| Double-stranded RNA virus | |
|---|---|

| |
| Electron micrograph of rotaviruses. The bar = 100 nm | |
| Virus classification | |
| Group: | Group III (dsRNA) |
| Kingdom: Phylum: Class | |
Double-stranded RNA viruses (dsRNA viruses) are a polyphyletic group of viruses that have double-stranded genomes made of ribonucleic acid. The double-stranded genome is used as a template by the viral RNA-dependent RNA polymerase (RdRp) to transcribe a positive-strand RNA functioning as messenger RNA (mRNA) for the host cell's ribosomes, which translate it into viral proteins. The positive-strand RNA can also be replicated by the RdRp to create a new double-stranded viral genome.
A distinguishing feature of the dsRNA viruses is their ability to carry out transcription of the dsRNA segments within the capsid, and the required enzymes are part of the virion structure.
Double-stranded RNA viruses are classified into two phyla, Duplornaviricota and Pisuviricota (specifically class Duplopiviricetes), in the kingdom Orthornavirae and realm Riboviria. The two phyla do not share a common dsRNA virus ancestor, but evolved their double strands two separate times from positive-strand RNA viruses. In the Baltimore classification system, dsRNA viruses belong to Group III.
Virus group members vary widely in host range (animals, plants, fungi, and bacteria), genome segment number (one to twelve), and virion organization (T-number, capsid layers, or turrets). Double-stranded RNA viruses include the rotaviruses, known globally as a common cause of gastroenteritis in young children, and bluetongue virus, an economically significant pathogen of cattle and sheep. The family Reoviridae is the largest and most diverse dsRNA virus family in terms of host range.
Classification
Two clades of dsRNA viruses exist: the phylum Duplornaviricota and the class Duplopiviricetes, which is in the phylum Pisuviricota. Both are included in the kingdom Orthornavirae in the realm Riboviria. Based on phylogenetic analysis of RdRp, the two clades do not share a common dsRNA ancestor but are instead separately descended from different positive-sense, single-stranded RNA viruses. In the Baltimore classification system, which groups viruses together based on their manner of mRNA synthesis, dsRNA viruses are group III.
Duplornaviricota
Duplornaviricota contains most dsRNA viruses, including reoviruses, which infect a diverse range of eukaryotes, and cystoviruses, which are the only dsRNA viruses known to infect prokaryotes. Apart from RdRp, viruses in Duplornaviricota also share icosahedral capsids that contain 60 homo- or heterodimers of the capsid protein organized on a pseudo T=2 lattice. The phylum is divided into three classes: Chrymotiviricetes, which primarily contains fungal and protozoan viruses, Resentoviricetes, which contains reoviruses, and Vidaverviricetes, which contains cystoviruses.
Duplopiviricetes
The class Duplopiviricetes is the second clade of dsRNA viruses and is in the phylum Pisuviricota, which also contains positive-sense single-stranded RNA viruses. Duplopiviricetes mostly contains plant and fungal viruses and includes the following four families: Amalgaviridae, Hypoviridae, Partitiviridae, and Picobirnaviridae.
Notes on selected species
Reoviridae
Reoviridae are currently classified into nine genera. The genomes of these viruses consist of 10 to 12 segments of dsRNA, each generally encoding one protein. The mature virions are non-enveloped. Their capsids, formed by multiple proteins, have icosahedral symmetry and are arranged generally in concentric layers.
Orthoreoviruses
The orthoreoviruses (reoviruses) are the prototypic members of the virus Reoviridae family and representative of the turreted members, which comprise about half the genera. Like other members of the family, the reoviruses are non-enveloped and characterized by concentric capsid shells that encapsidate a segmented dsRNA genome. In particular, reovirus has eight structural proteins and ten segments of dsRNA. A series of uncoating steps and conformational changes accompany cell entry and replication. High-resolution structures are known for almost all of the proteins of mammalian reovirus (MRV), which is the best-studied genotype. Electron cryo-microscopy (cryoEM) and X-ray crystallography have provided a wealth of structural information about two specific MRV strains, type 1 Lang (T1L) and type 3 Dearing (T3D).
Cypovirus
The cytoplasmic polyhedrosis viruses (CPVs) form the genus Cypovirus of the family Reoviridae. CPVs are classified into 14 species based on the electrophoretic migration profiles of their genome segments. Cypovirus has only a single capsid shell, which is similar to the orthoreovirus inner core. CPV exhibits striking capsid stability and is fully capable of endogenous RNA transcription and processing. The overall folds of CPV proteins are similar to those of other reoviruses. However, CPV proteins have insertional domains and unique structures that contribute to their extensive intermolecular interactions. The CPV turret protein contains two methylase domains with a highly conserved helix-pair/β-sheet/helix-pair sandwich fold but lacks the β-barrel flap present in orthoreovirus λ2. The stacking of turret protein functional domains and the presence of constrictions and A spikes along the mRNA release pathway indicate a mechanism that uses pores and channels to regulate the highly coordinated steps of RNA transcription, processing, and release.
Rotavirus
Rotavirus is the most common cause of acute gastroenteritis in infants and young children worldwide. This virus contains a dsRNA genome and is a member of the Reoviridae family. The genome of rotavirus consists of eleven segments of dsRNA. Each genome segment codes for one protein with the exception of segment 11, which codes for two proteins. Among the twelve proteins, six are structural and six are non-structural proteins. It is a double-stranded RNA non-enveloped virus.
Bluetongue virus
The members of genus Orbivirus within the Reoviridae family are arthropod borne viruses and are responsible for high morbidity and mortality in ruminants. Bluetongue virus (BTV) which causes disease in livestock (sheep, goat, cattle) has been in the forefront of molecular studies for the last three decades and now represents the best understood orbivirus at the molecular and structural levels. BTV, like other members of the family, is a complex non-enveloped virus with seven structural proteins and a RNA genome consisting of 10 variously sized dsRNA segments.
Phytoreoviruses
Phytoreoviruses are non-turreted reoviruses that are major agricultural pathogens, particularly in Asia. One member of this family, Rice Dwarf Virus (RDV), has been extensively studied by electron cryomicroscopy and x-ray crystallography. From these analyses, atomic models of the capsid proteins and a plausible model for capsid assembly have been derived. While the structural proteins of RDV share no sequence similarity to other proteins, their folds and the overall capsid structure are similar to those of other Reoviridae.
Saccharomyces cerevisiae virus L-A
The L-A dsRNA virus of the yeast Saccharomyces cerevisiae has a single 4.6 kb genomic segment that encodes its major coat protein, Gag (76 kDa) and a Gag-Pol fusion protein (180 kDa) formed by a -1 ribosomal frameshift. L-A can support the replication and encapsidation in separate viral particles of any of several satellite dsRNAs, called M dsRNAs, each of which encodes a secreted protein toxin (the killer toxin) and immunity to that toxin. L-A and M are transmitted from cell to cell by the cytoplasmic mixing that occurs in the process of mating. Neither is naturally released from the cell or enters cells by other mechanisms, but the high frequency of yeast mating in nature results in the wide distribution of these viruses in natural isolates. Moreover, the structural and functional similarities with dsRNA viruses of mammals has made it useful to consider these entities as viruses.
Infectious bursal disease virus
Infectious bursal disease virus (IBDV) is the best-characterized member of the family Birnaviridae. These viruses have bipartite dsRNA genomes enclosed in single layered icosahedral capsids with T = 13l geometry. IBDV shares functional strategies and structural features with many other icosahedral dsRNA viruses, except that it lacks the T = 1 (or pseudo T = 2) core common to the Reoviridae, Cystoviridae, and Totiviridae. The IBDV capsid protein exhibits structural domains that show homology to those of the capsid proteins of some positive-sense single-stranded RNA viruses, such as the nodaviruses and tetraviruses, as well as the T = 13 capsid shell protein of the Reoviridae. The T = 13 shell of the IBDV capsid is formed by trimers of VP2, a protein generated by removal of the C-terminal domain from its precursor, pVP2. The trimming of pVP2 is performed on immature particles as part of the maturation process. The other major structural protein, VP3, is a multifunctional component lying under the T = 13 shell that influences the inherent structural polymorphism of pVP2. The virus-encoded RNA-dependent RNA polymerase, VP1, is incorporated into the capsid through its association with VP3. VP3 also interacts extensively with the viral dsRNA genome.
Bacteriophage Φ6
Bacteriophage Φ6, is a member of the Cystoviridae family. It infects Pseudomonas bacteria (typically plant-pathogenic P. syringae). It has a three-part, segmented, double-stranded RNA genome, totalling ~13.5 kb in length. Φ6 and its relatives have a lipid membrane around their nucleocapsid, a rare trait among bacteriophages. It is a lytic phage, though under certain circumstances has been observed to display a delay in lysis which may be described as a "carrier state".
Anti-virals
Since cells do not produce double-stranded RNA during normal nucleic acid metabolism, natural selection has favored the evolution of enzymes that destroy dsRNA on contact. The best known class of this type of enzymes is Dicer. It is hoped that broad-spectrum anti-virals could be synthesized that take advantage of this vulnerability of double-stranded RNA viruses.
See also
References
- "Double-stranded RNA virus replication". ViralZone. Swiss Institute of Bioinformatics. Retrieved 6 August 2020.
- ^ Patton 2008
- ^ Koonin EV, Dolja VV, Krupovic M, Varsani A, Wolf YI, Yutin N, Zerbini M, Kuhn JH (18 October 2019). "Create a megataxonomic framework, filling all principal taxonomic ranks, for realm Riboviria" (docx). International Committee on Taxonomy of Viruses (ICTV). Retrieved 15 August 2020.
- ^ Wolf YI, Kazlauskas D, Iranzo J, Lucia-Sanz A, Kuhn JH, Krupovic M, Dolja VV, Kooning EV (27 November 2018). "Origins and Evolution of the Global RNA Virome". mBio. 9 (6): e02329-18. doi:10.1128/mBio.02329-18. PMC 6282212. PMID 30482837.
- Dryden KA, Coombs KM, Yeager M (2008). "Ch. 1: The Structure of Orthoreoviruses". Patton 2008. Caister Academic Press. pp. 3–. ISBN 9781904455219.
- Z. Hong Zhou (2008). "Ch. 2: Cypovirus". Patton 2008. Caister Academic Press. pp. 27–. ISBN 9781904455219.
- Xiaofang Jiang, Crawford SE, Estes MK, Prasad BV (2008). "Ch. 3: Rotavirus Structure". Patton 2008. Caister Academic Press. pp. 45–. ISBN 9781904455219.
- Roy P (2008). "Structure and Function of Bluetongue Virus and its Proteins". Segmented Double-stranded RNA Viruses: Structure and Molecular Biology. Caister Academic Press. ISBN 978-1-904455-21-9.
- Mettenleiter T, Sobrino F, eds. (2008). Animal Viruses: Molecular Biology. Caister Academic. ISBN 978-1-904455-22-6.
- Baker ML, Z. Hong Zhou, Wah Chiu (2008). "Ch. 5: Structures of Phytoreoviruses". Patton 2008. Caister Academic Press. pp. 89–. ISBN 9781904455219.
- Wickner, et al. (2008). "The Yeast dsRNA Virus L-A Resembles Mammalian dsRNA Virus Cores". Segmented Double-stranded RNA Viruses: Structure and Molecular Biology. Caister Academic Press. ISBN 978-1-904455-21-9.
- Castón JR, Rodríguez JF, Carrascosa JL (2008). "Infectious Bursal Disease Virus (IBDV): A Segmented Double-Stranded RNA Virus With a T=13 Capsid That Lacks a T=1 Core". Patton 2008. Caister Academic Press. pp. 133–. ISBN 9781904455219.
- Koivunen MR, Sarin LP, Bamford DH (2008). "Ch. 14: Structure-Function Insights Into the RNA-Dependent RNA Polymerase of the dsRNA Bacteriophage Φ6". Patton 2008. Caister Academic Press. pp. 239–. ISBN 9781904455219.
- Rider TH, Zook CE, Boettcher TL, Wick ST, Pancoast JS, Zusman BD (2011). Sambhara S (ed.). "Broad-Spectrum Antiviral Therapeutics". PLoS ONE. 6 (7): e22572. Bibcode:2011PLoSO...622572R. doi:10.1371/journal.pone.0022572. PMC 3144912. PMID 21818340.
...a new broad-spectrum antiviral approach, dubbed Double-stranded RNA (dsRNA) Activated Caspase Oligomerizer (DRACO)...
Bibliography
- Patton JT, ed. (2008). Segmented Double-stranded RNA Viruses: Structure and Molecular Biology. Caister Academic. ISBN 978-1-904455-21-9.
| Microbiology: Virus | ||
|---|---|---|
| Components | 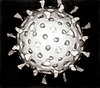 | |
| Viral life cycle | ||
| Genetics | ||
| By host | ||
| Other | ||
| Self-replicating organic structures | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Cellular life | |||||||||||
| Virus | |||||||||||
| Subviral agents |
| ||||||||||
| Nucleic acid self-replication |
| ||||||||||
| Endosymbiosis | |||||||||||
| Abiogenesis | |||||||||||
| See also | |||||||||||