pelletreau KN, Price DC, Sarver KE, Rumpho ME (August 2013). "Genome analysis of Elysia chlorotica Egg DNA provides no evidence for horizontal gene transfer into the germ line of this Kleptoplastic Mollusc". Molecular Biology and Evolution. 30 (8): 1843–52. doi:10.1093/molbev/mst084. PMC 3708498. PMID 23645554.
- Xia J, Guo Z, Yang Z, Han H, Wang S, Xu H, et al. (April 2021). "Whitefly hijacks a plant detoxification gene that neutralizes plant toxins". Cell. 184 (7): 1693–1705.e17. doi:10.1016/j.cell.2021.02.014. PMID 33770502. S2CID 232359463.
- Armijos Jaramillo VD, Vargas WA, Sukno SA, Thon MR (November 2013). "New insights into the evolution and structure of Colletotrichum plant-like subtilisins (CPLSs)". Communicative & Integrative Biology. 6 (6): e25727. doi:10.4161/cib.25727. PMC 3917961. PMID 24563701.
- ^ Nikolaidis N, Doran N, Cosgrove DJ (February 2014). "Plant expansins in bacteria and fungi: evolution by horizontal gene transfer and independent domain fusion". Molecular Biology and Evolution. 31 (2): 376–86. doi:10.1093/molbev/mst206. PMID 24150040.
- ^ Moran NA, Jarvik T (April 2010). "Lateral transfer of genes from fungi underlies carotenoid production in aphids". Science. 328 (5978): 624–7. Bibcode:2010Sci...328..624M. doi:10.1126/science.1187113. PMID 20431015. S2CID 14785276.
- Fukatsu T (April 2010). "Evolution. A fungal past to insect color". Science. 328 (5978): 574–5. Bibcode:2010Sci...328..574F. doi:10.1126/science.1190417. PMID 20431000. S2CID 23686682.
- Luo H, Hallen-Adams HE, Lüli Y, Sgambelluri RM, Li X, Smith M, et al. (May 2022). "Genes and evolutionary fates of the amanitin biosynthesis pathway in poisonous mushrooms". Proceedings of the National Academy of Sciences of the United States of America. 119 (20): e2201113119. Bibcode:2022PNAS..11901113L. doi:10.1073/pnas.2201113119. PMC 9171917. PMID 35533275. S2CID 248668772.
- McDonald MC, Taranto AP, Hill E, Schwessinger B, Liu Z, Simpfendorfer S, et al. (2019-10-29). Di Pietro A (ed.). "Transposon-Mediated Horizontal Transfer of the Host-Specific Virulence Protein ToxA between Three Fungal Wheat Pathogens". mBio. 10 (5). doi:10.1128/mBio.01515-19. ISSN 2161-2129. PMC 6737239. PMID 31506307.
- Cheeseman K, Ropars J, Renault P, Dupont J, Gouzy J, Branca A, et al. (2014-01-10). "Multiple recent horizontal transfers of a large genomic region in cheese making fungi". Nature Communications. 5 (1): 2876. Bibcode:2014NatCo...5.2876C. doi:10.1038/ncomms3876. ISSN 2041-1723. PMC 3896755. PMID 24407037.
- Richards TA, Dacks JB, Jenkinson JM, Thornton CR, Talbot NJ (September 2006). "Evolution of filamentous plant pathogens: gene exchange across eukaryotic kingdoms". Current Biology. 16 (18): 1857–1864. Bibcode:2006CBio...16.1857R. doi:10.1016/j.cub.2006.07.052. PMID 16979565.
- Richards TA, Soanes DM, Jones MD, Vasieva O, Leonard G, Paszkiewicz K, et al. (September 2011). "Horizontal gene transfer facilitated the evolution of plant parasitic mechanisms in the oomycetes". Proceedings of the National Academy of Sciences of the United States of America. 108 (37): 15258–15263. Bibcode:2011PNAS..10815258R. doi:10.1073/pnas.1105100108. PMC 3174590. PMID 21878562.
- Wilcox C (2021-06-09). "DNA Jumps Between Animal Species. No One Knows How Often". Quanta Magazine. Archived from the original on 2021-06-14. Retrieved 2021-06-15.
- Martin JP, Fridovich I (June 1981). "Evidence for a natural gene transfer from the ponyfish to its bioluminescent bacterial symbiont Photobacter leiognathi. The close relationship between bacteriocuprein and the copper-zinc superoxide dismutase of teleost fishes". The Journal of Biological Chemistry. 256 (12): 6080–6089. doi:10.1016/S0021-9258(19)69131-3. PMID 6787049.
- Bar D (16 February 2011). "Evidence of Massive Horizontal Gene Transfer Between Humans and Plasmodium vivax". Nature Precedings. doi:10.1038/npre.2011.5690.1. Archived from the original on 31 March 2019. Retrieved 13 May 2011.
- "Human beings' ancestors have routinely stolen genes from other species". The Economist. 14 March 2015. Archived from the original on 16 March 2015. Retrieved 17 March 2015.
- Andersson DI, Hughes D (July 2014). "Microbiological effects of sublethal levels of antibiotics". Nature Reviews. Microbiology. 12 (7): 465–478. doi:10.1038/nrmicro3270. PMID 24861036. S2CID 3351736.
- ^ Wang Y, Lu J, Zhang S, Li J, Mao L, Yuan Z, et al. (September 2021). "Non-antibiotic pharmaceuticals promote the transmission of multidrug resistance plasmids through intra- and intergenera conjugation". The ISME Journal. 15 (9): 2493–2508. Bibcode:2021ISMEJ..15.2493W. doi:10.1038/s41396-021-00945-7. PMC 8397710. PMID 33692486.
- ^ Jiao YN, Chen H, Gao RX, Zhu YG, Rensing C (October 2017). "Organic compounds stimulate horizontal transfer of antibiotic resistance genes in mixed wastewater treatment systems". Chemosphere. 184: 53–61. Bibcode:2017Chmsp.184...53J. doi:10.1016/j.chemosphere.2017.05.149. PMID 28578196.
- ^ Ma X, Zhang X, Xia J, Sun H, Zhang X, Ye L (December 2021). "Phenolic compounds promote the horizontal transfer of antibiotic resistance genes in activated sludge". The Science of the Total Environment. 800: 149549. Bibcode:2021ScTEn.80049549M. doi:10.1016/j.scitotenv.2021.149549. PMID 34392203.
- ^ Zhang Y, Gu AZ, Cen T, Li X, He M, Li D, et al. (June 2018). "Sub-inhibitory concentrations of heavy metals facilitate the horizontal transfer of plasmid-mediated antibiotic resistance genes in water environment". Environmental Pollution. 237: 74–82. Bibcode:2018EPoll.237...74Z. doi:10.1016/j.envpol.2018.01.032. PMID 29477117. S2CID 4911120.
- Schaack S, Gilbert C, Feschotte C (2010-09-01). "Promiscuous DNA: horizontal transfer of transposable elements and why it matters for eukaryotic evolution". Trends in Ecology & Evolution. 25 (9): 537–546. Bibcode:2010TEcoE..25..537S. doi:10.1016/j.tree.2010.06.001. ISSN 0169-5347. PMC 2940939. PMID 20591532.
- Stegemann S, Hartmann S, Ruf S, Bock R (2003-07-22). "High-frequency gene transfer from the chloroplast genome to the nucleus". Proceedings of the National Academy of Sciences. 100 (15): 8828–8833. Bibcode:2003PNAS..100.8828S. doi:10.1073/pnas.1430924100. PMC 166398. PMID 12817081.
- Cerutti H, Jagendorf A (1995-11-01). "Movement of DNA across the chloroplast envelope: Implications for the transfer of promiscuous DNA". Photosynthesis Research. 46 (1): 329–337. Bibcode:1995PhoRe..46..329C. doi:10.1007/BF00020448. ISSN 1573-5079. PMID 24301600.
- ^ Sacerdot C, Casaregola S, Lafontaine I, Tekaia F, Dujon B, Ozier-Kalogeropoulos O (1 September 2008). "Promiscuous DNA in the nuclear genomes of hemiascomycetous yeasts". academic.oup.com.
- Ellis J (October 1982). "Promiscuous DNA—chloroplast genes inside plant mitochondria". Nature. 299 (5885): 678–679. Bibcode:1982Natur.299..678E. doi:10.1038/299678a0. ISSN 1476-4687. PMID 7121600.
- Stern DB, Lonsdale DM (October 1982). "Mitochondrial and chloroplast genomes of maize have a 12-kilobase DNA sequence in common". Nature. 299 (5885): 698–702. Bibcode:1982Natur.299..698S. doi:10.1038/299698a0. ISSN 1476-4687. PMID 6889685.
- Zeltz P, Kadowaki Ki, Kubo N, Maier RM, Hirai A, Kössel H (1996-06-01). "A promiscuous chloroplast DNA fragment is transcribed in plant mitochondria but the encoded RNA is not edited". Plant Molecular Biology. 31 (3): 647–656. doi:10.1007/BF00042236. ISSN 1573-5028. PMID 8790296.
- Park HS, Jayakodi M, Lee SH, Jeon JH, Lee HO, Park JY, et al. (2020-04-09). "Mitochondrial plastid DNA can cause DNA barcoding paradox in plants". Scientific Reports. 10 (1): 6112. Bibcode:2020NatSR..10.6112P. doi:10.1038/s41598-020-63233-y. ISSN 2045-2322. PMC 7145815. PMID 32273595.
- Ayliffe MA, Scott NS, Timmis JN (1 June 1998). ""Analysis of plastid DNA-like sequences within the nuclear genomes of higher plants."". Molecular Biology and Evolution. 15 (6): 738–745. doi:10.1093/oxfordjournals.molbev.a025977. PMID 9615455 – via Oxford Academic.
- Suzuki H, Yano H, Brown CJ, Top EM (27 September 2010). "Predicting Plasmid Promiscuity Based on Genomic Signature". Journal of Bacteriology. 192 (22): 6045–6055. doi:10.1128/JB.00277-10. PMC 2976448. PMID 20851899.
- Cerutti H, Jagendorf A (1995-11-01). "Movement of DNA across the chloroplast envelope: Implications for the transfer of promiscuous DNA". Photosynthesis Research. 46 (1): 329–337. Bibcode:1995PhoRe..46..329C. doi:10.1007/BF00020448. ISSN 1573-5079. PMID 24301600.
- Harutyunyan T (7 October 2023). "The known unknowns of mitochondrial carcinogenesis: de novo NUMTs and intercellular mitochondrial transfer". Oxford Academic.
- Namasivayam S, Sun C, Bah AB, Oberstaller J, Pierre-Louis E, Etheridge RD, et al. (2023-11-07). "Massive invasion of organellar DNA drives nuclear genome evolution in Toxoplasma". Proceedings of the National Academy of Sciences. 120 (45): e2308569120. Bibcode:2023PNAS..12008569N. doi:10.1073/pnas.2308569120. PMC 10636329. PMID 37917792.
- Michalovova M, Vyskot B, Kejnovsky E (October 2013). "Analysis of plastid and mitochondrial DNA insertions in the nucleus (NUPTs and NUMTs) of six plant species: size, relative age and chromosomal localization". Heredity. 111 (4): 314–320. doi:10.1038/hdy.2013.51. ISSN 1365-2540. PMC 3807264. PMID 23715017.
- Ivics Z, Hackett PB, Plasterk RH, Izsvák Z (November 1997). "Molecular reconstruction of Sleeping Beauty, a Tc1-like transposon from fish, and its transposition in human cells". Cell. 91 (4): 501–510. doi:10.1016/S0092-8674(00)80436-5. PMID 9390559. S2CID 17908472.
- Plasterk RH (1996). "The Tc1/Mariner Transposon Family". In Saedler H, Gierl A (eds.). Transposable Elements. Current Topics in Microbiology and Immunology. Vol. 204. Berlin, Heidelberg: Springer. pp. 125–143. doi:10.1007/978-3-642-79795-8_6. ISBN 978-3-642-79797-2. PMID 8556864.
- Izsvák Z, Ivics Z, Plasterk RH (September 2000). "Sleeping Beauty, a wide host-range transposon vector for genetic transformation in vertebrates". Journal of Molecular Biology. 302 (1): 93–102. doi:10.1006/jmbi.2000.4047. PMID 10964563.
- Kurtti TJ, Mattila JT, Herron MJ, Felsheim RF, Baldridge GD, Burkhardt NY, et al. (October 2008). "Transgene expression and silencing in a tick cell line: A model system for functional tick genomics". Insect Biochemistry and Molecular Biology. 38 (10): 963–968. Bibcode:2008IBMB...38..963K. doi:10.1016/j.ibmb.2008.07.008. PMC 2581827. PMID 18722527.
- Lawton G (21 January 2009). "Why Darwin was wrong about the tree of life". New Scientist Magazine. Archived from the original on 2015-04-14.
- Badger JH, Eisen JA, Ward NL (May 2005). "Genomic analysis of Hyphomonas neptunium contradicts 16S rRNA gene-based phylogenetic analysis: implications for the taxonomy of the orders 'Rhodobacterales' and Caulobacterales". International Journal of Systematic and Evolutionary Microbiology. 55 (Pt 3): 1021–1026. doi:10.1099/ijs.0.63510-0. PMID 15879228.
- Zhaxybayeva O, Gogarten JP (April 2004). "Cladogenesis, coalescence and the evolution of the three domains of life". Trends in Genetics. 20 (4): 182–7. doi:10.1016/j.tig.2004.02.004. PMID 15041172.
- ^ Doolittle WF (February 2000). "Uprooting the tree of life". Scientific American. 282 (2): 90–5. Bibcode:2000SciAm.282b..90D. doi:10.1038/scientificamerican0200-90. PMID 10710791.
- Woese CR (June 2004). "A new biology for a new century". Microbiology and Molecular Biology Reviews. 68 (2): 173–86. doi:10.1128/MMBR.68.2.173-186.2004. PMC 419918. PMID 15187180.
- Theobald DL (May 2010). "A formal test of the theory of universal common ancestry". Nature. 465 (7295): 219–22. Bibcode:2010Natur.465..219T. doi:10.1038/nature09014. PMID 20463738. S2CID 4422345.
- ^ Harris HM, Hill C (2021). "A Place for Viruses on the Tree of Life". Frontiers in Microbiology. 11: 604048. doi:10.3389/fmicb.2020.604048. PMC 7840587. PMID 33519747.
- Huang J, Gogarten JP (2009). "Ancient Gene Transfer as a Tool in Phylogenetic Reconstruction". Horizontal Gene Transfer. Methods in Molecular Biology. Vol. 532. Humana Press. pp. 127–39. doi:10.1007/978-1-60327-853-9_7. ISBN 978-1-60327-852-2. PMID 19271182.
- Davín AA, Tannier E, Williams TA, Boussau B, Daubin V, Szöllősi GJ (May 2018). "Gene transfers can date the tree of life". Nature Ecology & Evolution. 2 (5): 904–909. Bibcode:2018NatEE...2..904D. doi:10.1038/s41559-018-0525-3. PMC 5912509. PMID 29610471.
- Wolfe JM, Fournier GP (May 2018). "Horizontal gene transfer constrains the timing of methanogen evolution". Nature Ecology & Evolution. 2 (5): 897–903. Bibcode:2018NatEE...2..897W. doi:10.1038/s41559-018-0513-7. hdl:1721.1/118329. PMID 29610466. S2CID 4968981.
- Oliveira PH, Touchon M, Cury J, Rocha EP (October 2017). "The chromosomal organization of horizontal gene transfer in bacteria". Nature Communications. 8 (1): 841. Bibcode:2017NatCo...8..841O. doi:10.1038/s41467-017-00808-w. PMC 5635113. PMID 29018197.
- Bryant DA, Frigaard NU (November 2006). "Prokaryotic photosynthesis and phototrophy illuminated". Trends in Microbiology. 14 (11): 488–96. doi:10.1016/j.tim.2006.09.001. PMID 16997562.
- Avrain L, Vernozy-Rozand C, Kempf I (2004). "Evidence for natural horizontal transfer of tetO gene between Campylobacter jejuni strains in chickens". Journal of Applied Microbiology. 97 (1): 134–40. doi:10.1111/j.1365-2672.2004.02306.x. PMID 15186450. S2CID 19184139.
- Darkened Forests, Ferns Stole Gene From an Unlikely Source — and Then From Each Other Archived 2016-03-07 at the Wayback Machine by Jennifer Frazer (May 6, 2014). Scientific American.
- Li FW, Rothfels CJ, Melkonian M, Villarreal JC, Stevenson DW, Graham SW, et al. (2015). "The origin and evolution of phototropins". Frontiers in Plant Science. 6: 637. doi:10.3389/fpls.2015.00637. PMC 4532919. PMID 26322073.
- Wybouw N, Dermauw W, Tirry L, Stevens C, Grbić M, Feyereisen R, et al. (April 2014). "A gene horizontally transferred from bacteria protects arthropods from host plant cyanide poisoning". eLife. 3: e02365. doi:10.7554/eLife.02365. PMC 4011162. PMID 24843024.
- Yong E (2011-02-16). "Gonorrhea has picked up human DNA (and that's just the beginning)". National Geographic. Archived from the original on January 6, 2013. Retrieved 2016-07-14.
Further reading
- Quammen D (2018). The Tangled Tree: A Radical New History of Life. Simon & Schuster. ISBN 978-1-4767-7662-0.
- Gyles C, Boerlin P (March 2014). "Horizontally transferred genetic elements and their role in pathogenesis of bacterial disease". Veterinary Pathology. 51 (2): 328–40. doi:10.1177/0300985813511131. PMID 24318976. S2CID 206510894.
- – Papers by Dr Michael Syvanen on Horizontal Gene Transfer Archived 2022-02-27 at the Wayback Machine
- Salzberg SL, White O, Peterson J, Eisen JA (June 2001). "Microbial genes in the human genome: lateral transfer or gene loss?" (PDF). Science. 292 (5523): 1903–6. Bibcode:2001Sci...292.1903S. doi:10.1126/science.1061036. PMID 11358996. S2CID 17016011. Archived from the original (PDF) on 2006-09-01. Retrieved 2005-12-29.
About 40 genes were found to be exclusively shared by humans and bacteria and are candidate examples of horizontal transfer from bacteria to vertebrates. Gene loss combined with sample size effects and evolutionary rate variation provide an alternative, more biologically plausible explanation
- Qi Z, Cui Y, Fang W, Ling L, Chen R (January 2004). "Autosomal similarity revealed by eukaryotic genomic comparison". Journal of Biological Physics. 30 (4): 305–12. doi:10.1007/s10867-004-0996-0. PMC 3456315. PMID 23345874.
- Woese CR (June 2002). "On the evolution of cells". Proceedings of the National Academy of Sciences of the United States of America. 99 (13): 8742–7. Bibcode:2002PNAS...99.8742W. doi:10.1073/pnas.132266999. PMC 124369. PMID 12077305. This article seeks to shift the emphasis in early phylogenic adaptation from vertical to horizontal gene transfer. He uses the term "Darwinian Threshold" for the time of major transition of evolutionary mechanisms from mostly horizontal to mostly vertical transfer, and the "origin of speciation".
- Snel B, Bork P, Huynen MA (January 1999). "Genome phylogeny based on gene content". Nature Genetics. 21 (1): 108–10. doi:10.1038/5052. PMID 9916801. S2CID 10296406. This article proposes using the presence or absence of a set of genes to infer phylogenies, in order to avoid confounding factors such as horizontal gene transfer.
- "Webfocus in Nature with free review articles". Archived from the original on 2005-11-02.
- Patil PB, Sonti RV (October 2004). "Variation suggestive of horizontal gene transfer at a lipopolysaccharide (lps) biosynthetic locus in Xanthomonas oryzae pv. oryzae, the bacterial leaf blight pathogen of rice". BMC Microbiology. 4 (1): 40. doi:10.1186/1471-2180-4-40. PMC 524487. PMID 15473911.
- Jin G, Nakhleh L, Snir S, Tuller T (November 2006). "Maximum likelihood of phylogenetic networks". Bioinformatics. 22 (21): 2604–11. doi:10.1093/bioinformatics/btl452. PMID 16928736.
- Jain R, Rivera MC, Lake JA (March 1999). "Horizontal gene transfer among genomes: the complexity hypothesis". Proceedings of the National Academy of Sciences of the United States of America. 96 (7): 3801–6. Bibcode:1999PNAS...96.3801J. doi:10.1073/pnas.96.7.3801. PMC 22375. PMID 10097118.
- Ochman H, Lawrence JG, Groisman EA (May 2000). "Lateral gene transfer and the nature of bacterial innovation". Nature. 405 (6784): 299–304. Bibcode:2000Natur.405..299O. doi:10.1038/35012500. PMID 10830951. S2CID 85739173.
- Preston R (July 12, 1999). "The Demon in the Freezer". The New Yorker. pp. 44–61. Archived from the original on March 18, 2023. Retrieved February 20, 2020.
Smallpox knows how to make a mouse protein. How did smallpox learn that? 'The poxviruses are promiscuous at capturing genes from their hosts,' Esposito said. 'It tells you that smallpox was once inside a mouse or some other small rodent.'
- Szpirer C, Top E, Couturier M, Mergeay M (December 1999). "Retrotransfer or gene capture: a feature of conjugative plasmids, with ecological and evolutionary significance". Microbiology. 145 (Pt 12): 3321–3329. doi:10.1099/00221287-145-12-3321. PMID 10627031. Archived from the original on 2007-02-23. Retrieved 2006-05-12.
- "Can transgenes from genetically modified plants be absorbed by micro-organisms and spread in this way?". GMO Safety: Results of research into horizontal gene transfer. Archived from the original on 2011-07-21.
- Whitaker JW, McConkey GA, Westhead DR (2009). "The transferome of metabolic genes explored: analysis of the horizontal transfer of enzyme encoding genes in unicellular eukaryotes". Genome Biology. 10 (4): R36. doi:10.1186/gb-2009-10-4-r36. PMC 2688927. PMID 19368726.
External links
- Citizendium:Horizontal gene transfer
- Citizendium:Horizontal gene transfer in prokaryotes
- Citizendium:Horizontal gene transfer in plants
- Citizendium:Horizontal gene transfer (History)
| Genetics: homologous recombination / mobile genetic elements | |
|---|---|
| Primarily prokaryotic | |
| Occurs in eukaryotes | |
| Viral | |