| dopamine beta-monooxygenase | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Identifiers | |||||||||
| EC no. | 1.14.17.1 | ||||||||
| CAS no. | 9013-38-1 | ||||||||
| Databases | |||||||||
| IntEnz | IntEnz view | ||||||||
| BRENDA | BRENDA entry | ||||||||
| ExPASy | NiceZyme view | ||||||||
| KEGG | KEGG entry | ||||||||
| MetaCyc | metabolic pathway | ||||||||
| PRIAM | profile | ||||||||
| PDB structures | RCSB PDB PDBe PDBsum | ||||||||
| Gene Ontology | AmiGO / QuickGO | ||||||||
| |||||||||
Dopamine beta-hydroxylase (DBH), also known as dopamine beta-monooxygenase, is an enzyme (EC 1.14.17.1) that in humans is encoded by the DBH gene. Dopamine beta-hydroxylase catalyzes the conversion of dopamine to norepinephrine.

The three substrates of the enzyme are dopamine, vitamin C (ascorbate), and O2. The products are norepinephrine, dehydroascorbate, and H2O.
DBH is a 290 kDa copper-containing oxygenase consisting of four identical subunits, and its activity requires ascorbate as a cofactor.
It is the only enzyme involved in the synthesis of small-molecule neurotransmitters that is membrane-bound, making norepinephrine the only known transmitter synthesized inside vesicles. It is expressed in noradrenergic neurons of the central nervous system (i.e. locus coeruleus) and peripheral nervous systems (i.e. sympathetic ganglia), as well as in chromaffin cells of the adrenal medulla.
Mechanism of catalysis
Based on the observations of what happens when there is no substrate, or oxygen, the following steps seem to constitute the hydroxylation reaction.

Although details of DBH mechanism are yet to be confirmed, DBH is homologous to another enzyme, peptidylglycine α-hydroxylating monooxygenase (PHM). Because DBH and PHM share similar structures, it is possible to model DBH mechanism based on what is known about PHM mechanism.
Substrate specificity
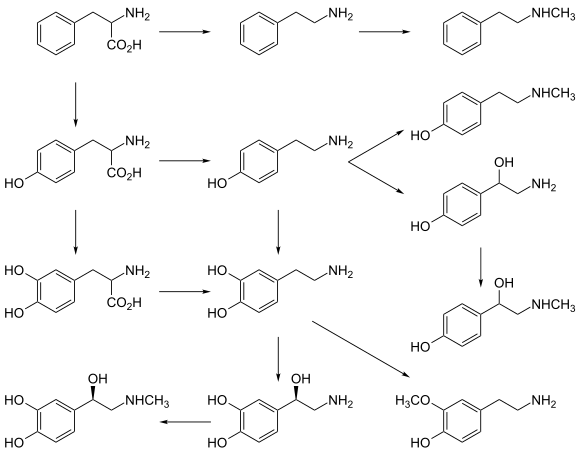
Biosynthetic pathways for catecholamines and trace amines in the human brain
 L-Phenylalanine
L-Tyrosine
L-DOPA
Epinephrine
Phenethylamine
p-Tyramine
Dopamine
Norepinephrine
N-Methylphenethylamine
N-Methyltyramine
p-Octopamine
Synephrine
3-Methoxytyramine
AADC
AADC
AADC
primary
L-Phenylalanine
L-Tyrosine
L-DOPA
Epinephrine
Phenethylamine
p-Tyramine
Dopamine
Norepinephrine
N-Methylphenethylamine
N-Methyltyramine
p-Octopamine
Synephrine
3-Methoxytyramine
AADC
AADC
AADC
primarypathway PNMT PNMT PNMT PNMT AAAH AAAH brain CYP2D6 minor pathway COMT DBH DBH |
Dopamine beta-hydroxylase catalyzes the hydroxylation of not only dopamine but also other phenylethylamine derivatives when available. The minimum requirement seems to be the phenylethylamine skeleton: a benzene ring with a two-carbon side chain that terminates in an amino group.
Assays for DBH activity in human serum and cerebrospinal fluid
DBH activity in human serum could be estimated by a spectrophotometric method or with the aid of Ultra high performance liquid chromatography with Photo Diode Array detector (UHPLC-PDA). A sensitive assay for the detection of DBH activity in cerebrospinal fluid using High-performance liquid chromatography with Electrochemical detector(HPLC-ECD) was also described earlier.
Expression quantitative trait loci (eQTLs) at DBH loci
Genetic variants such as single-nucleotide polymorphisms(SNPs) at DBH loci were found to be associated with DBH activity and are well known expression quantitative trait loci. Allele variants at two regulatory SNPs namely rs1611115 and rs1989787 were shown to affect transcription of this gene. Mutations identified in dopamine beta hydroxylase deficiency and non-synonymous SNPs such as rs6271 in this gene were found to cause defective secretion of the protein from the endoplasmic reticulum.
Clinical significance
DBH primarily contributes to catecholamine and trace amine biosynthesis. It also participates in the metabolism of xenobiotics related to these substances; for example, the human DBH enzyme catalyzes the beta-hydroxylation of amphetamine and para-hydroxyamphetamine, producing norephedrine and para-hydroxynorephedrine respectively.
DBH has been implicated as correlating factor in conditions associated with decision making and addictive drugs, e.g., alcoholism and smoking, attention deficit hyperactivity disorder, schizophrenia, and Alzheimer's disease. Inadequate DBH is called dopamine beta hydroxylase deficiency.
The proximal promoter SNPs rs1989787 and rs1611115 were found to be associated with cognition in schizophrenia subjects. Further these SNPs (rs1989787;rs1611115) and a distal promoter variant 19bp Ins/Del(rs141116007) were associated with scores of Abnormal Involuntary Movement Scale in tardive dyskinesia positive schizophrenia subjects. Of the three variants, the proximal promoter SNP(rs1611115) was associated with Positive and Negative Syndrome Scale(PANSS) scores in tardive dyskinesia positive schizophrenia subjects. The main effect of a putative splice variant in Dopamine beta-hydroxylase namely rs1108580 was found to be associated with Working memory Processing speed in a north Indian Schizophrenia case control study where the G/G genotype of that single-nucleotide polymorphism(SNP) was found to have lower cognitive scores than those with A/A and A/G genotypes. Furthermore the same SNP was associated with Emotion accuracy in healthy controls.
Structure

It was difficult to obtain a stable crystal of dopamine beta-hydroxylase. Hence an homology model based on the primary sequence and comparison to PHM is available.
However, a crystal structure was also put forward in 2016.
Regulation and inhibition
This protein may use the morpheein model of allosteric regulation.
Inhibitors
| HYD | HP | QCA | IQCA | BI | IAA | |
|---|---|---|---|---|---|---|
| Competitive | Ascorbate | Ascorbate | Ascorbate | Ascorbate | Ascorbate | Ascorbate |
| Uncompetitive | Tyramine | Tyramine | ||||
| Mixed | Tyramine | Tyramine | Tyramine | Tyramine | ||
Ascorbate is cofactor; tyramine is substitute for dopamine, DBH's namesake substrate
| ||||||
DBH is inhibited by disulfiram, tropolone, and, most selectively, by nepicastat. It is also inhibited by etamicastat and zamicastat.
DBH is reversibly inhibited by l-2H-Phthalazine hydrazone (hydralazine; HYD), 2-1H-pyridinone hydrazone (2-hydrazinopyridine; HP), 2-quinoline-carboxylic acid (QCA), l-isoquinolinecarboxylic acid (IQCA), 2,2'-bi-lH-imidazole (2,2'-biimidazole; BI), and IH-imidazole-4-acetic acid (imidazole-4-acetic acid; IAA). HYD, QCA, and IAA are allosteric competitive.
Nomenclature
The systematic name of this enzyme class is 3,4-dihydroxyphenethylamine, ascorbate:oxygen oxidoreductase (beta-hydroxylating).
Other names in common use include:
- dopamine beta-monooxygenase
- dopamine beta-hydroxylase
- membrane-associated dopamine beta-monooxygenase (MDBH)
- soluble dopamine beta-monooxygenase (SDBH)
- dopamine-B-hydroxylase
- 3,4-dihydroxyphenethylamine beta-oxidase
- 4-(2-aminoethyl) pyrocatechol beta-oxidase
- dopa beta-hydroxylase
- dopamine beta-oxidase
- dopamine hydroxylase
- phenylamine beta-hydroxylase
- (3,4-dihydroxyphenethylamine) beta-mono-oxygenase
References
- ^ GRCh38: Ensembl release 89: ENSG00000123454 – Ensembl, May 2017
- ^ GRCm38: Ensembl release 89: ENSMUSG00000000889 – Ensembl, May 2017
- "Human PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- "Mouse PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- Rush RA, Geffen LB (1980). "Dopamine beta-hydroxylase in health and disease". Critical Reviews in Clinical Laboratory Sciences. 12 (3): 241–77. doi:10.3109/10408368009108731. PMID 6998654.
- ^ Kaufman S, Bridgers WF, Baron J (1968). "The Mechanism of Action of Dopamine β-Hydroxylase". Oxidation of Organic Compounds. Advances in Chemistry. Vol. 77. pp. 172–176. doi:10.1021/ba-1968-0077.ch073. ISBN 0-8412-0078-5.
- Friedman S, Kaufman S (May 1966). "An electron paramagnetic resonance study of 3,4-dihydroxyphenylethylamine beta-hydroxylase". The Journal of Biological Chemistry. 241 (10): 2256–9. doi:10.1016/S0021-9258(18)96614-7. PMID 4287853.
- Prigge ST, Mains RE, Eipper BA, Amzel LM (August 2000). "New insights into copper monooxygenases and peptide amidation: structure, mechanism and function". Cellular and Molecular Life Sciences. 57 (8–9): 1236–59. doi:10.1007/pl00000763. PMC 11146793. PMID 11028916. S2CID 12738480.
- Broadley KJ (March 2010). "The vascular effects of trace amines and amphetamines". Pharmacology & Therapeutics. 125 (3): 363–375. doi:10.1016/j.pharmthera.2009.11.005. PMID 19948186.
- Lindemann L, Hoener MC (May 2005). "A renaissance in trace amines inspired by a novel GPCR family". Trends in Pharmacological Sciences. 26 (5): 274–281. doi:10.1016/j.tips.2005.03.007. PMID 15860375.
- Wang X, Li J, Dong G, Yue J (February 2014). "The endogenous substrates of brain CYP2D". European Journal of Pharmacology. 724: 211–218. doi:10.1016/j.ejphar.2013.12.025. PMID 24374199.
- Nagatsu T, Udenfriend S (1972). "Photometric Assay of Dopamine-β-Hydroxylase Activity in Human Blood". Clinical Chemistry. 18 (9): 980–983. doi:10.1093/clinchem/18.9.980. PMID 5052101.
- Punchaichira TJ, Deshpande SN, Thelma BK (2018). "Determination of Dopamine-β-hydroxylase Activity in Human Serum Using UHPLC-PDA Detection". Neurochemical Research. 43 (12): 2324–2332. doi:10.1007/s11064-018-2653-1. PMID 30357655. S2CID 53024826.
- Matsui H, Kato T, Yamamoto C, Fujita K, Nagatsu T (1981). "Highly sensitive assay for dopamine-beta-hydroxylase activity in human cerebrospinal fluid by high performance liquid chromatography-electrochemical detection: properties of the enzyme". Journal of Neurochemistry. 37 (2): 289–296. doi:10.1111/j.1471-4159.1981.tb00454.x. PMID 7264660. S2CID 42736106.
- Zabetian CP, Anderson GM, Buxbaum SG, Elston RC, Ichinose H, Nagatsu T, Kim KS, Kim CH, Malison RT, Gelernter J, Cubells JF (2001). "A quantitative-trait analysis of human plasma-dopamine beta-hydroxylase activity: evidence for a major functional polymorphism at the DBH locus". American Journal of Human Genetics. 68 (2): 515–22. doi:10.1086/318198. PMC 1235285. PMID 11170900.
- Punchaichira TJ, Prasad S, Deshpande SN, Thelma BK (2016). "Deep sequencing identifies novel regulatory variants in the distal promoter region of the dopamine-beta-hydroxylase gene". Pharmacogenetics and Genomics. 26 (7): 311–23. doi:10.1097/FPC.0000000000000214. PMID 26959714. S2CID 205601803.
- Chen Y, Wen G, Rao F, Zhang K, Wang L, Rodriguez-Flores JL, Sanchez, AP, Mahata M, Taupenot L, Sun P, Mahata SK, Tayo B, Schork NJ, Ziegler MG, Hamilton BA, O'Connor DT (2010). "Human dopamine beta-hydroxylase (DBH) regulatory polymorphism that influences enzymatic activity, autonomic function, and blood pressure". Journal of Hypertension. 28 (1): 76–86. doi:10.1097/HJH.0b013e328332bc87. PMC 2860271. PMID 20009769.
- Chen Y, Zhang K, Wen G, Rao F, Sanchez AP, Wang L, Rodriguez-Flores JL, Mahata M, Mahata SK, Waalen J, Ziegler MG, Hamilton BA, O'Connor DT (2011). "Human dopamine beta-hydroxylase promoter variant alters transcription in chromaffin cells, enzyme secretion, and blood pressure". American Journal of Hypertension. 24 (1): 24–32. doi:10.1038/ajh.2010.186. PMC 4906639. PMID 20814407.
- Kim CH, Leung A, Huh YH, Yang E, Kim DJ, Leblanc P, Ryu H, Kim K, Kim DW, Garland EM, Raj SR, Biaggioni I, Robertson D, Kim KS (2011). "Norepinephrine deficiency is caused by combined abnormal mRNA processing and defective protein trafficking of dopamine beta-hydroxylase". Journal of Biological Chemistry. 286 (11): 9196–204. doi:10.1074/jbc.M110.192351. PMC 3059068. PMID 21209083.
- Punchaichira TJ, Dey SK, Mukhopadhyay A, Kundu S, Thelma BK (2017). "Characterization of SNPs in the dopamine-beta-hydroxylase gene providing new insights into its structure-function relationship". Neurogenetics. 18 (3): 155–168. doi:10.1007/s10048-017-0519-3. PMID 28707163. S2CID 5259134.
- Glennon RA (2013). "Phenylisopropylamine stimulants: amphetamine-related agents". In Lemke TL, Williams DA, Roche VF, Zito W (eds.). Foye's principles of medicinal chemistry (7th ed.). Philadelphia, US: Wolters Kluwer Health/Lippincott Williams & Wilkins. pp. 646–648. ISBN 9781609133450. Archived from the original on 8 March 2024. Retrieved 11 September 2015.
The phase 1 metabolism of amphetamine analogs is catalyzed by two systems: cytochrome P450 and flavin monooxygenase. ... Amphetamine can also undergo aromatic hydroxylation to p-hydroxyamphetamine. ... Subsequent oxidation at the benzylic position by DA β-hydroxylase affords p-hydroxynorephedrine. Alternatively, direct oxidation of amphetamine by DA β-hydroxylase can afford norephedrine.
- Taylor KB (January 1974). "Dopamine-beta-hydroxylase. Stereochemical course of the reaction" (PDF). J. Biol. Chem. 249 (2): 454–458. doi:10.1016/S0021-9258(19)43051-2. PMID 4809526. Archived (PDF) from the original on 7 October 2018. Retrieved 6 November 2014.
Dopamine-β-hydroxylase catalyzed the removal of the pro-R hydrogen atom and the production of 1-norephedrine, (2S,1R)-2-amino-1-hydroxyl-1-phenylpropane, from d-amphetamine.
- Horwitz D, Alexander RW, Lovenberg W, Keiser HR (May 1973). "Human serum dopamine-β-hydroxylase. Relationship to hypertension and sympathetic activity". Circ. Res. 32 (5): 594–599. doi:10.1161/01.RES.32.5.594. PMID 4713201.
Subjects with exceptionally low levels of serum dopamine-β-hydroxylase activity showed normal cardiovascular function and normal β-hydroxylation of an administered synthetic substrate, hydroxyamphetamine.
- Mutschler J, Abbruzzese E, Witt SH, Dirican G, Nieratschker V, Frank J, Grosshans M, Rietschel M, Kiefer F (August 2012). "Functional polymorphism of the dopamine β-hydroxylase gene is associated with increased risk of disulfiram-induced adverse effects in alcohol-dependent patients". Journal of Clinical Psychopharmacology. 32 (4): 578–80. doi:10.1097/jcp.0b013e31825ddbe6. PMID 22760354.
- Ella E, Sato N, Nishizawa D, Kageyama S, Yamada H, Kurabe N, Ishino K, Tao H, Tanioka F, Nozawa A, Renyin C, Shinmura K, Ikeda K, Sugimura H (June 2012). "Association between dopamine beta hydroxylase rs5320 polymorphism and smoking behaviour in elderly Japanese". Journal of Human Genetics. 57 (6): 385–90. doi:10.1038/jhg.2012.40. PMID 22513716.
- Bhaduri N, Sinha S, Chattopadhyay A, Gangopadhyay PK, Singh M, Mukhopadhyay KK (February 2005). "Analysis of polymorphisms in the dopamine beta hydroxylase gene: association with attention deficit hyperactivity disorder in Indian children". Indian Pediatrics. 42 (2): 123–9. PMID 15767706.
- Cubells JF, Sun X, Li W, Bonsall RW, McGrath JA, Avramopoulos D, Lasseter VK, Wolyniec PS, Tang YL, Mercer K, Pulver AE, Elston RC (November 2011). "Linkage analysis of plasma dopamine β-hydroxylase activity in families of patients with schizophrenia". Human Genetics. 130 (5): 635–43. doi:10.1007/s00439-011-0989-6. PMC 3193571. PMID 21509519.
- Combarros O, Warden DR, Hammond N, Cortina-Borja M, Belbin O, Lehmann MG, Wilcock GK, Brown K, Kehoe PG, Barber R, Coto E, Alvarez V, Deloukas P, Gwilliam R, Heun R, Kölsch H, Mateo I, Oulhaj A, Arias-Vásquez A, Schuur M, Aulchenko YS, Ikram MA, Breteler MM, van Duijn CM, Morgan K, Smith AD, Lehmann DJ (2010). "The dopamine β-hydroxylase -1021C/T polymorphism is associated with the risk of Alzheimer's disease in the Epistasis Project". BMC Medical Genetics. 11 (161): 162. doi:10.1186/1471-2350-11-162. PMC 2994840. PMID 21070631.
- ^ Punchaichira TJ, Mukhopadhyay A, Kukshal P, Bhatia T, Deshpande SN, Thelma BK (2020). "Association of regulatory variants of dopamine β-hydroxylase with cognition and tardive dyskinesia in schizophrenia subjects". Journal of Psychopharmacology. 34 (3): 358–369. doi:10.1177/0269881119895539. PMC 7150076. PMID 31913053.
- Punchaichira TJ, Kukshal P, Bhatia T, Deshpande SN (2023). "Effect of rs1108580 of DBH and rs1006737 of CACNA1C on Cognition and Tardive Dyskinesia in a North Indian Schizophrenia Cohort". Molecular Neurobiology. 60 (12): 6826–6839. doi:10.1007/s12035-023-03496-4. PMID 37493923. S2CID 260162784.
- ^ Kapoor A, Shandilya M, Kundu S (2011). "Structural insight of dopamine β-hydroxylase, a drug target for complex traits, and functional significance of exonic single nucleotide polymorphisms". PLOS ONE. 6 (10): e26509. Bibcode:2011PLoSO...626509K. doi:10.1371/journal.pone.0026509. PMC 3197665. PMID 22028891.
- Vendelboe TV, Harris P, Zhao Y, Walter TS, Harlos K, Omari KE, Christensen HM (2016). "The crystal structure of human dopamine β-hydroxylase at 2.9 Å resolution". Science Advances. 2 (4): e1500980. Bibcode:2016SciA....2E0980V. doi:10.1126/sciadv.1500980. PMC 4846438. PMID 27152332.
- Selwood T, Jaffe EK (March 2012). "Dynamic dissociating homo-oligomers and the control of protein function". Archives of Biochemistry and Biophysics. 519 (2): 131–43. doi:10.1016/j.abb.2011.11.020. PMC 3298769. PMID 22182754.
- Goldstein M, Anagnoste B, Lauber E, Mckeregham MR (July 1964). "Inhibition of dopamine- β -hydroxylase by disulfiram". Life Sciences. 3 (7): 763–7. doi:10.1016/0024-3205(64)90031-1. PMID 14203977.
- Goldstein M, Lauber E, Mckereghan MR (July 1964). "The inhibitionof dopamine-β-hydroxylase by tropolone and other chelating agents". Biochemical Pharmacology. 13 (7): 1103–6. doi:10.1016/0006-2952(64)90109-1. PMID 14201135.
- Stanley WC, Li B, Bonhaus DW, Johnson LG, Lee K, Porter S, Walker K, Martinez G, Eglen RM, Whiting RL, Hegde SS (August 1997). "Catecholamine modulatory effects of nepicastat (RS-25560-197), a novel, potent and selective inhibitor of dopamine-beta-hydroxylase". British Journal of Pharmacology. 121 (8): 1803–9. doi:10.1038/sj.bjp.0701315. PMC 1564872. PMID 9283721.
- Dey SK, Saini M, Prabhakar P, Kundu S (September 2020). "Dopamine β hydroxylase as a potential drug target to combat hypertension". Expert Opin Investig Drugs. 29 (9): 1043–1057. doi:10.1080/13543784.2020.1795830. PMID 32658551.
- Townes S, Titone C, Rosenberg RC (February 1990). "Inhibition of dopamine beta-hydroxylase by bidentate chelating agents". Biochimica et Biophysica Acta (BBA) - Protein Structure and Molecular Enzymology. 1037 (2): 240–7. doi:10.1016/0167-4838(90)90174-E. PMID 2306475.
Further reading
- Friedman S, Kaufman S (December 1965). "3,4-dihydroxyphenylethylamine beta-hydroxylase. Physical properties, copper content, and role of copper in the catalytic activity". The Journal of Biological Chemistry. 240 (12): 4763–73. doi:10.1016/S0021-9258(18)97021-3. PMID 5846992.
- Levin EY, Levenberg B, Kaufman S (1960). "The enzymatic conversion of 3,4-dihydroxyphenylethylamine to norepinephrine". J. Biol. Chem. 235 (7): 2080–2086. doi:10.1016/S0021-9258(18)69366-4. PMID 14416204.
External links
- GeneReviews/NIH/NCBI/UW entry on Dopamine Beta-Hydroxylase Deficiency
- Dopamine+beta-Hydroxylase at the U.S. National Library of Medicine Medical Subject Headings (MeSH)
| Enzymes involved in neurotransmission | |||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| monoamine |
| ||||||||||||||||||||
| arginine→NO | |||||||||||||||||||||
| choline→Acetylcholine |
| ||||||||||||||||||||
| Oxidoreductases: dioxygenases, including steroid hydroxylases (EC 1.14) | |
|---|---|
| 1.14.11: 2-oxoglutarate | |
| 1.14.13: NADH or NADPH | |
| 1.14.14: reduced flavin or flavoprotein | |
| 1.14.15: reduced iron–sulfur protein | |
| 1.14.16: reduced pteridine (BH4 dependent) | |
| 1.14.17: reduced ascorbate | |
| 1.14.18-19: other | |
| 1.14.99 - miscellaneous | |
| Enzymes | |
|---|---|
| Activity | |
| Regulation | |
| Classification | |
| Kinetics | |
| Types |
|
| Monoamine metabolism modulators | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Non-specific |
| ||||||||||
| Phenethylamines (dopamine, epinephrine, norepinephrine) |
| ||||||||||
| Tryptamines (serotonin, melatonin) |
| ||||||||||
| Histamine |
| ||||||||||
| See also: Receptor/signaling modulators • Adrenergics • Dopaminergics • Melatonergics • Serotonergics • Monoamine reuptake inhibitors • Monoamine releasing agents • Monoamine neurotoxins | |||||||||||
Categories: