Hypoxia-inducible factor 1-alpha, also known as HIF-1-alpha, is a subunit of a heterodimeric transcription factor hypoxia-inducible factor 1 (HIF-1) that is encoded by the HIF1A gene. The Nobel Prize in Physiology or Medicine 2019 was awarded for the discovery of HIF.
HIF1A is a basic helix-loop-helix PAS domain containing protein, and is considered as the master transcriptional regulator of cellular and developmental response to hypoxia. The dysregulation and overexpression of HIF1A by either hypoxia or genetic alternations have been heavily implicated in cancer biology, as well as a number of other pathophysiologies, specifically in areas of vascularization and angiogenesis, energy metabolism, cell survival, and tumor invasion. The presence of HIF1A in a hypoxic environment is required to push forward normal placental development in early gestation. Two other alternative transcripts encoding different isoforms have been identified.
Structure
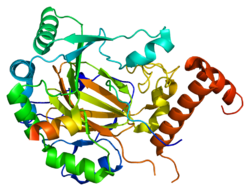
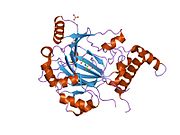
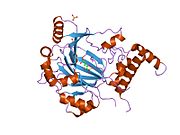
HIF1 is a heterodimeric basic helix-loop-helix structure that is composed of HIF1A, the alpha subunit (this protein), and the aryl hydrocarbon receptor nuclear translocator (Arnt), the beta subunit. HIF1A contains a basic helix-loop-helix domain near the C-terminal, followed by two distinct PAS (PER-ARNT-SIM) domains, and a PAC (PAS-associated C-terminal) domain. The HIF1A polypeptide also contains a nuclear localization signal motif, two transactivating domains CTAD and NTAD, and an intervening inhibitory domain (ID) that can repress the transcriptional activities of CTAD and NTAD. There are a total of three HIF1A isoforms formed by alternative splicing, however isoform1 has been chosen as the canonical structure, and is the most extensively studied isoform in structure and function.
Gene and expression
The human HIF1A gene encodes for the alpha subunit, HIF1A of the transcription factor hypoxia-inducible factor (HIF1). Its protein expression level can be measured by antibodies against HIF-1-alpha through various biological detection methods including western blot or immunostaining. HIF1A expression level is dependent on its GC-rich promoter activation. In most cells, HIF1A gene is constitutively expressed in low levels under normoxic conditions, however, under hypoxia, HIF1A transcription is often significantly upregulated. Typically, oxygen-independent pathway regulates protein expression, and oxygen-dependent pathway regulates degradation. In hypoxia-independent ways, HIF1A expression may be upregulated through a redox-sensitive mechanism.
Function

The transcription factor HIF-1 plays an important role in cellular response to systemic oxygen levels in mammals. HIF1A activity is regulated by a host of post-translational modifications: hydroxylation, acetylation, and phosphorylation. HIF-1 is known to induce transcription of more than 60 genes, including VEGF and erythropoietin that are involved in biological processes such as angiogenesis and erythropoiesis, which assist in promoting and increasing oxygen delivery to hypoxic regions. HIF-1 also induces transcription of genes involved in cell proliferation and survival, as well as glucose and iron metabolism. In accordance with its dynamic biological role, HIF-1 responds to systemic oxygen levels by undergoing conformational changes, and associates with HRE regions of promoters of hypoxia-responsive genes to induce transcription.
HIF1A stability, subcellular localization, as well as transcriptional activity are especially affected by oxygen level. The alpha subunit forms a heterodimer with the beta subunit. Under normoxic conditions, VHL-mediated ubiquitin protease pathway rapidly degrades HIF1A; however, under hypoxia, HIF1A protein degradation is prevented and HIF1A levels accumulate to associate with HIF1B to exert transcriptional roles on target genes Enzymes prolyl hydroxylase (PHD) and HIF prolyl hydroxylase (HPH) are involved in specific post-translational modification of HIF1A proline residues (P402 and P564 within the ODD domain), which allows for VHL association with HIF1A. The enzymatic activity of oxygen sensor dioxygenase PHD is dependent on oxygen level as it requires oxygen as one of its main substrates to transfer to the proline residue of HIF1A. The hydroxylated proline residue of HIF1A is then recognized and buried in the hydrophobic core of von Hippel-Lindau tumor suppressor protein (VHL), which itself is part of a ubiquitin ligase enzyme. Once the hydrolylated HIF1A is buried in the VHL protein, VHL will transport it to a proteasome to digest and destroy HIF1A. This prevents HIF1A from entering into the cell nucleus to carry out the transcription of many different regulatory pathways. Many of these pathways are necessary for proper placental development in early gestation. Under normoxic conditions the HIF1A will be hydroxylated and destroyed, which leads to placental tissue necrosis, disorganization, and overgrowth. The hydroxylation of HIF1A proline residue also regulates its ability to associate with co-activators under hypoxia. Function of HIF1A gene can be effectively examined by siRNA knockdown based on an independent validation.
Repair, regeneration and rejuvenation
In normal circumstances after injury HIF1A is degraded by prolyl hydroxylases (PHDs). In June 2015, scientists found that the continued up-regulation of HIF1A via PHD inhibitors regenerates lost or damaged tissue in mammals that have a repair response; and the continued down-regulation of HIF1A results in healing with a scarring response in mammals with a previous regenerative response to the loss of tissue. The act of regulating HIF1A can either turn off, or turn on the key processes of mammalian regeneration. One such regenerative process in which HIF1A is involved is peripheral nerve regeneration. Following axon injury, HIF1A activates VEGFA to promote regeneration and functional recovery. HIF1A also controls skin healing. Researchers at the Stanford University School of Medicine demonstrated that HIF1A activation was able to prevent and treat chronic wounds in diabetic and aged mice. Not only did the wounds in the mice heal more quickly, but the quality of the new skin was even better than the original. Additionally the regenerative effect of HIF-1A modulation on aged skin cells was described and a rejuvenating effect on aged facial skin was demonstrated in patients. HIF modulation has also been linked to a beneficial effect on hair loss. The biotech company Tomorrowlabs GmbH, founded in Vienna in 2016 by the physician Dominik Duscher and pharmacologist Dominik Thor, makes use of this mechanism. Based on the patent-pending HSF ("HIF strengthening factor") active ingredient, products have been developed that are supposed to promote skin and hair regeneration.
Regulation
HIF1A abundance (and its subsequent activity) is regulated transcriptionally in an NF-κB-dependent manner. In addition, the coordinated activity of the prolyl hydroxylases (PHDs) maintain the appropriate balance of HIF1A protein in the post-translation phase.
PHDs rely on iron among other molecules to hydroxylate HIF1A; as such, iron chelators such as desferrioxamine (DFO) have proven successful in HIF1A stabilization. HBO (Hyperbaric oxygen therapy) and HIF1A imitators such as cobalt chloride have also been successfully utilized.
Factors increasing HIF1A
- Modulator of Degradation:
- Oxygen-Dependent:
- EPF UCP (degrades pHVL)
- VDU2 (de-ubiquitinates HIF1A)
- SUMOylation (via RSUME)
- DeSUMOylation ( via SENP1)
- Oxygen-independent:
- Calcineurin A ( Ca2+-dependent via RACK1)
- Oxygen-Dependent:
- Modulators of translation:
Factors decreasing HIF1A
- Modulator of Degradation:
- Modulators of translation:
- Calcium signaling
- miRNAs
Role in cancer
HIF1A is overexpressed in many human cancers. HIF1A overexpression is heavily implicated in promoting tumor growth and metastasis through its role in initiating angiogenesis and regulating cellular metabolism to overcome hypoxia. Hypoxia promotes apoptosis in both normal and tumor cells. However, hypoxic conditions in tumor microenvironment especially, along with accumulation of genetic alternations often contribute to HIF1A overexpression.
Significant HIF1A expression has been noted in most solid tumors studied, which include cancers of the gastric, colon, breast, pancreas, kidneys, prostate, ovary, brain, and bladder. Clinically, elevated HIF1A levels in a number of cancers, including cervical cancer, non-small-cell lung carcinoma, breast cancer (LV-positive and negative), oligodendroglioma, oropharyngeal cancer, ovarian cancer, endometrial cancer, esophageal cancer, head and neck cancer, and stomach cancer, have been associated with aggressive tumor progression, and thus has been implicated as a predictive and prognostic marker for resistance to radiation treatment, chemotherapy, and increased mortality. HIF1A expression may also regulate breast tumor progression. Elevated HIF1A levels may be detected in early cancer development, and have been found in early ductal carcinoma in situ, a pre-invasive stage in breast cancer development, and is also associated with increased microvasculature density in tumor lesions. Moreover, despite histologically-determined low-grade, lymph-node negative breast tumor in a subset of patients examined, detection of significant HIF1A expression was able to independently predict poor response to therapy. Similar findings have been reported in brain cancer and ovarian cancer studies as well, and suggest at regulatory role of HIF1A in initiating angiogenesis through interactions with pro-angiogenic factors such as VEGF. Studies of glioblastoma multiforme show striking similarity between HIF1A expression pattern and that of VEGF gene transcription level. In addition, high-grade glioblastoma multiform tumors with high VEGF expression pattern, similar to breast cancer with HIF1A overexpression, display significant signs of tumor neovascularization. This further suggests the regulatory role of HIF1A in promoting tumor progression, likely through hypoxia-induced VEGF expression pathways.
HIF1A overexpression in tumors may also occur in a hypoxia-independent pathway. In hemangioblastoma, HIF1A expression is found in most cells sampled from the well-vascularized tumor. Although in both renal carcinoma and hemangioblastoma, the von Hippel-Lindau gene is inactivated, HIF1A is still expressed at high levels. In addition to VEGF overexpression in response elevated HIF1A levels, the PI3K/AKT pathway is also involved in tumor growth. In prostate cancers, the commonly occurring PTEN mutation is associated with tumor progression toward aggressive stage, increased vascular density and angiogenesis.
During hypoxia, tumor suppressor p53 overexpression may be associated with HIF1A-dependent pathway to initiate apoptosis. Moreover, p53-independent pathway may also induce apoptosis through the Bcl-2 pathway. However, overexpression of HIF1A is cancer- and individual-specific, and depends on the accompanying genetic alternations and levels of pro- and anti-apoptotic factors present. One study on epithelial ovarian cancer shows HIF1A and nonfunctional tumor suppressor p53 is correlated with low levels of tumor cell apoptosis and poor prognosis. Further, early-stage esophageal cancer patients with demonstrated overexpression of HIF1 and absence of BCL2 expression also failed photodynamic therapy.
While research efforts to develop therapeutic drugs to target hypoxia-associated tumor cells have been ongoing for many years, there has not yet been any breakthrough that has shown selectivity and effectiveness at targeting HIF1A pathways to decrease tumor progression and angiogenesis. Successful therapeutic approaches in the future may also be highly case-specific to particular cancers and individuals, and seem unlikely to be widely applicable due to the genetically heterogenous nature of the many cancer types and subtypes.
Interactions
HIF1A has been shown to interact with:
See also
References
- ^ GRCh38: Ensembl release 89: ENSG00000100644 – Ensembl, May 2017
- ^ GRCm38: Ensembl release 89: ENSMUSG00000021109 – Ensembl, May 2017
- "Human PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- "Mouse PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- Semenza GL, Rue EA, Iyer NV, Pang MG, Kearns WG (June 1996). "Assignment of the hypoxia-inducible factor 1alpha gene to a region of conserved synteny on mouse chromosome 12 and human chromosome 14q". Genomics. 34 (3): 437–9. doi:10.1006/geno.1996.0311. PMID 8786149.
- ^ Hogenesch JB, Chan WK, Jackiw VH, Brown RC, Gu YZ, Pray-Grant M, et al. (March 1997). "Characterization of a subset of the basic-helix-loop-helix-PAS superfamily that interacts with components of the dioxin signaling pathway". The Journal of Biological Chemistry. 272 (13): 8581–93. doi:10.1074/jbc.272.13.8581. PMID 9079689. S2CID 14908247.
- ^ "Entrez Gene: HIF1A hypoxia-inducible factor 1, alpha subunit (basic helix-loop-helix transcription factor)".
- ^ Wang GL, Jiang BH, Rue EA, Semenza GL (June 1995). "Hypoxia-inducible factor 1 is a basic-helix-loop-helix-PAS heterodimer regulated by cellular O2 tension". Proceedings of the National Academy of Sciences of the United States of America. 92 (12): 5510–4. Bibcode:1995PNAS...92.5510W. doi:10.1073/pnas.92.12.5510. PMC 41725. PMID 7539918.
- Iyer NV, Kotch LE, Agani F, Leung SW, Laughner E, Wenger RH, et al. (January 1998). "Cellular and developmental control of O2 homeostasis by hypoxia-inducible factor 1 alpha". Genes & Development. 12 (2): 149–62. doi:10.1101/gad.12.2.149. PMC 316445. PMID 9436976.
- ^ Semenza GL (October 2003). "Targeting HIF-1 for cancer therapy". Nature Reviews. Cancer. 3 (10): 721–32. doi:10.1038/nrc1187. PMID 13130303. S2CID 2448376.
- Soares MJ, Iqbal K, Kozai K (October 2017). "Hypoxia and Placental Development". Birth Defects Research. 109 (17): 1309–1329. doi:10.1002/bdr2.1135. PMC 5743230. PMID 29105383.
- Wang FS, Wang CJ, Chen YJ, Chang PR, Huang YT, Sun YC, et al. (March 2004). "Ras induction of superoxide activates ERK-dependent angiogenic transcription factor HIF-1alpha and VEGF-A expression in shock wave-stimulated osteoblasts". The Journal of Biological Chemistry. 279 (11): 10331–7. doi:10.1074/jbc.M308013200. PMID 14681237. S2CID 23881074.
- Jiang BH, Zheng JZ, Leung SW, Roe R, Semenza GL (August 1997). "Transactivation and inhibitory domains of hypoxia-inducible factor 1alpha. Modulation of transcriptional activity by oxygen tension". The Journal of Biological Chemistry. 272 (31): 19253–60. doi:10.1074/jbc.272.31.19253. PMID 9235919. S2CID 19885003.
- Iyer NV, Leung SW, Semenza GL (September 1998). "The human hypoxia-inducible factor 1alpha gene: HIF1A structure and evolutionary conservation". Genomics. 52 (2): 159–65. doi:10.1006/geno.1998.5416. PMID 9782081.
- "Hypoxia-inducible factor 1-alpha". 2014.
- "HIF1A". National Center for Biotechnology Information.
- "Anti-HIF1 alpha antibody (GTX127309) | GeneTex". www.genetex.com. Retrieved 2019-10-28.
- ^ Minet E, Ernest I, Michel G, Roland I, Remacle J, Raes M, et al. (August 1999). "HIF1A gene transcription is dependent on a core promoter sequence encompassing activating and inhibiting sequences located upstream from the transcription initiation site and cis elements located within the 5'UTR". Biochemical and Biophysical Research Communications. 261 (2): 534–40. doi:10.1006/bbrc.1999.0995. PMID 10425220.
- Danon A, Assouline G (1979). "Antiulcer activity of hypertonic solutions in the rat: possible role of prostaglandins". European Journal of Pharmacology. 58 (4): 425–431. doi:10.1016/0014-2999(79)90313-3. PMID 41725.
- Ladoux A, Frelin C (November 1997). "Cardiac expressions of HIF-1 alpha and HLF/EPAS, two basic loop helix/PAS domain transcription factors involved in adaptative responses to hypoxic stresses". Biochemical and Biophysical Research Communications. 240 (3): 552–6. doi:10.1006/bbrc.1997.7708. PMID 9398602.
- Wiener CM, Booth G, Semenza GL (August 1996). "In vivo expression of mRNAs encoding hypoxia-inducible factor 1". Biochemical and Biophysical Research Communications. 225 (2): 485–8. doi:10.1006/bbrc.1996.1199. PMID 8753788.
- Palmer LA, Semenza GL, Stoler MH, Johns RA (February 1998). "Hypoxia induces type II NOS gene expression in pulmonary artery endothelial cells via HIF-1". The American Journal of Physiology. 274 (2 Pt 1): L212–9. doi:10.1152/ajplung.1998.274.2.L212. PMID 9486205.
- Wenger RH, Kvietikova I, Rolfs A, Gassmann M, Marti HH (February 1997). "Hypoxia-inducible factor-1 alpha is regulated at the post-mRNA level". Kidney International. 51 (2): 560–3. doi:10.1038/ki.1997.79. PMID 9027739.
- Bonello S, Zähringer C, BelAiba RS, Djordjevic T, Hess J, Michiels C, et al. (April 2007). "Reactive oxygen species activate the HIF-1alpha promoter via a functional NFkappaB site". Arteriosclerosis, Thrombosis, and Vascular Biology. 27 (4): 755–61. doi:10.1161/01.ATV.0000258979.92828.bc. PMID 17272744. S2CID 15292804.
- Semenza GL (1999). "Regulation of mammalian O2 homeostasis by hypoxia-inducible factor 1". Annual Review of Cell and Developmental Biology. 15: 551–78. doi:10.1146/annurev.cellbio.15.1.551. PMID 10611972.
- Semenza GL (April 2000). "HIF-1: mediator of physiological and pathophysiological responses to hypoxia". Journal of Applied Physiology. 88 (4): 1474–80. doi:10.1152/jappl.2000.88.4.1474. PMID 10749844. S2CID 2395367.
- ^ Lee JW, Bae SH, Jeong JW, Kim SH, Kim KW (February 2004). "Hypoxia-inducible factor (HIF-1)alpha: its protein stability and biological functions". Experimental & Molecular Medicine. 36 (1): 1–12. doi:10.1038/emm.2004.1. PMID 15031665. S2CID 41613739.
- Semenza GL (2002). "HIF-1 and tumor progression: pathophysiology and therapeutics". Trends in Molecular Medicine. 8 (4 Suppl): S62–7. doi:10.1016/s1471-4914(02)02317-1. PMID 11927290.
- Bruick RK, McKnight SL (November 2001). "A conserved family of prolyl-4-hydroxylases that modify HIF". Science. 294 (5545): 1337–40. Bibcode:2001Sci...294.1337B. doi:10.1126/science.1066373. PMID 11598268. S2CID 9695199.
- ^ Epstein AC, Gleadle JM, McNeill LA, Hewitson KS, O'Rourke J, Mole DR, et al. (October 2001). "C. elegans EGL-9 and mammalian homologs define a family of dioxygenases that regulate HIF by prolyl hydroxylation". Cell. 107 (1): 43–54. doi:10.1016/s0092-8674(01)00507-4. PMID 11595184. S2CID 18372306.
- Ivan M, Kondo K, Yang H, Kim W, Valiando J, Ohh M, et al. (April 2001). "HIFalpha targeted for VHL-mediated destruction by proline hydroxylation: implications for O2 sensing". Science. 292 (5516): 464–8. Bibcode:2001Sci...292..464I. doi:10.1126/science.1059817. PMID 11292862. S2CID 33725562.
- Jaakkola P, Mole DR, Tian YM, Wilson MI, Gielbert J, Gaskell SJ, et al. (April 2001). "Targeting of HIF-alpha to the von Hippel-Lindau ubiquitylation complex by O2-regulated prolyl hydroxylation". Science. 292 (5516): 468–72. Bibcode:2001Sci...292..468J. doi:10.1126/science.1059796. PMID 11292861. S2CID 20914281.
- ^ Masson N, Willam C, Maxwell PH, Pugh CW, Ratcliffe PJ (September 2001). "Independent function of two destruction domains in hypoxia-inducible factor-alpha chains activated by prolyl hydroxylation". The EMBO Journal. 20 (18): 5197–206. doi:10.1093/emboj/20.18.5197. PMC 125617. PMID 11566883.
- Huang LE, Arany Z, Livingston DM, Bunn HF (December 1996). "Activation of hypoxia-inducible transcription factor depends primarily upon redox-sensitive stabilization of its alpha subunit". The Journal of Biological Chemistry. 271 (50): 32253–9. doi:10.1074/jbc.271.50.32253. PMID 8943284. S2CID 11397503.
- Kallio PJ, Pongratz I, Gradin K, McGuire J, Poellinger L (May 1997). "Activation of hypoxia-inducible factor 1alpha: posttranscriptional regulation and conformational change by recruitment of the Arnt transcription factor". Proceedings of the National Academy of Sciences of the United States of America. 94 (11): 5667–72. Bibcode:1997PNAS...94.5667K. doi:10.1073/pnas.94.11.5667. PMC 20836. PMID 9159130.
- Jewell UR, Kvietikova I, Scheid A, Bauer C, Wenger RH, Gassmann M (May 2001). "Induction of HIF-1alpha in response to hypoxia is instantaneous". FASEB Journal. 15 (7): 1312–4. doi:10.1096/fj.00-0732fje. PMID 11344124. S2CID 32080596.
- Hon WC, Wilson MI, Harlos K, Claridge TD, Schofield CJ, Pugh CW, et al. (June 2002). "Structural basis for the recognition of hydroxyproline in HIF-1 alpha by pVHL". Nature. 417 (6892): 975–8. doi:10.1038/nature00767. PMID 12050673. S2CID 4388644.
- ^ Min JH, Yang H, Ivan M, Gertler F, Kaelin WG, Pavletich NP (June 2002). "Structure of an HIF-1alpha -pVHL complex: hydroxyproline recognition in signaling". Science. 296 (5574): 1886–9. Bibcode:2002Sci...296.1886M. doi:10.1126/science.1073440. PMID 12004076. S2CID 19641938.
- Zhao H, Wong RJ, Stevenson DK (September 2021). "The Impact of Hypoxia in Early Pregnancy on Placental Cells". International Journal of Molecular Sciences. 22 (18): 2–8. doi:10.3390/ijms22189675. PMC 8466283. PMID 34575844.
- Hung TH, Charnock-Jones DS, Skepper JN, Burton GJ (March 2004). "Secretion of tumor necrosis factor-alpha from human placental tissues induced by hypoxia-reoxygenation causes endothelial cell activation in vitro: a potential mediator of the inflammatory response in preeclampsia". The American Journal of Pathology. 164 (3): 1049–1061. doi:10.1016/S0002-9440(10)63192-6. PMC 1614718. PMID 14982858.
- ^ Lando D, Peet DJ, Whelan DA, Gorman JJ, Whitelaw ML (February 2002). "Asparagine hydroxylation of the HIF transactivation domain a hypoxic switch". Science. 295 (5556): 858–61. Bibcode:2002Sci...295..858L. doi:10.1126/science.1068592. PMID 11823643. S2CID 24045310.
- Sang N, Fang J, Srinivas V, Leshchinsky I, Caro J (May 2002). "Carboxyl-terminal transactivation activity of hypoxia-inducible factor 1 alpha is governed by a von Hippel-Lindau protein-independent, hydroxylation-regulated association with p300/CBP". Molecular and Cellular Biology. 22 (9): 2984–92. doi:10.1128/mcb.22.9.2984-2992.2002. PMC 133771. PMID 11940656.
- Munkácsy G, Sztupinszki Z, Herman P, Bán B, Pénzváltó Z, Szarvas N, et al. (September 2016). "Validation of RNAi Silencing Efficiency Using Gene Array Data shows 18.5% Failure Rate across 429 Independent Experiments". Molecular Therapy: Nucleic Acids. 5 (9): e366. doi:10.1038/mtna.2016.66. PMC 5056990. PMID 27673562.
- eurekalert.org staff (3 June 2015). "Scientist at LIMR leads study demonstrating drug-induced tissue regeneration". eurekalert.org. Lankenau Institute for Medical Research (LIMR). Retrieved 3 July 2015.
- Zhang Y, Strehin I, Bedelbaeva K, Gourevitch D, Clark L, Leferovich J, et al. (June 2015). "Drug-induced regeneration in adult mice". Science Translational Medicine. 7 (290): 290ra92. doi:10.1126/scitranslmed.3010228. PMC 4687906. PMID 26041709.
- Cho Y, Shin JE, Ewan EE, Oh YM, Pita-Thomas W, Cavalli V (November 2015). "Activating Injury-Responsive Genes with Hypoxia Enhances Axon Regeneration through Neuronal HIF-1α". Neuron. 88 (4): 720–34. doi:10.1016/j.neuron.2015.09.050. PMC 4655162. PMID 26526390.
- Mahar M, Cavalli V (June 2018). "Intrinsic mechanisms of neuronal axon regeneration". Nature Reviews. Neuroscience. 19 (6): 323–337. doi:10.1038/s41583-018-0001-8. PMC 5987780. PMID 29666508.
- Hong WX, Hu MS, Esquivel M, Liang GY, Rennert RC, McArdle A, et al. (May 2014). "The Role of Hypoxia-Inducible Factor in Wound Healing". Advances in Wound Care. 3 (5): 390–399. doi:10.1089/wound.2013.0520. PMC 4005494. PMID 24804159.
- "Skin patch could help heal, prevent diabetic ulcers, study finds". Welcome to Bio-X. © Stanford University, Stanford, California 94305. 2015-01-23. Retrieved 2020-12-04.
- Duscher D, Neofytou E, Wong VW, Maan ZN, Rennert RC, Inayathullah M, et al. (January 2015). "Transdermal deferoxamine prevents pressure-induced diabetic ulcers". Proceedings of the National Academy of Sciences of the United States of America. 112 (1): 94–9. Bibcode:2015PNAS..112...94D. doi:10.1073/pnas.1413445112. PMC 4291638. PMID 25535360.
- Duscher D, Trotsyuk AA, Maan ZN, Kwon SH, Rodrigues M, Engel K, et al. (August 2019). "Optimization of transdermal deferoxamine leads to enhanced efficacy in healing skin wounds". Journal of Controlled Release. 308: 232–239. doi:10.1016/j.jconrel.2019.07.009. PMID 31299261. S2CID 196350143.
- Bonham CA, Rodrigues M, Galvez M, Trotsyuk A, Stern-Buchbinder Z, Inayathullah M, et al. (May 2018). "Deferoxamine can prevent pressure ulcers and accelerate healing in aged mice". Wound Repair and Regeneration. 26 (3): 300–305. doi:10.1111/wrr.12667. PMC 6238634. PMID 30152571.
- Pagani A, Aitzetmüller MM, Brett EA, König V, Wenny R, Thor D, et al. (April 2018). "Skin Rejuvenation through HIF-1α Modulation". Plastic and Reconstructive Surgery. 141 (4): 600e–607e. doi:10.1097/PRS.0000000000004256. PMID 29596193. S2CID 4473259.
- Pagani A, Kirsch BM, Hopfner U, Aitzetmueller MM, Brett EA, Thor D, et al. (June 2020). "Deferiprone Stimulates Aged Dermal Fibroblasts Via HIF-1α Modulation". Aesthetic Surgery Journal. 41 (4): 514–524. doi:10.1093/asj/sjaa142. ISSN 1090-820X. PMID 32479616.
- Duscher D, Maan ZN, Hu MS, Thor D (November 2020). "A single-center blinded randomized clinical trial to evaluate the anti-aging effects of a novel HSF™-based skin care formulation". Journal of Cosmetic Dermatology. 19 (11): 2936–2945. doi:10.1111/jocd.13356. PMID 32306525. S2CID 216031505.
- Houschyar KS, Borrelli MR, Tapking C, Popp D, Puladi B, Ooms M, et al. (2020). "Molecular Mechanisms of Hair Growth and Regeneration: Current Understanding and Novel Paradigms". Dermatology. 236 (4): 271–280. doi:10.1159/000506155. PMID 32163945. S2CID 212693280.
- Tomorrowlabs. "Tomorrowlabs". Tomorrowlabs. Retrieved 2020-12-04.
- "Kosmetikbranche: Wie das Beauty-Start-up Tomorrowlabs den Markt erobert". www.handelsblatt.com (in German). Retrieved 2020-12-04.
- "Ein Protein gegen das Altern und für das Geldverdienen". nachrichten.at (in German). Retrieved 2020-12-04.
- "Das neue Beauty-Investment von Michael Pieper - HZ". Handelszeitung (in German). Retrieved 2020-12-04.
- andrea.hodoschek (2020-08-03). "Milliardenmarkt Anti-Aging: Start-up aus Österreich mischt mit". kurier.at (in German). Retrieved 2020-12-04.
- van Uden P, Kenneth NS, Rocha S (June 2008). "Regulation of hypoxia-inducible factor-1alpha by NF-kappaB". The Biochemical Journal. 412 (3): 477–84. doi:10.1042/BJ20080476. PMC 2474706. PMID 18393939.
- Rius, J., Guma, M., Schachtrup, C. et al. NF-κB links innate immunity to the hypoxic response through transcriptional regulation of HIF-1α. Nature 453, 807–811 (2008). https://doi.org/10.1038/nature06905
- Semenza GL (August 2004). "Hydroxylation of HIF-1: oxygen sensing at the molecular level". Physiology. 19 (4): 176–82. doi:10.1152/physiol.00001.2004. PMID 15304631. S2CID 2434206.
- ^ Xiao H, Gu Z, Wang G, Zhao T (2013). "The possible mechanisms underlying the impairment of HIF-1α pathway signaling in hyperglycemia and the beneficial effects of certain therapies". International Journal of Medical Sciences. 10 (10): 1412–21. doi:10.7150/ijms.5630. PMC 3752727. PMID 23983604.
- ^ Yee Koh M, Spivak-Kroizman TR, Powis G (November 2008). "HIF-1 regulation: not so easy come, easy go". Trends in Biochemical Sciences. 33 (11): 526–34. doi:10.1016/j.tibs.2008.08.002. PMID 18809331.
- ^ Zhong H, De Marzo AM, Laughner E, Lim M, Hilton DA, Zagzag D, et al. (November 1999). "Overexpression of hypoxia-inducible factor 1alpha in common human cancers and their metastases". Cancer Research. 59 (22): 5830–5. PMID 10582706.
- ^ Talks KL, Turley H, Gatter KC, Maxwell PH, Pugh CW, Ratcliffe PJ, et al. (August 2000). "The expression and distribution of the hypoxia-inducible factors HIF-1alpha and HIF-2alpha in normal human tissues, cancers, and tumor-associated macrophages". The American Journal of Pathology. 157 (2): 411–21. doi:10.1016/s0002-9440(10)64554-3. PMC 1850121. PMID 10934146.
- ^ Bos R, van der Groep P, Greijer AE, Shvarts A, Meijer S, Pinedo HM, et al. (March 2003). "Levels of hypoxia-inducible factor-1alpha independently predict prognosis in patients with lymph node negative breast carcinoma". Cancer. 97 (6): 1573–81. doi:10.1002/cncr.11246. PMID 12627523. S2CID 32635739.
- ^ Vaupel P, Mayer A (June 2007). "Hypoxia in cancer: significance and impact on clinical outcome". Cancer and Metastasis Reviews. 26 (2): 225–39. doi:10.1007/s10555-007-9055-1. PMID 17440684. S2CID 21902400.
- ^ Ezzeddini R, Taghikhani M, Somi MH, Samadi N, Rasaee, MJ (May 2019). "Clinical importance of FASN in relation to HIF-1α and SREBP-1c in gastric adenocarcinoma". Life Sciences. 224: 169–176. doi:10.1016/j.lfs.2019.03.056. PMID 30914315. S2CID 85532042.
- Aebersold DM, Burri P, Beer KT, Laissue J, Djonov V, Greiner RH, et al. (April 2001). "Expression of hypoxia-inducible factor-1alpha: a novel predictive and prognostic parameter in the radiotherapy of oropharyngeal cancer". Cancer Research. 61 (7): 2911–6. PMID 11306467.
- Höckel M, Vaupel P (February 2001). "Tumor hypoxia: definitions and current clinical, biologic, and molecular aspects". Journal of the National Cancer Institute. 93 (4): 266–76. doi:10.1093/jnci/93.4.266. PMID 11181773.
- Dvorák K (May 1990). "". Vnitrni Lekarstvi (in Czech). 36 (5): 426–34. PMID 2375073.
- ^ Birner P, Schindl M, Obermair A, Breitenecker G, Oberhuber G (June 2001). "Expression of hypoxia-inducible factor 1alpha in epithelial ovarian tumors: its impact on prognosis and on response to chemotherapy". Clinical Cancer Research. 7 (6): 1661–8. PMID 11410504.
- Ezzeddini R, Taghikhani M, Salek Farrokhi A, Somi MH, Samadi N, Esfahani A, et al. (May 2021). "Downregulation of fatty acid oxidation by involvement of HIF-1α and PPARγ in human gastric adenocarcinoma and its related clinical significance". Journal of Physiology and Biochemistry. 77 (2): 249–260. doi:10.1007/s13105-021-00791-3. PMID 33730333. S2CID 232300877.
- Bos R, Zhong H, Hanrahan CF, Mommers EC, Semenza GL, Pinedo HM, et al. (February 2001). "Levels of hypoxia-inducible factor-1 alpha during breast carcinogenesis". Journal of the National Cancer Institute. 93 (4): 309–14. doi:10.1093/jnci/93.4.309. PMID 11181778.
- ^ Zagzag D, Zhong H, Scalzitti JM, Laughner E, Simons JW, Semenza GL (June 2000). "Expression of hypoxia-inducible factor 1alpha in brain tumors: association with angiogenesis, invasion, and progression". Cancer. 88 (11): 2606–18. doi:10.1002/1097-0142(20000601)88:11<2606::aid-cncr25>3.0.co;2-w. PMID 10861440. S2CID 85168033.
- Neufeld G, Kessler O, Vadasz Z, Gluzman-Poltorak Z (April 2001). "The contribution of proangiogenic factors to the progression of malignant disease: role of vascular endothelial growth factor and its receptors". Surgical Oncology Clinics of North America. 10 (2): 339–56, ix. doi:10.1016/S1055-3207(18)30069-3. PMID 11382591.
- ^ Powis G, Kirkpatrick L (May 2004). "Hypoxia inducible factor-1alpha as a cancer drug target". Molecular Cancer Therapeutics. 3 (5): 647–54. doi:10.1158/1535-7163.647.3.5. PMID 15141023.
- Pietsch T, Valter MM, Wolf HK, von Deimling A, Huang HJ, Cavenee WK, et al. (February 1997). "Expression and distribution of vascular endothelial growth factor protein in human brain tumors". Acta Neuropathologica. 93 (2): 109–17. doi:10.1007/s004010050591. PMID 9039457. S2CID 20164007.
- ^ Krieg M, Haas R, Brauch H, Acker T, Flamme I, Plate KH (November 2000). "Up-regulation of hypoxia-inducible factors HIF-1alpha and HIF-2alpha under normoxic conditions in renal carcinoma cells by von Hippel-Lindau tumor suppressor gene loss of function". Oncogene. 19 (48): 5435–43. doi:10.1038/sj.onc.1203938. PMID 11114720. S2CID 28480163.
- Zundel W, Schindler C, Haas-Kogan D, Koong A, Kaper F, Chen E, et al. (February 2000). "Loss of PTEN facilitates HIF-1-mediated gene expression". Genes & Development. 14 (4): 391–6. doi:10.1101/gad.14.4.391. PMC 316386. PMID 10691731.
- Koukourakis MI, Giatromanolaki A, Skarlatos J, Corti L, Blandamura S, Piazza M, et al. (March 2001). "Hypoxia inducible factor (HIF-1a and HIF-2a) expression in early esophageal cancer and response to photodynamic therapy and radiotherapy". Cancer Research. 61 (5): 1830–2. PMID 11280732.
- Liu XW, Cai TY, Zhu H, Cao J, Su Y, Hu YZ, et al. (January 2014). "Q6, a novel hypoxia-targeted drug, regulates hypoxia-inducible factor signaling via an autophagy-dependent mechanism in hepatocellular carcinoma". Autophagy. 10 (1): 111–22. doi:10.4161/auto.26838. PMC 4389865. PMID 24220190.
- Hogenesch JB, Gu YZ, Jain S, Bradfield CA (May 1998). "The basic-helix-loop-helix-PAS orphan MOP3 forms transcriptionally active complexes with circadian and hypoxia factors". Proceedings of the National Academy of Sciences of the United States of America. 95 (10): 5474–9. Bibcode:1998PNAS...95.5474H. doi:10.1073/pnas.95.10.5474. PMC 20401. PMID 9576906.
- Woods SL, Whitelaw ML (March 2002). "Differential activities of murine single minded 1 (SIM1) and SIM2 on a hypoxic response element. Cross-talk between basic helix-loop-helix/per-Arnt-Sim homology transcription factors". The Journal of Biological Chemistry. 277 (12): 10236–43. doi:10.1074/jbc.M110752200. PMID 11782478. S2CID 25125998.
- Ema M, Hirota K, Mimura J, Abe H, Yodoi J, Sogawa K, et al. (April 1999). "Molecular mechanisms of transcription activation by HLF and HIF1alpha in response to hypoxia: their stabilization and redox signal-induced interaction with CBP/p300". The EMBO Journal. 18 (7): 1905–14. doi:10.1093/emboj/18.7.1905. PMC 1171276. PMID 10202154.
- Bhattacharya S, Michels CL, Leung MK, Arany ZP, Kung AL, Livingston DM (January 1999). "Functional role of p35srj, a novel p300/CBP binding protein, during transactivation by HIF-1". Genes & Development. 13 (1): 64–75. doi:10.1101/gad.13.1.64. PMC 316375. PMID 9887100.
- ^ Park YK, Ahn DR, Oh M, Lee T, Yang EG, Son M, et al. (July 2008). "Nitric oxide donor, (+/-)-S-nitroso-N-acetylpenicillamine, stabilizes transactive hypoxia-inducible factor-1alpha by inhibiting von Hippel-Lindau recruitment and asparagine hydroxylation". Molecular Pharmacology. 74 (1): 236–45. doi:10.1124/mol.108.045278. PMID 18426857. S2CID 31675735.
- Freedman SJ, Sun ZY, Poy F, Kung AL, Livingston DM, Wagner G, et al. (April 2002). "Structural basis for recruitment of CBP/p300 by hypoxia-inducible factor-1 alpha". Proceedings of the National Academy of Sciences of the United States of America. 99 (8): 5367–72. Bibcode:2002PNAS...99.5367F. doi:10.1073/pnas.082117899. PMC 122775. PMID 11959990.
- ^ Mahon PC, Hirota K, Semenza GL (October 2001). "FIH-1: a novel protein that interacts with HIF-1alpha and VHL to mediate repression of HIF-1 transcriptional activity". Genes & Development. 15 (20): 2675–86. doi:10.1101/gad.924501. PMC 312814. PMID 11641274.
- ^ Chen D, Li M, Luo J, Gu W (April 2003). "Direct interactions between HIF-1 alpha and Mdm2 modulate p53 function". The Journal of Biological Chemistry. 278 (16): 13595–8. doi:10.1074/jbc.C200694200. PMID 12606552. S2CID 85351036.
- ^ Ravi R, Mookerjee B, Bhujwalla ZM, Sutter CH, Artemov D, Zeng Q, et al. (January 2000). "Regulation of tumor angiogenesis by p53-induced degradation of hypoxia-inducible factor 1alpha". Genes & Development. 14 (1): 34–44. doi:10.1101/gad.14.1.34. PMC 316350. PMID 10640274.
- ^ Kim BY, Kim H, Cho EJ, Youn HD (February 2008). "Nur77 upregulates HIF-alpha by inhibiting pVHL-mediated degradation". Experimental & Molecular Medicine. 40 (1): 71–83. doi:10.3858/emm.2008.40.1.71. PMC 2679322. PMID 18305400.
- Hansson LO, Friedler A, Freund S, Rudiger S, Fersht AR (August 2002). "Two sequence motifs from HIF-1alpha bind to the DNA-binding site of p53". Proceedings of the National Academy of Sciences of the United States of America. 99 (16): 10305–9. Bibcode:2002PNAS...9910305H. doi:10.1073/pnas.122347199. PMC 124909. PMID 12124396.
- An WG, Kanekal M, Simon MC, Maltepe E, Blagosklonny MV, Neckers LM (March 1998). "Stabilization of wild-type p53 by hypoxia-inducible factor 1alpha". Nature. 392 (6674): 405–8. Bibcode:1998Natur.392..405A. doi:10.1038/32925. PMID 9537326. S2CID 4423081.
- Cho S, Choi YJ, Kim JM, Jeong ST, Kim JH, Kim SH, et al. (June 2001). "Binding and regulation of HIF-1alpha by a subunit of the proteasome complex, PSMA7". FEBS Letters. 498 (1): 62–6. Bibcode:2001FEBSL.498...62C. doi:10.1016/S0014-5793(01)02499-1. PMID 11389899. S2CID 83756271.
- ^ Jung JE, Kim HS, Lee CS, Shin YJ, Kim YN, Kang GH, et al. (October 2008). "STAT3 inhibits the degradation of HIF-1alpha by pVHL-mediated ubiquitination". Experimental & Molecular Medicine. 40 (5): 479–85. doi:10.3858/emm.2008.40.5.479. PMC 2679355. PMID 18985005.
- ^ André H, Pereira TS (October 2008). "Identification of an alternative mechanism of degradation of the hypoxia-inducible factor-1alpha". The Journal of Biological Chemistry. 283 (43): 29375–84. doi:10.1074/jbc.M805919200. PMC 2662024. PMID 18694926.
- Corn PG, McDonald ER, Herman JG, El-Deiry WS (November 2003). "Tat-binding protein-1, a component of the 26S proteasome, contributes to the E3 ubiquitin ligase function of the von Hippel-Lindau protein". Nature Genetics. 35 (3): 229–37. doi:10.1038/ng1254. PMID 14556007. S2CID 22798700.
- Li Z, Wang D, Na X, Schoen SR, Messing EM, Wu G (April 2003). "The VHL protein recruits a novel KRAB-A domain protein to repress HIF-1alpha transcriptional activity". The EMBO Journal. 22 (8): 1857–67. doi:10.1093/emboj/cdg173. PMC 154465. PMID 12682018.
- Tanimoto K, Makino Y, Pereira T, Poellinger L (August 2000). "Mechanism of regulation of the hypoxia-inducible factor-1 alpha by the von Hippel-Lindau tumor suppressor protein". The EMBO Journal. 19 (16): 4298–309. doi:10.1093/emboj/19.16.4298. PMC 302039. PMID 10944113.
- Yu F, White SB, Zhao Q, Lee FS (August 2001). "HIF-1alpha binding to VHL is regulated by stimulus-sensitive proline hydroxylation". Proceedings of the National Academy of Sciences of the United States of America. 98 (17): 9630–5. Bibcode:2001PNAS...98.9630Y. doi:10.1073/pnas.181341498. PMC 55503. PMID 11504942.
- Haase VH (2009). "The VHL tumor suppressor: master regulator of HIF". Current Pharmaceutical Design. 15 (33): 3895–903. doi:10.2174/138161209789649394. PMC 3622710. PMID 19671042.
- Sun YY, Wang CY, Hsu MF, Juan SH, Chang CY, Chou CM, et al. (July 2010). "Glucocorticoid protection of oligodendrocytes against excitotoxin involving hypoxia-inducible factor-1alpha in a cell-type-specific manner". The Journal of Neuroscience. 30 (28): 9621–30. doi:10.1523/JNEUROSCI.2295-10.2010. PMC 6632428. PMID 20631191.
- Menshanov PN, Bannova AV, Dygalo NN (January 2017). "Anoxia ameliorates the dexamethasone-induced neurobehavioral alterations in the neonatal male rat pups". Hormones and Behavior. 87: 122–128. doi:10.1016/j.yhbeh.2016.11.013. PMID 27865789. S2CID 4108143.
Further reading
- Semenza GL (August 2000). "HIF-1 and human disease: one highly involved factor". Genes & Development. 14 (16): 1983–91. doi:10.1101/gad.14.16.1983. PMID 10950862. S2CID 12788170.
- Semenza G (September 2002). "Signal transduction to hypoxia-inducible factor 1". Biochemical Pharmacology. 64 (5–6): 993–8. doi:10.1016/S0006-2952(02)01168-1. PMID 12213597.
- Arbeit JM (2002). "Quiescent hypervascularity mediated by gain of HIF-1 alpha function". Cold Spring Harbor Symposia on Quantitative Biology. 67: 133–42. doi:10.1101/sqb.2002.67.133. PMID 12858534.
- Sitkovsky M, Lukashev D (September 2005). "Regulation of immune cells by local-tissue oxygen tension: HIF1 alpha and adenosine receptors". Nature Reviews. Immunology. 5 (9): 712–21. doi:10.1038/nri1685. PMID 16110315. S2CID 30400163.
- Mobasheri A, Richardson S, Mobasheri R, Shakibaei M, Hoyland JA (October 2005). "Hypoxia inducible factor-1 and facilitative glucose transporters GLUT1 and GLUT3: putative molecular components of the oxygen and glucose sensing apparatus in articular chondrocytes". Histology and Histopathology. 20 (4): 1327–38. doi:10.14670/HH-20.1327. PMID 16136514.
- Schipani E (2006). "Hypoxia and HIF-1 alpha in chondrogenesis". Seminars in Cell & Developmental Biology. 16 (4–5): 539–46. doi:10.1016/j.semcdb.2005.03.003. PMID 16144691.
- Haase VH (August 2006). "Hypoxia-inducible factors in the kidney". American Journal of Physiology. Renal Physiology. 291 (2): F271–81. doi:10.1152/ajprenal.00071.2006. PMC 4232221. PMID 16554418.
- Liang D, Kong X, Sang N (November 2006). "Effects of histone deacetylase inhibitors on HIF-1". Cell Cycle. 5 (21): 2430–5. doi:10.4161/cc.5.21.3409. PMC 4505804. PMID 17102633.
External links
- Overview of all the structural information available in the PDB for UniProt: Q16665 (Human Hypoxia-inducible factor 1-alpha) at the PDBe-KB.
- Overview of all the structural information available in the PDB for UniProt: Q61221 (Mouse Hypoxia-inducible factor 1-alpha) at the PDBe-KB.
- Scientific animation of HIF-1alpha in complex with ARNT on DNA: https://www.youtube.com/watch?v=azIEzLXXyHM
| PDB gallery | |
|---|---|
|
| Transcription factors and intracellular receptors | |||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| |||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||
| see also transcription factor/coregulator deficiencies | |||||||||||||||||||||||||||||||