The heteronuclear single quantum coherence or heteronuclear single quantum correlation experiment, normally abbreviated as HSQC, is used frequently in NMR spectroscopy of organic molecules and is of particular significance in the field of protein NMR. The experiment was first described by Geoffrey Bodenhausen and D. J. Ruben in 1980. The resulting spectrum is two-dimensional (2D) with one axis for proton (H) and the other for a heteronucleus (an atomic nucleus other than a proton), which is usually C or N. The spectrum contains a peak for each unique proton attached to the heteronucleus being considered. The 2D HSQC can also be combined with other experiments in higher-dimensional NMR experiments, such as NOESY-HSQC or TOCSY-HSQC.
General scheme
The HSQC experiment is a highly sensitive 2D-NMR experiment and was first described in a H—N system, but is also applicable to other nuclei such as H—C and H—P. The basic scheme of this experiment involves the transfer of magnetization on the proton to the second nucleus, which may be N, C or P, via an INEPT (Insensitive nuclei enhanced by polarization transfer) step. After a time delay (t1), the magnetization is transferred back to the proton via a retro-INEPT step and the signal is then recorded. In HSQC, a series of experiments is recorded where the time delay t1 is incremented. The H signal is detected in the directly measured dimension in each experiment, while the chemical shift of N or C is recorded in the indirect dimension which is formed from the series of experiments.
HSQC in protein NMR
Main article: protein NMRH—N HSQC

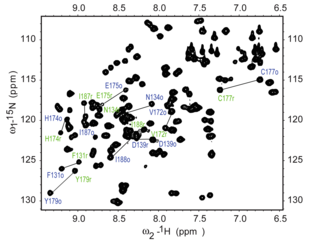
The N HSQC experiment is one of the most frequently recorded experiments in protein NMR. The HSQC experiment can be performed using the natural abundance of the N isotope, but normally for protein NMR, isotopically labeled proteins are used. Such labelled proteins are usually produced by expressing the protein in cells grown in N-labelled media.
Each residue of the protein, with the exception of proline, has an amide proton attached to a nitrogen in the peptide bond. The HSQC provides the correlation between the nitrogen and amide proton, and each amide yields a peak in the HSQC spectra. Each residue (except proline) therefore can produce an observable peak in the spectra, although in practice not all the peaks are always seen due to a number of factors. Normally the N-terminal residue (which has an NH3 group attached) is not readily observable due to exchange with solvent. In addition to the backbone amide resonances, sidechains with nitrogen-bound protons will also produce peaks.
In a typical HSQC spectrum, the NH2 peaks from the sidechains of asparagine and glutamine appear as doublets on the top right corner, and a smaller peak may appear on top of each peak due to deuterium exchange from the D2O normally added to an NMR sample, giving these sidechain peaks a distinctive appearance. The sidechain amine peaks from tryptophan are usually shifted downfield and appear near the bottom left corner. The backbone amide peaks of glycine normally appear near the top of the spectrum.
The N HSQC is normally the first heteronuclear spectrum acquired for the assignment of resonances where each amide peak is assigned to a particular residue in the protein. If the protein is folded, the peaks are usually well-dispersed, and most of the individual peaks can be distinguished. If there is a large cluster of severely overlapped peaks around the middle of the spectrum, that would indicate the presence of significant unstructured elements in the protein. In such cases where there are severe overlap of resonances the assignment of resonances in the spectra can be difficult. The assignment of the HSQC spectrum requires other experiments, ideally using triple resonance experiments with N and C-labelled proteins, that provide sequential connectivities between residues so that the resonances can be linked to particular residues and sequentially assigned. The assignment of the spectrum is essential for a meaningful interpretation of more advanced NMR experiments such as structure determination and relaxation analysis.
Chemicals labelled with N isotope are relatively inexpensive, and the N HSQC is a sensitive experiment whereby a spectrum can be acquired in a relatively short time, the N HSQC is therefore often used to screen candidates for their suitability for structure determination by NMR, as well as optimization of the sample conditions. The time-consuming process of structure determination is usually not undertaken until a good HSQC spectrum can be obtained. The HSQC experiment is also useful for detecting binding interface in protein-protein interaction, as well the interactions with ligands such as drugs. By comparing the HSQC of the free protein with the one bound to the ligand, changes in the chemical shifts of some peaks may be observed, and these peaks are likely to lie on the binding surface where the binding perturbed their chemical shifts. The N HSQC may also be used in relaxation analysis in the studies of molecular dynamics of proteins, the determination of ionization constant, and other studies.
H—C HSQC
This experiment provides correlations between a carbon and its attached protons. The constant time (CT) version of H—C HSQC is normally used as it circumvents the issue of splitting of signal due to homonuclear C—C J couplings which reduces spectral resolution. The "constant time" refers to the entire evolution period between the two INEPT steps which is kept constant in this experiment. If this evolution period is set to be the inverse of the J-coupling constant, then the sign of the magnetization of those carbons with an odd number of aliphatic carbon attached will be opposite to those with an even number. For example, if the Cβ of leucine appears as a positive peak (2 aliphatic carbons attached), then the Cγ (3 aliphatic carbons attached) and Cα (1 aliphatic carbons attached) would appear negative.
HSQC in lipid NMR
H—P HSQC
The use of H—P HSQC is relatively uncommon in lipidomics, however use of P in lipidomics dates back to the 1990s. The use of this technique is limited with respect to mass spectrometry due to its requirement for much bigger sample size, however the combination of H—P HSQC with mass spectrometry is regarded as a thorough approach to lipidomics and techniques for 'dual spectroscopy' are becoming available.
See also
References
- Bodenhausen, G.; Ruben, D.J. (1980). "Natural abundance nitrogen-15 NMR by enhanced heteronuclear spectroscopy". Chemical Physics Letters. 69 (1): 185–189. Bibcode:1980CPL....69..185B. doi:10.1016/0009-2614(80)80041-8. S2CID 96730420.
- Wu, Bin; Skarina, Tatiana, Yee, Adelinda, Jobin, Marie-Claude, DiLeo, Rosa, Semesi, Anthony, Fares, Christophe, Lemak, Alexander, Coombes, Brian K., Arrowsmith, Cheryl H., Singer, Alexander U., Savchenko, Alexei, Stebbins, C. Erec (June 2010). "NleG Type 3 Effectors from Enterohaemorrhagic Escherichia coli Are U-Box E3 Ubiquitin Ligases". PLOS Pathogens. 6 (6): e1000960. doi:10.1371/journal.ppat.1000960. PMC 2891834. PMID 20585566.
{{cite journal}}: CS1 maint: multiple names: authors list (link) - Steven M. Pascal (2008). NMR Primer: An HSQC-based Approach with Vector Animations. IM Publications LLP. pp. 29–31. ISBN 978-1901019087.
- Williamson, Mike P. (2013-08-01). "Using chemical shift perturbation to characterise ligand binding". Progress in Nuclear Magnetic Resonance Spectroscopy. 73: 1–16. Bibcode:2013PNMRS..73....1W. doi:10.1016/j.pnmrs.2013.02.001. ISSN 0079-6565. PMID 23962882.
- Geerten W. Vuister; Ad Bax (1992). "Resolution Enhancement and Spectral Editing of Uniformly 13CEnriched Proteins by Homonuclear Broadband 13C Decoupling" (PDF). Journal of Magnetic Resonance. 98 (2): 428–435. Bibcode:1992JMagR..98..428V. doi:10.1016/0022-2364(92)90144-v.
- Bosco, M.; Culeddu, N.; Toffanin, R.; Pollesello, P. (1997). "Organic Solvent Systems for 31P Nuclear Magnetic Resonance Analysis of Lecithin Phospholipids: Applications to Two-Dimensional Gradient-Enhanced1H-Detected Heteronuclear Multiple Quantum Coherence Experiments". Analytical Biochemistry. 245 (1): 38–47. doi:10.1006/abio.1996.9907. PMID 9025966.
- Furse, Samuel; Fernandez-Twinn, Denise; Jenkins, Benjamin; Meek, Claire L.; Williams, Huw E.; Smith, Gordon C. S.; Charnock-Jones, D. Stephen; Ozanne Susan, E.; Koulman, Albert (2020). "A high throughput platform for detailed lipidomic analysis of a range of mouse and human tissues". Analytical and Bioanalytical Chemistry. 412 (12): 2851–2862. doi:10.1007/s00216-020-02511-0. PMC 7196091. PMID 32144454.
General references
- Protein NMR Spectroscopy : Principles and Practice (1995) John Cavanagh, Wayne J. Fairbrother, Arthur G. Palmer III, Nicholas J. Skelton, Academic Press
External links
- Protein NMR Protein NMR spectra