In molecular biology and genetics, the sense of a nucleic acid molecule, particularly of a strand of DNA or RNA, refers to the nature of the roles of the strand and its complement in specifying a sequence of amino acids. Depending on the context, sense may have slightly different meanings. For example, the negative-sense strand of DNA is equivalent to the template strand, whereas the positive-sense strand is the non-template strand whose nucleotide sequence is equivalent to the sequence of the mRNA transcript.
DNA sense
Because of the complementary nature of base-pairing between nucleic acid polymers, a double-stranded DNA molecule will be composed of two strands with sequences that are reverse complements of each other. To help molecular biologists specifically identify each strand individually, the two strands are usually differentiated as the "sense" strand and the "antisense" strand. An individual strand of DNA is referred to as positive-sense (also positive (+) or simply sense) if its nucleotide sequence corresponds directly to the sequence of an RNA transcript which is translated or translatable into a sequence of amino acids (provided that any thymine bases in the DNA sequence are replaced with uracil bases in the RNA sequence). The other strand of the double-stranded DNA molecule is referred to as negative-sense (also negative (−) or antisense), and is reverse complementary to both the positive-sense strand and the RNA transcript. It is actually the antisense strand that is used as the template from which RNA polymerases construct the RNA transcript, but the complementary base-pairing by which nucleic acid polymerization occurs means that the sequence of the RNA transcript will look identical to the positive-sense strand, apart from the RNA transcript's use of uracil instead of thymine.
Sometimes the phrases coding strand and template strand are encountered in place of sense and antisense, respectively, and in the context of a double-stranded DNA molecule the usage of these terms is essentially equivalent. However, the coding/sense strand need not always contain a code that is used to make a protein; both protein-coding and non-coding RNAs may be transcribed.
The terms "sense" and "antisense" are relative only to the particular RNA transcript in question, and not to the DNA strand as a whole. In other words, either DNA strand can serve as the sense or antisense strand. Most organisms with sufficiently large genomes make use of both strands, with each strand functioning as the template strand for different RNA transcripts in different places along the same DNA molecule. In some cases, RNA transcripts can be transcribed in both directions (i.e. on either strand) from a common promoter region, or be transcribed from within introns on either strand (see "ambisense" below).
Sense DNA
The DNA sense strand looks like the messenger RNA (mRNA) transcript, and can therefore be used to read the expected codon sequence that will ultimately be used during translation (protein synthesis) to build an amino acid sequence and then a protein. For example, the sequence "ATG" within a DNA sense strand corresponds to an "AUG" codon in the mRNA, which codes for the amino acid methionine. However, the DNA sense strand itself is not used as the template for the mRNA; it is the DNA antisense strand that serves as the source for the protein code, because, with bases complementary to the DNA sense strand, it is used as a template for the mRNA. Since transcription results in an RNA product complementary to the DNA template strand, the mRNA is complementary to the DNA antisense strand.
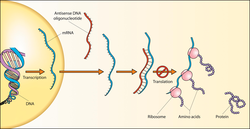
Hence, a base triplet 3′-TAC-5′ in the DNA antisense strand (complementary to the 5′-ATG-3′ of the DNA sense strand) is used as the template which results in a 5′-AUG-3′ base triplet in the mRNA. The DNA sense strand will have the triplet ATG, which looks similar to the mRNA triplet AUG but will not be used to make methionine because it will not be directly used to make mRNA. The DNA sense strand is called a "sense" strand not because it will be used to make protein (it won't be), but because it has a sequence that corresponds directly to the RNA codon sequence. By this logic, the RNA transcript itself is sometimes described as "sense".
Example with double-stranded DNA
- DNA strand 1: antisense strand (transcribed to) → RNA strand (sense)
- DNA strand 2: sense strand
Some regions within a double-stranded DNA molecule code for genes, which are usually instructions specifying the order in which amino acids are assembled to make proteins, as well as regulatory sequences, splicing sites, non-coding introns, and other gene products. For a cell to use this information, one strand of the DNA serves as a template for the synthesis of a complementary strand of RNA. The transcribed DNA strand is called the template strand, with antisense sequence, and the mRNA transcript produced from it is said to be sense sequence (the complement of antisense). The untranscribed DNA strand, complementary to the transcribed strand, is also said to have sense sequence; it has the same sense sequence as the mRNA transcript (though T bases in DNA are substituted with U bases in RNA).
3′CGCTATAGCGTTT 5′ |
DNA antisense strand (template/noncoding) | Used as a template for transcription. |
5′GCGATATCGCAAA 3′ |
DNA sense strand (nontemplate/coding) | Complementary to the template strand. |
5′GCGAUAUCGCAAA 3′ |
mRNA sense transcript | RNA strand that is transcribed from the noncoding (template/antisense) strand. Note: Except for the fact that all thymines are now uracils (T → U), it is complementary to the noncoding (template/antisense) DNA strand and identical to the coding (nontemplate/sense) DNA strand.
|
3′CGCUAUAGCGUUU 5′ |
mRNA antisense transcript | RNA strand that is transcribed from the coding (nontemplate/sense) strand. Note: Except for the fact that all thymines are now uracils (T → U), it is complementary to the coding (nontemplate/sense) DNA strand and identical to the noncoding (template/antisense) DNA strand.
|
The names assigned to each strand actually depend on which direction you are writing the sequence that contains the information for proteins (the "sense" information), not on which strand is depicted as "on the top" or "on the bottom" (which is arbitrary). The only biological information that is important for labeling strands is the relative locations of the terminal 5′ phosphate group and the terminal 3′ hydroxyl group (at the ends of the strand or sequence in question), because these ends determine the direction of transcription and translation. A sequence written 5′-CGCTAT-3′ is equivalent to a sequence written 3′-TATCGC-5′ as long as the 5′ and 3′ ends are noted. If the ends are not labeled, convention is to assume that both sequences are written in the 5′-to-3′ direction. The "Watson strand" refers to 5′-to-3′ top strand (5′→3′), whereas the "Crick strand" refers to the 5′-to-3′ bottom strand (3′←5′). Both Watson and Crick strands can be either sense or antisense strands depending on the specific gene product made from them.
For example, the notation "YEL021W", an alias of the URA3 gene used in the National Center for Biotechnology Information (NCBI) database, denotes that this gene is in the 21st open reading frame (ORF) from the centromere of the left arm (L) of Yeast (Y) chromosome number V (E), and that the expression coding strand is the Watson strand (W). "YKL074C" denotes the 74th ORF to the left of the centromere of chromosome XI and that the coding strand is the Crick strand (C). Another confusing term referring to "Plus" and "Minus" strand is also widely used. Whether the strand is sense (positive) or antisense (negative), the default query sequence in NCBI BLAST alignment is "Plus" strand.
Ambisense
A single-stranded genome that is used in both positive-sense and negative-sense capacities is said to be ambisense. Some viruses have ambisense genomes. Bunyaviruses have three single-stranded RNA (ssRNA) fragments, some of them containing both positive-sense and negative-sense sections; arenaviruses are also ssRNA viruses with an ambisense genome, as they have three fragments that are mainly negative-sense except for part of the 5′ ends of the large and small segments of their genome.
Antisense RNA
Main article: Antisense RNAAn RNA sequence that is complementary to an endogenous mRNA transcript is sometimes called "antisense RNA". In other words, it is a non-coding strand complementary to the coding sequence of RNA; this is similar to negative-sense viral RNA. When mRNA forms a duplex with a complementary antisense RNA sequence, translation is blocked. This process is related to RNA interference. Cells can produce antisense RNA molecules naturally, called microRNAs, which interact with complementary mRNA molecules and inhibit their expression. The concept has also been exploited as a molecular biology technique, by artificially introducing a transgene coding for antisense RNA in order to block the expression of a gene of interest. Radioactively or fluorescently labelled antisense RNA can be used to show the level of transcription of genes in various cell types.
Some alternative antisense structural types have been experimentally applied as antisense therapy. In the United States, the Food and Drug Administration (FDA) has approved the phosphorothioate antisense oligonucleotides fomivirsen (Vitravene) and mipomersen (Kynamro) for human therapeutic use.
RNA sense in viruses
In virology, the term "sense" has a slightly different meaning. The genome of an RNA virus can be said to be either positive-sense, also known as a "plus-strand", or negative-sense, also known as a "minus-strand". In most cases, the terms "sense" and "strand" are used interchangeably, making terms such as "positive-strand" equivalent to "positive-sense", and "plus-strand" equivalent to "plus-sense". Whether a viral genome is positive-sense or negative-sense can be used as a basis for classifying viruses.
Positive-sense
Main article: Positive-sense single-stranded RNA virusPositive-sense (5′-to-3′) viral RNA signifies that a particular viral RNA sequence may be directly translated into viral proteins (e.g., those needed for viral replication). Therefore, in positive-sense RNA viruses, the viral RNA genome can be considered viral mRNA, and can be immediately translated by the host cell. Unlike negative-sense RNA, positive-sense RNA is of the same sense as mRNA. Some viruses (e.g. Coronaviridae) have positive-sense genomes that can act as mRNA and be used directly to synthesize proteins without the help of a complementary RNA intermediate. Because of this, these viruses do not need to have an RNA polymerase packaged into the virion—the RNA polymerase will be one of the first proteins produced by the host cell, since it is needed in order for the virus's genome to be replicated.
Negative-sense
Main article: Negative-sense single-stranded RNA virusNegative-sense (3′-to-5′) viral RNA is complementary to the viral mRNA, thus a positive-sense RNA must be produced by an RNA-dependent RNA polymerase from it prior to translation. Like DNA, negative-sense RNA has a nucleotide sequence complementary to the mRNA that it encodes; also like DNA, this RNA cannot be translated into protein directly. Instead, it must first be transcribed into a positive-sense RNA that acts as an mRNA. Some viruses (e.g. influenza viruses) have negative-sense genomes and so must carry an RNA polymerase inside the virion.
Antisense oligonucleotides
Gene silencing can be achieved by introducing into cells a short "antisense oligonucleotide" that is complementary to an RNA target. This experiment was first done by Zamecnik and Stephenson in 1978 and continues to be a useful approach, both for laboratory experiments and potentially for clinical applications (antisense therapy). Several viruses, such as influenza viruses Respiratory syncytial virus (RSV) and SARS coronavirus (SARS-CoV), have been targeted using antisense oligonucleotides to inhibit their replication in host cells.
If the antisense oligonucleotide contains a stretch of DNA or a DNA mimic (phosphorothioate DNA, 2′F-ANA, or others) it can recruit RNase H to degrade the target RNA. This makes the mechanism of gene silencing catalytic. Double-stranded RNA can also act as a catalytic, enzyme-dependent antisense agent through the RNAi/siRNA pathway, involving target mRNA recognition through sense-antisense strand pairing followed by target mRNA degradation by the RNA-induced silencing complex (RISC). The R1 plasmid hok/sok system provides yet another example of an enzyme-dependent antisense regulation process through enzymatic degradation of the resulting RNA duplex.
Other antisense mechanisms are not enzyme-dependent, but involve steric blocking of their target RNA (e.g. to prevent translation or to induce alternative splicing). Steric blocking antisense mechanisms often use oligonucleotides that are heavily modified. Since there is no need for RNase H recognition, this can include chemistries such as 2′-O-alkyl, peptide nucleic acid (PNA), locked nucleic acid (LNA), and Morpholino oligomers.
See also
- Antisense therapy
- Directionality (molecular biology)
- DNA codon table
- RNA virus
- Transcription (genetics)
- Translation (genetics)
- Viral replication
References
- Anne-Lise Haenni (2003). "Expression strategies of ambisense viruses". Virus Research. 93 (2): 141–150. doi:10.1016/S0168-1702(03)00094-7. PMID 12782362.
- Kakutani T; Hayano Y; Hayashi T; Minobe Y. (1991). "Ambisense segment 3 of rice stripe virus: the first instance of a virus containing two ambisense segments". J Gen Virol. 72 (2): 465–8. doi:10.1099/0022-1317-72-2-465. PMID 1993885.
- Zhu Y; Hayakawa T; Toriyama S; Takahashi M. (1991). "Complete nucleotide sequence of RNA 3 of rice stripe virus: an ambisense coding strategy". J Gen Virol. 72 (4): 763–7. doi:10.1099/0022-1317-72-4-763. PMID 2016591.
- Cartwright, Reed; Dan Graur (Feb 8, 2011). "The multiple personalities of Watson and Crick strands". Biology Direct. 6: 7. doi:10.1186/1745-6150-6-7. PMC 3055211. PMID 21303550.
- "FDA approves fomivirsen for CMV". healio. 1 October 1998. Retrieved 18 September 2020.
- "FDA approves orphan drug for inherited cholesterol disorder". Drug Topics. 30 January 2013. Retrieved 18 September 2020.
- Zamecnik, P.C.; Stephenson, M.L. (1978). "Inhibition of Rous sarcoma Virus Replication and Cell Transformation by a Specific Oligodeoxynucleotide". Proc. Natl. Acad. Sci. USA. 75 (1): 280–284. Bibcode:1978PNAS...75..280Z. doi:10.1073/pnas.75.1.280. PMC 411230. PMID 75545.
- Watts, J.K.; Corey, D.R. (2012). "Silencing Disease Genes in the Laboratory and in the Clinic". J. Pathol. 226 (2): 365–379. doi:10.1002/path.2993. PMC 3916955. PMID 22069063.
- ^ Kumar, Binod; Khanna, Madhu; Meseko, Clement A.; Sanicas, Melvin; Kumar, Prashant; Asha, Kumari; Asha, Kumari; Kumar, Prashant; Sanicas, Melvin (January 2019). "Advancements in Nucleic Acid Based Therapeutics against Respiratory Viral Infections". Journal of Clinical Medicine. 8 (1): 6. doi:10.3390/jcm8010006. PMC 6351902. PMID 30577479.
- Kumar, Binod; Asha, Kumari; Khanna, Madhu; Ronsard, Larance; Meseko, Clement Adebajo; Sanicas, Melvin (2018-01-10). "The emerging influenza virus threat: status and new prospects for its therapy and control". Archives of Virology. 163 (4): 831–844. doi:10.1007/s00705-018-3708-y. ISSN 0304-8608. PMC 7087104. PMID 29322273.
- Kumar, Prashant; Kumar, Binod; Rajput, Roopali; Saxena, Latika; Banerjea, Akhil C.; Khanna, Madhu (2013-06-02). "Cross-Protective Effect of Antisense Oligonucleotide Developed Against the Common 3′ NCR of Influenza A Virus Genome". Molecular Biotechnology. 55 (3): 203–211. doi:10.1007/s12033-013-9670-8. ISSN 1073-6085. PMID 23729285. S2CID 24496875.
- Kumar, B.; Khanna, Madhu; Kumar, P.; Sood, V.; Vyas, R.; Banerjea, A. C. (2011-07-09). "Nucleic Acid-Mediated Cleavage of M1 Gene of Influenza A Virus Is Significantly Augmented by Antisense Molecules Targeted to Hybridize Close to the Cleavage Site". Molecular Biotechnology. 51 (1): 27–36. doi:10.1007/s12033-011-9437-z. ISSN 1073-6085. PMID 21744034. S2CID 45686564.