| This article includes a list of general references, but it lacks sufficient corresponding inline citations. Please help to improve this article by introducing more precise citations. (February 2016) (Learn how and when to remove this message) |
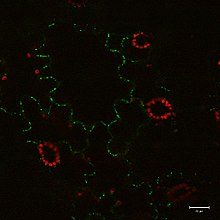
A movement protein (MP) is a specific virus-encoded protein that is thought to be a general feature of plant genomes. For a virus to infect a plant, it must be able to move between cells so it can spread throughout the plant. Plant cell walls make this moving/spreading quite difficult and therefore, for this to occur, movement proteins must be present. Movement proteins allow for local and systemic viral spread throughout a plant. MPs were first studied in the Tobacco Mosaic Virus (TMV), where it was found that viruses were unable to spread without the presence of a specific protein. In general, the plant viruses first move within the cell from replication sites to the plasmodesmata (PD). Then, the virus can go through the PD and spread to other cells. This process is controlled through MPs. Different MPs use different mechanisms and pathways to regulate the spread of some viruses. Nearly all plants express at least one MP, while some can encode many different MPs which help with cell-to-cell viral transmission. They serve to increase the size exclusion limits (SEL) of plasmodesmata to allow for greater spread of the virus.
Plant viral movement protein regulation
Viral MPs can undergo some sort of regulation. They can be phosphorylated by plant protein kinases which can inactivate the viral MPs and provide an avenue for post-translational modification and regulation of viral movement. Phosphorylation also can assist in regulating viral infectivity. Plasmodesmata function can regulate the stability of MP-vNA complexes which are formed for viruses to be transported via the movement protein. Phosphorylation during the tobacco mosaic virus-MP-vRNA transport could be responsible for playing a role in regulating the degree of infectivity of the virus.
The function of movement proteins
Movement proteins can assist in unraveling key mechanisms that help control and regulate macromolecule transport within and between plant cells. MPs can use plasmodesmata, however, they are also able to alter and intercept intercellular channels based on if they are fully differentiated or if they are developing cells. When MPs are actively being expressed, the cell wall barrier to the movement of plant viruses is eliminated which can imply that movement proteins can play a role in changing cell architecture. MPs and other viral components can interact with the endomembrane system along with the cytoskeletal network right before the virus crosses the cell wall. These interactions occur in order to identify the viral genome and direct it to the cell wall for transport. Different viral encoded MPs are responsible for interfering with plasmodesmal gating. Research has suggested that there could even be plasmodesmal targeting sequences within movement proteins and that these proteins could even serve as tools to identify certain components of the plasmodesmata. There have not been extensive similarities in sequences in MPs that belong to different plant virus taxonomic groups. Additionally, some transport systems for viruses just need a single MP while others may need additional virus encoded proteins in order to facilitate the transport of viral genomes.
Mechanisms of movement proteins
There are multiple different mechanisms that MPs can use. The 30-kDa MP found in the TMV, the prototype of the 30K MP superfamily, has been shown to alter the size exclusion limit of PD. It is also able to bind ssRNAs and also may pass through plasmodesmata as an RNP complex containing virus genomic RNA. Some MPs have the necessary protein motifs to undergo cell to cell movement without the help of other virus-specific proteins. These MPs are able to sequence non-specific RNA binding and help the movement of other viruses that are unable to transport themselves. Another type of MP mechanism involves the movement of the plasmodesmata internal structures such as the desmotubule and the transmission of entire virions, from infected cells to adjacent cells.
Origin and evolution
Movement proteins of the 30K superfamily, one of the most widespread groups of MPs found in viruses from 16 different families, share the conserved jelly-roll fold domain with the capsid proteins (CPs) of small icosahedral RNA and DNA viruses, in particular, those infecting plants. It has been suggested that the 30K MPs evolved via duplication or horizontal acquisition of the CP gene in a virus that infected an ancestor of vascular plants, followed by exaptation of one of the paralogous CPs. During the subsequent coevolution of viruses with diversifying vascular plants, the 30K MP genes underwent explosive horizontal spread among emerging plant viruses, driving the diversification of the 30K MP superfamily and molding the contemporary plant virome.
References
- ^ Mushegian, A. R.; Elena, S. F. (2015-07-01). "Evolution of plant virus movement proteins from the 30K superfamily and of their homologs integrated in plant genomes". Virology. 72 (476): 304–315. doi:10.1016/j.virol.2014.12.012. hdl:10261/109217. PMID 25576984.
- ^ Taliansky, Michael; Torrance, Lesley; Kalinina, Natalia O. (2008), Foster, Gary D.; Johansen, I. Elisabeth; Hong, Yiguo; Nagy, Peter D. (eds.), "Role of Plant Virus Movement Proteins", Plant Virology Protocols, Methods in Molecular Biology, vol. 451, Totowa, NJ: Humana Press, pp. 33–54, doi:10.1007/978-1-59745-102-4_3, ISBN 978-1-58829-827-0, PMID 18370246, retrieved 2022-05-04
- ^ Lucas, William J. (2006). "Plant viral movement proteins: Agents for cell-to-cell trafficking of viral genomes". Virology. 344 (1): 169–184. doi:10.1016/j.virol.2005.09.026. PMID 16364748.
- ^ Lazarowitz, Sondra G.; Beachy, Roger N. (1999). "Viral Movement Proteins as Probes for Intracellular and Intercellular Trafficking in Plants". The Plant Cell. 11 (4): 535–548. doi:10.1105/tpc.11.4.535. ISSN 1040-4651. PMC 144200. PMID 10213776.
- Lee, Jung-Youn; Lucas, William J (2001). "Phosphorylation of viral movement proteins – regulation of cell-to-cell trafficking". Trends in Microbiology. 9 (1): 5–8. doi:10.1016/S0966-842X(00)01901-6. PMID 11166222.
- Morozov, Sergey Y.; Solovyev, Andrey G.; Morozov, Sergey Y.; Solovyev, Andrey G. (2020). "Small hydrophobic viral proteins involved in intercellular movement of diverse plant virus genomes". AIMS Microbiology. 6 (3): 305–329. doi:10.3934/microbiol.2020019. ISSN 2471-1888. PMC 7595835. PMID 33134746.
- ^ Butkovic, A; Dolja, VV; Koonin, EV; Krupovic, M (2023). "Plant virus movement proteins originated from jelly-roll capsid proteins". PLOS Biology. 21 (6): e3002157. doi:10.1371/journal.pbio.3002157. PMC 10306228. PMID 37319262.
 This article incorporates text from this source, which is available under the CC0 license.
This article incorporates text from this source, which is available under the CC0 license.