| 3-mercaptopyruvate sulfurtransferase | |||||||||
|---|---|---|---|---|---|---|---|---|---|
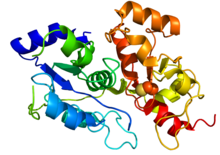 Crystallographic structure of 3-mercaptopyruvate sulfurtransferase based on PDB: 3OLH. Rainbow colored (N-terminus blue and C-terminus red). The catalytic cysteine is depicted by spheres. Crystallographic structure of 3-mercaptopyruvate sulfurtransferase based on PDB: 3OLH. Rainbow colored (N-terminus blue and C-terminus red). The catalytic cysteine is depicted by spheres. | |||||||||
| Identifiers | |||||||||
| EC no. | 2.8.1.2 | ||||||||
| CAS no. | 9026-05-5 | ||||||||
| Databases | |||||||||
| IntEnz | IntEnz view | ||||||||
| BRENDA | BRENDA entry | ||||||||
| ExPASy | NiceZyme view | ||||||||
| KEGG | KEGG entry | ||||||||
| MetaCyc | metabolic pathway | ||||||||
| PRIAM | profile | ||||||||
| PDB structures | RCSB PDB PDBe PDBsum | ||||||||
| Gene Ontology | AmiGO / QuickGO | ||||||||
| |||||||||
| 3-mercaptopyruvate sulfurtransferase | |||||||
|---|---|---|---|---|---|---|---|
| Identifiers | |||||||
| Symbol | MPST | ||||||
| Alt. symbols | MST, TST2, TUM1 | ||||||
| Alt. names | human liver rhodanese | ||||||
| NCBI gene | 4357 | ||||||
| HGNC | 7223 | ||||||
| OMIM | 602496 | ||||||
| PDB | 3OLH | ||||||
| RefSeq | NP_066949 | ||||||
| UniProt | P25325 | ||||||
| Other data | |||||||
| EC number | 2.8.1.2 | ||||||
| Locus | Chr. 22 q123 | ||||||
| |||||||
In enzymology, a 3-mercaptopyruvate sulfurtransferase (EC 2.8.1.2) is an enzyme that catalyzes the chemical reactions of 3-mercaptopyruvate. This enzyme belongs to the family of transferases, specifically the sulfurtransferases. This enzyme participates in cysteine metabolism. It is encoded by the MPST gene.
The enzyme is of interest because it provides a pathway for detoxification of cyanide, especially since it occurs widely in the cytosol and distributed broadly.
Nomenclature
The systematic name of this enzyme class is 3-mercaptopyruvate:cyanide sulfurtransferase. This enzyme is also called beta-mercaptopyruvate sulfurtransferase and in the older literature, human liver rhodanese.
Structure
Gene
The MPST gene lies on the chromosome location of 22q12.3 and consists of 6 exons. Alternatively spliced transcript variants encoding the same protein have been identified.
Protein
The encoded cytoplasmic protein is a member of the rhodanese family but is not rhodanese itself, which is found only in mitochondria. MPST protein consists of 317 amino acid residues and weighs 35250Da. MPST contains two rhodanese domains with similar secondary structures suggesting a common evolutionary origin. The catalytic cysteine residue only exists in the C-terminal rhodanese domain. The protein can function as a monomer or as a disulfide-linked homodimer.
Function and mechanism
The biological function of MPST remains unclear. It may be involved in cyanide detoxification, biosynthesis of thiosulfate, production of the signalling molecule hydrogen sulfide, or the degradation of cysteine.
MPST reacts with 3-MP to convert a cysteinyl residue to the persulfide-containing intermediate:
- RSH + HSCH2C(O)CO2 → RSSH + CH3C(O)CO2
The persulfide group is labile, and can transfer S to other groups, such as cyanide. It is also susceptible to reduction to release hydrogen sulfide.
H2S is produced from l-cysteine by cystathionine-γ-lyase (CSE) and MPST. l-cysteine-dependent production of H2S by MPST is a two-step reaction. In the first step, cysteine transaminase converts l-cysteine along with α-ketoglutarate into 3-mercaptopyruvate (3-MP). Subsequently, MPST converts 3-MP into H2S. MPST catalyzes the transfer of a sulfur atom from mercaptopyruvate to sulfur acceptors like cyanides or thiol compounds. Thus it is also considered to participate in cysteine degradation. MPST generates H2S in coronary artery, mediating its effects through direct modulation of NO, namely vasodilation. This has important implications for H2S-based therapy in healthy and diseased coronary arteries.
Biological significance
MPST is expressed in a number of tissues, including kidney, liver, lung, heart, muscle, spleen, and brain.
3-Mercaptopyruvate (3-MP), along with α-ketoglutarate, is derived from cysteine by the action of an aminotransferase. Thus MPST may participate in cysteine degradation.
Biomedical
Hydrogen sulfide production
H2S is produced from MPST (as well as by cystathionine-γ-lyase). MPST generates H2S in coronary artery, mediating its effects through direct modulation of NO, namely vasodilation. This has important implications for H2S-based therapy in healthy and diseased coronary arteries.
Mercaptolactate-cysteine disulfiduria
Deficiency in MPST has been found related to mercaptolactate-cysteine disulfiduria (or Ampola syndrome), a rare inheritable disorder associated with oversecretion of mercaptolactate-cysteine disulfide in urine. Fewer than 1,000 people have this rare disorder in the United States. Symptoms and signs include: high forehead, seizures, arachnodactyly, genu valgum, hypolasia of the ear cartilage and umbilical hernia.
Bladder cancer
Immunoreactivity for MPST was detected in malignant uroepithelium and muscular layer of bladder cancer samples. Protein levels and catalytic activities of MPST increased with the increase of malignant degrees in human bladder tissues and human urothelial cell carcinoma of bladder (UCB) cell lines and this may promote the application of MPST novel enzymes to UCB diagnosis or treatment.
References
- ^ Yadav PK, Yamada K, Chiku T, Koutmos M, Banerjee R (July 2013). "Structure and kinetic analysis of H2S production by human mercaptopyruvate sulfurtransferase". The Journal of Biological Chemistry. 288 (27): 20002–13. doi:10.1074/jbc.M113.466177. PMC 3707699. PMID 23698001.
- ^ "Entrez Gene: MPST mercaptopyruvate sulfurtransferase".
- ^ Patterson SE, Moeller B, Nagasawa HT, Vince R, Crankshaw DL, Briggs J, Stutelberg MW, Vinnakota CV, Logue BA (June 2016). "Development of sulfanegen for mass cyanide casualties". Annals of the New York Academy of Sciences. 1374 (1): 202–9. Bibcode:2016NYASA1374..202P. doi:10.1111/nyas.13114. PMC 4940216. PMID 27308865.
- Pallini R, Guazzi GC, Cannella C, Cacace MG (October 1991). "Cloning and sequence analysis of the human liver rhodanese: comparison with the bovine and chicken enzymes". Biochemical and Biophysical Research Communications. 180 (2): 887–93. doi:10.1016/S0006-291X(05)81148-9. PMID 1953758.
- ^ Nagahara N, Okazaki T, Nishino T (July 1995). "Cytosolic mercaptopyruvate sulfurtransferase is evolutionarily related to mitochondrial rhodanese. Striking similarity in active site amino acid sequence and the increase in the mercaptopyruvate sulfurtransferase activity of rhodanese by site-directed mutagenesis". The Journal of Biological Chemistry. 270 (27): 16230–5. doi:10.1074/jbc.270.27.16230. PMID 7608189.
- ^ Shibuya N, Mikami Y, Kimura Y, Nagahara N, Kimura H (November 2009). "Vascular endothelium expresses 3-mercaptopyruvate sulfurtransferase and produces hydrogen sulfide". Journal of Biochemistry. 146 (5): 623–6. doi:10.1093/jb/mvp111. PMID 19605461.
- Vachek H, Wood JL (January 1972). "Purification and properties of mercaptopyruvate sulfur transferase of Escherichia coli". Biochimica et Biophysica Acta (BBA) - Enzymology. 258 (1): 133–46. doi:10.1016/0005-2744(72)90973-4. PMID 4550801.
- ^ Kuo MM, Kim DH, Jandu S, Bergman Y, Tan S, Wang H, Pandey DR, Abraham TP, Shoukas AA, Berkowitz DE, Santhanam L (January 2016). "MPST but not CSE is the primary regulator of hydrogen sulfide production and function in the coronary artery". American Journal of Physiology. Heart and Circulatory Physiology. 310 (1): H71–9. doi:10.1152/ajpheart.00574.2014. PMC 4796461. PMID 26519030.
- Billaut-Laden I, Rat E, Allorge D, Crunelle-Thibaut A, Cauffiez C, Chevalier D, Lo-Guidice JM, Broly F (August 2006). "Evidence for a functional genetic polymorphism of the human mercaptopyruvate sulfurtransferase (MPST), a cyanide detoxification enzyme". Toxicology Letters. 165 (2): 101–11. doi:10.1016/j.toxlet.2006.02.002. PMID 16545926.
- Ubuka T, Hosaki Y, Nishina H, Ikeda T (1985). "3-Mercaptopyruvate sulfurtransferase activity in guinea pig and rat tissues". Physiological Chemistry and Physics and Medical NMR. 17 (1): 41–3. PMID 3862140.
- Nagahara N, Ito T, Kitamura H, Nishino T (September 1998). "Tissue and subcellular distribution of mercaptopyruvate sulfurtransferase in the rat: confocal laser fluorescence and immunoelectron microscopic studies combined with biochemical analysis". Histochemistry and Cell Biology. 110 (3): 243–50. doi:10.1007/s004180050286. PMID 9749958. S2CID 25430792.
- Sörbo B (1987). "3-Mercaptopyruvate, 3-mercaptolactate and mercaptoacetate". Sulfur and Sulfur Amino Acids. Methods in Enzymology. Vol. 143. pp. 178–82. doi:10.1016/0076-6879(87)43033-4. ISBN 9780121820435. PMID 3657534.
- "Beta-mercaptolactate cysteine disulfiduria - About the Disease - Genetic and Rare Diseases Information Center". rarediseases.info.nih.gov. Retrieved 2024-03-09.
- Gai JW, Qin W, Liu M, Wang HF, Zhang M, Li M, Zhou WH, Ma QT, Liu GM, Song WH, Jin J, Ma HS (April 2016). "Expression profile of hydrogen sulfide and its synthases correlates with tumor stage and grade in urothelial cell carcinoma of bladder". Urologic Oncology. 34 (4): 166.e15–20. doi:10.1016/j.urolonc.2015.06.020. PMID 26847849.
Further reading (mainly older literature)
- Fiedler H, Wood JL (September 1956). "Specificity studies on the beta-mercaptopyruvate-cyanide transsulfuration system". The Journal of Biological Chemistry. 222 (1): 387–97. doi:10.1016/S0021-9258(19)50803-1. PMID 13367011.
- Hylin JW, Wood JL (August 1959). "Enzymatic formation of polysulfides from mercaptopyruvate". The Journal of Biological Chemistry. 234 (8): 2141–4. doi:10.1016/S0021-9258(18)69881-3. PMID 13673028.
- Sorbo B (May 1957). "Enzymic transfer of sulfur from mercaptopyruvate to sulfate or sulfinates". Biochimica et Biophysica Acta. 24 (2): 324–9. doi:10.1016/0006-3002(57)90201-9. PMID 13436433.
- Vachek H, Wood JL (January 1972). "Purification and properties of mercaptopyruvate sulfur transferase of Escherichia coli". Biochimica et Biophysica Acta (BBA) - Enzymology. 258 (1): 133–46. doi:10.1016/0005-2744(72)90973-4. PMID 4550801.
- Van den Hamer CJ, Morell AG, Scheinberg IH (May 1967). "A study of the cooper content of beta-mercaptopyruvate trans-sulfurase". The Journal of Biological Chemistry. 242 (10): 2514–6. doi:10.1016/S0021-9258(18)95992-2. PMID 6026243.
| Enzymes | |
|---|---|
| Activity | |
| Regulation | |
| Classification | |
| Kinetics | |
| Types |
|