| This article may be too technical for most readers to understand. Please help improve it to make it understandable to non-experts, without removing the technical details. (October 2021) (Learn how and when to remove this message) |

In genetics, a polygenic score (PGS) is a number that summarizes the estimated effect of many genetic variants on an individual's phenotype. The PGS is also called the polygenic index (PGI) or genome-wide score; in the context of disease risk, it is called a polygenic risk score (PRS or PR score) or genetic risk score. The score reflects an individual's estimated genetic predisposition for a given trait and can be used as a predictor for that trait. It gives an estimate of how likely an individual is to have a given trait based only on genetics, without taking environmental factors into account; and it is typically calculated as a weighted sum of trait-associated alleles.
Recent progress in genetics has developed polygenic predictors of complex human traits, including risk for many important complex diseases that are typically affected by many genetic variants, each of which confers a small effect on overall risk. In a polygenic risk predictor the lifetime (or age-range) risk for the disease is a numerical function captured by the score which depends on the states of thousands of individual genetic variants (i.e., single-nucleotide polymorphisms, or SNPs).
Polygenic scores are widely used in animal breeding and plant breeding due to their efficacy in improving livestock breeding and crops. In humans, polygenic scores are typically generated from data of genome-wide association study (GWAS). They are an active area of research spanning topics such as learning algorithms for genomic prediction; new predictor training; validation testing of predictors; and clinical application of PRS. In 2018, the American Heart Association named polygenic risk scores as one of the major breakthroughs in research in heart disease and stroke.
Background

DNA in living organisms is the molecular genetic code for life. Although polygenic risk scores from study in humans have gained the most attention, the basic idea was first introduced for selective plant and animal breeding. Similar to the latter-day approaches of constructing a polygenic risk score, an individual's—animal or plant—breeding value was calculated to be the combined weight of several single-nucleotide polymorphisms (SNPs) by their individual effects on a trait.
Human DNA contains about 3 billion bases. The human genome can be broadly separated into coding and non-coding sequences, where the coding genome encodes instructions for genes, including some of the sequence that codes for proteins. Genome-wide association studies enable mapping phenotypes to the variations in nucleotide bases in human populations. Improvements in methodology and studies with large cohorts have enabled the mapping of many traits—some of which are diseases—to the human genome. Learning which variations influence which specific traits and how strongly they do so, are the key targets for constructing polygenic scores in humans.
The methods were first considered for humans after the year 2000, and specifically by a proposal in 2007 that such scores could be used in human genetics to identify individuals at high risk for disease. The concept was successfully applied in 2009 by researchers who organized a genome-wide association study (GWAS) regarding schizophrenia with the objective of constructing scores of risk propensity. That study was the first to use the term polygenic score for a prediction drawn from a linear combination of single-nucleotide polymorphism (SNP) genotypes—which was able to explain 3% of the variance in schizophrenia.
Calculation with genome-wide association study
A PRS is constructed from the estimated effect sizes derived from a genome-wide association study (GWAS). In a GWAS, single-nucleotide polymorphisms (SNPs) are tested for an association between cases and controls, (see top graphic). The results from a GWAS estimate the strength of the association at each SNP, i.e., the effect size at the SNP, as well as a p-value for statistical significance. A typical score is then calculated by adding the number of risk-modifying alleles across a large number of SNPs, where the number of alleles for each SNP is multiplied by the weight for the SNP.
In mathematical form, the estimated polygenic score is obtained as the sum across m number of SNPs with risk-increasing alleles weighted by their weights, i.e., .
This idea can be generalized to the study of any trait, and is an example of the more general mathematical term regression analysis.
Key considerations
Methods for generating polygenic scores in humans are an active area of research. Two key considerations in developing polygenic scores are which SNPs and the number of SNPs to include. The simplest, the so-called "pruning and thresholding" method, sets weights equal to the coefficient estimates from a regression of the trait on each genetic variant. The included SNPs may be selected using an algorithm that attempts to ensure that each marker is approximately independent.
Independence of each SNP is important for the score's predictive accuracy. SNPs that are physically close to each other are more likely to be in linkage disequilibrium, meaning they typically are inherited together and therefore don't provide independent predictive power. That's what's referred to as 'pruning'. The 'thresholding' refers to using only SNPs that meet a specific p-value threshold.
Penalized regression can also be used to construct polygenic scores. From prior information penalized regression assigns probabilities on: 1) how many genetic variants are expected to affect a trait, and 2) the distribution of their effect sizes. These methods in effect "penalize" the large coefficients in a regression model and shrink them conservatively. One popular tool for this approach is "PRS-CS". Another is to use certain Bayesian methods, first proposed in 2001 that directly incorporate genetic features of a given trait as well as genomic features like linkage disequilibrium. (One Bayesian method uses "linkage disequilibrium prediction" or LDpred.)
More approaches for developing polygenic risk scores continue to be described. For example, by incorporating effect sizes from populations of different ancestry, the predictive ability of scores can be improved. Incorporating knowledge of the functional roles of specific genomic chunks can improve the utility of scores. Studies have examined the performances of these methods on standardized dataset.
Application to humans
As the number of genome-wide association studies has exploded, along with rapid advances in methods for calculating polygenic scores, its most obvious application is in clinical settings for disease prediction or risk stratification. It is important not to over- or under-state the value of polygenic scores. A key advantage of quantifying polygenic contribution for each individual is that the genetic liability does not change over an individual's lifespan. However, while a disease may have strong genetic contributions, the risk arising from one's genetics has to be interpreted in the context of environmental factors. For example, even if an individual has a high genetic risk for alcoholism, that risk is lessened if that individual is never exposed to alcohol.
Predictive performance in humans
For humans, while most polygenic scores are not predictive enough to diagnose disease, they could be used in addition to other covariates (such as age, BMI, smoking status) to improve estimates of disease susceptibility. However, even if a polygenic score might not make reliable diagnostic predictions across an entire population, it may still make very accurate predictions for outliers at extreme high or low risk. The clinical utility may therefore still be large even if average measures of prediction performance are moderate.
Although issues such as poorer predictive performance in individuals of non-European ancestry limit widespread use, several authors have noted that some causal variants for some conditions, but not others, are shared between Europeans and other groups across different continents for (e.g.) BMI and type 2 diabetes in African populations as well as schizophrenia in Chinese populations. Other researchers recognize that polygenic under-prediction in non-European population should galvanize new GWAS that prioritize greater genetic diversity in order to maximize the potential health benefits brought about by predictive polygenic scores. Significant scientific efforts are being made to this end.
Embryo genetic screening is common with millions biopsied and tested each year worldwide. Genotyping methods have been developed so that the embryo genotype can be determined to high precision. Testing for aneuploidy and monogenetic diseases has increasingly become established over decades, whereas tests for polygenic diseases have begun to be employed more recently, having been first used in embryo selection in 2019. The use of polygenic scores for embryo selection has been criticised due to alleged ethical and safety issues as well as limited practical utility. However, trait-specific evaluations claiming the contrary have been put forth and ethical arguments for PGS-based embryo selection have also been made. The topic continues to be an active area of research not only within genomics but also within clinical applications and ethics.
As of 2019, polygenic scores from well over a hundred phenotypes have been developed from genome-wide association statistics. These include scores that can be categorized as anthropometric, behavioural, cardiovascular, non-cancer illness, psychiatric/neurological, and response to treatment/medication.
Examples of disease prediction performance
When predicting disease risk, a PGS gives a continuous score that estimates the risk of having or getting the disease, within some pre-defined time span. A common metric for evaluating such continuous estimates of yes/no questions (see Binary classification) is the area under the ROC curve (AUC). Some example results of PGS performance, as measured in AUC (0 ≤ AUC ≤ 1 where a larger number implies better prediction), include:
- In 2018, AUC ≈ 0.64 for coronary disease using ~120,000 British individuals.
- In 2019, AUC ≈ 0.63 for breast cancer, developed from ~95,000 case subjects and ~75,000 controls of European ancestry.
- In 2019, AUC ≈ 0.71 for hypothyroidism for ~24,000 case subjects and ~463,00 controls of European ancestry.
- In 2020, AUC ≈ 0.71 for schizophrenia, using 90 cohorts including ~67,000 case subjects and ~94,000 controls with ~80% of European ancestry and ~20% of East Asian ancestry. Note that these results use purely genetic information as input; including additional information such as age and sex often greatly improves the predictions. The coronary disease predictor and the hypothyroidism predictor above achieve AUCs of ~ 0.80 and ~0.78, respectively, when also including age and sex.
Importance of sample size

The performance of a polygenic predictor is highly dependent on the size of the dataset that is available for analysis and ML training. Recent scientific progress in prediction power relies heavily on the creation and expansion of large biobanks containing data for both genotypes and phenotypes of very many individuals. As of 2021, there exist several biobanks with hundreds of thousands samples, i.e., data entries with both genetic and trait information for each individual (see for instance the incomplete list of biobanks).
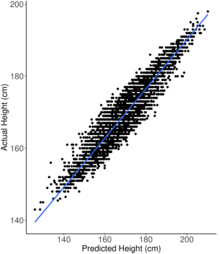
With the use of these growing biobanks, data from many thousands of individuals are used to detect the relevant variants for a specific trait. Exactly how many are required depends very much on the trait in question. Typically, increasing levels of prediction are observed until a plateau phase where the performance levels off and does not change much when increasing the sample size even further. This is the limit of how accurate a polygenic predictor that only uses genetic information can be and is set by the heritability of the specific trait. The sample size required to reach this performance level for a certain trait is determined by the complexity of the underlying genetic architecture and the distribution of genetic variance in the sampled population. This sample size dependence is illustrated in the figure for hypothyroidism, hypertension and type 2 diabetes.
Note again, that current methods to construct polygenic predictors are sensitive to the ancestries present in the data. As of 2021, most available data have been primarily of populations with European ancestry, which is the reason why PGS generally perform better within this ancestry. The construction of more diverse biobanks with successful recruitment from all ancestries is required to rectify this skewed access to and benefits from PGS-based medicine.
Clinical utility and current usage
A landmark study examining the role of polygenic risk scores in cardiovascular disease invigorated interest the clinical potential of polygenic scores. This study demonstrated that an individual with the highest polygenic risk score (top 1%) had a lifetime cardiovascular risk >10% which was comparable to those with rare genetic variants. This comparison is important because clinical practice can be influenced by knowing which individuals have this rare genetic cause of cardiovascular disease. Since this study, polygenic risk scores have shown promise for disease prediction across other traits. Polygenic risk scores have been studied heavily in obesity, coronary artery disease, diabetes, breast cancer, prostate cancer, Alzheimer's disease and psychiatric diseases.
As of January 2021 providing PRS directly to individuals was undergoing research trials in health systems around the world, but is not yet offered as standard of care. Most use is therefore through consumer genetic testing, where a number of private companies report PRS for a number of diseases and traits. Consumers download their genotype (genetic variant) data and upload them into online PRS calculators, e.g. Scripps Health, Impute.me or Color Genomics. The most frequently reported motivation for individuals to seek out PRS reports is general curiosity (98.2%), and the reactions are generally mixed with common misinterpretations. It is speculated that personal use of PRS could contribute to treatment choices, but that more data is needed. As of 2020 a more typical use was that clinicians face individuals with commercially derived disease-specific PRS in the expectation that the clinician will interpret them, something that may create extra burdens for the clinical care system.
Challenges and risks in clinical contexts
At a fundamental level, the use of polygenic scores in clinical context will have similar technical issues as existing tools. For example, if a tool is not validated in a diverse population, then it may exacerbate disparities with unequal efficacy across populations. This is especially important in genetics where, as of 2018, a majority of the studies to date have been done in Europeans. Other challenges that can arise include how precisely the polygenic risk score can be calculated and how precise it needs to be for clinical utility. Even if a polygenic score is accurately calculated and calibrated for a population, its interpretation must be approached with caution. First, it is important to realize that polygenic traits are different from monogenic traits; the latter stem from fewer genetic loci and can be detected more accurately. Genetic tests are often difficult to interpret and require genetic counseling. Currently, polygenic-score results are being shared with clinicians. Since monogenic genetic testing is far more mature than polygenic scores, we can look there for approximating the clinical impact of polygenic scores. While some studies have found negative effects of returning monogenic genetic results to patients, the majority of studies have that negative consequences are minor.
Benefits in humans
Unlike many other clinical laboratory or imaging methods, an individual's germ-line genetic risk can be calculated at birth for a variety of diseases after sequencing their DNA once. Thus, polygenic scores may ultimately be a cost-effective measure that can be informative for clinical management. Moreover, the polygenic risk score may be informative across an individual's lifespan helping to quantify the genetic lifelong risk for certain diseases. For many diseases, having a strong genetic risk can results in an earlier onset of presentation (e.g. Familial Hypercholesterolemia). Recognizing an increased genetic burden earlier can allow clinicians to intervene earlier and avoid delayed diagnoses. Polygenic score can be combined with traditional risk factors to increase clinical utility. For example, polygenic risk scores may help improve diagnosis of diseases. This is especially evident in distinguishing Type 1 from Type 2 Diabetes. Likewise, a polygenic risk score based approach may reduce invasive diagnostic procedures as demonstrated in Celiac disease. Polygenic scores may also empower individuals to alter their lifestyles to reduce risk for diseases. While there is some evidence for behavior modification as a result of knowing one's genetic predisposition, more work is required to evaluate risk-modifying behaviors across a variety of different disease states. Population level screening is another use case for polygenic scores. The goal of population-level screening is to identify patients at high risk for a disease who would benefit from an existing treatment. Polygenic scores can identify a subset of the population at high risk that could benefit from screening. Several clinical studies are being done in breast cancer and heart disease is another area that could benefit from a polygenic score based screening program.
Non-predictive applications
A variety of applications exists for polygenic scores. In humans, polygenic scores were originally computed in an effort to predict the prevalence and etiology of complex, heritable diseases, which are typically affected by many genetic variants that individually confer a small effect to overall risk. Additionally, a polygenic score can be used in several different ways: as a lower bound to test whether heritability estimates may be biased; as a measure of genetic overlap of traits (genetic correlation), which might indicate e.g. shared genetic bases for groups of mental disorders; as a means to assess group differences in a trait such as height, or to examine changes in a trait over time due to natural selection indicative of a soft selective sweep (as e.g. for intelligence where the changes in frequency would be too small to detect on each individual hit but not on the overall polygenic score); in Mendelian randomization (assuming no pleiotropy with relevant traits); to detect & control for the presence of genetic confounds in outcomes (e.g. the correlation of schizophrenia with poverty); or to investigate gene–environment interactions and correlations. Polygenic scores also have useful statistical properties in (genomic) association testing, for instance to account for outcome-specific background effects and/or improve statistical power.
Applications in non-human species
The benefit of polygenic scores is that they can be used to predict the future for crops, animal breeding, and humans alike. Although the same basic concepts underlie these areas of prediction, they face different challenges that require different methodologies. The ability to produce very large family size in nonhuman species, accompanied by deliberate selection, leads to a smaller effective population, higher degrees of linkage disequilibrium among individuals, and a higher average genetic relatedness among individuals within a population. For example, members of plant and animal breeds that humans have effectively created, such as modern maize or domestic cattle, are all technically "related". In human genomic prediction, by contrast, unrelated individuals in large populations are selected to estimate the effects of common SNPs. Because of smaller effective population in livestock, the mean coefficient of relationship between any two individuals is likely high, and common SNPs will tag causal variants at greater physical distance than for humans; this is the major reason for lower SNP-based heritability estimates for humans compared to livestock. In both cases, however, sample size is key for maximizing the accuracy of genomic prediction.
While modern genomic prediction scoring in humans is generally referred to as a "polygenic score" (PGS) or a "polygenic risk score" (PRS), in livestock the more common term is "genomic estimated breeding value", or GEBV (similar to the more familiar "EBV", but with genotypic data). Conceptually, a GEBV is the same as a PGS: a linear function of genetic variants that are each weighted by the apparent effect of the variant. Despite this, polygenic prediction in livestock is useful for a fundamentally different reason than for humans. In humans, a PRS is used for the prediction of individual phenotype, while in livestock a GEBV is typically used to predict the offspring's average value of a phenotype of interest in terms of the genetic material it inherited from a parent. In this way, a GEBV can be understood as the average of the offspring of an individual or pair of individual animals. GEBVs are also typically communicated in the units of the trait of interest. For example, the expected increase in milk production of the offspring of a specific parent compared to the offspring from a reference population might be a typical way of using a GEBV in dairy cow breeding and selection.
Notes
- A. Preprint lists AUC for pure PRS while the published version of the paper only lists AUC for PGS combined with age, sex and genotyping array information.
References
- Gregory, Gillian; Das Gupta, Kuheli; Meiser, Bettina; Barlow-Stewart, Kristine; Geelan-Small, Peter; Kaur, Rajneesh; Scheepers-Joynt, Maatje; McInerny, Simone; Taylor, Shelby; Antill, Yoland; Salmon, Lucinda; Smyth, Courtney; Young, Mary-Anne; James, Paul A; Yanes, Tatiane (February 2022). "Polygenic risk in familial breast cancer: Changing the dynamics of communicating genetic risk". Journal of Genetic Counseling. 31 (1): 120–129. doi:10.1002/jgc4.1458. hdl:1959.4/unsworks_78566. PMID 34223688. S2CID 235732957.
- ^ de Vlaming R, Groenen PJ (2015). "The Current and Future Use of Ridge Regression for Prediction in Quantitative Genetics". BioMed Research International. 2015: 143712. doi:10.1155/2015/143712. PMC 4529984. PMID 26273586.
- Lewis CM, Vassos E (November 2017). "Prospects for using risk scores in polygenic medicine". Genome Medicine. 9 (1): 96. doi:10.1186/s13073-017-0489-y. PMC 5683372. PMID 29132412.
- ^ Khera AV, Chaffin M, Aragam KG, Haas ME, Roselli C, Choi SH, et al. (September 2018). "Genome-wide polygenic scores for common diseases identify individuals with risk equivalent to monogenic mutations". Nature Genetics. 50 (9): 1219–1224. doi:10.1038/s41588-018-0183-z. PMC 6128408. PMID 30104762.
- Yanes T, Meiser B, Kaur R, Scheepers-Joynt M, McInerny S, Taylor S, et al. (March 2020). "Uptake of polygenic risk information among women at increased risk of breast cancer". Clinical Genetics. 97 (3): 492–501. doi:10.1111/cge.13687. hdl:11343/286783. PMID 31833054. S2CID 209342044.
- ^ Vilhjálmsson BJ, Yang J, Finucane HK, Gusev A, Lindström S, Ripke S, et al. (October 2015). "Modeling Linkage Disequilibrium Increases Accuracy of Polygenic Risk Scores". American Journal of Human Genetics. 97 (4): 576–592. doi:10.1016/j.ajhg.2015.09.001. PMC 4596916. PMID 26430803.
- Dudbridge F (March 2013). "Power and predictive accuracy of polygenic risk scores". PLOS Genetics. 9 (3): e1003348. doi:10.1371/journal.pgen.1003348. PMC 3605113. PMID 23555274.
- ^ Torkamani A, Wineinger NE, Topol EJ (September 2018). "The personal and clinical utility of polygenic risk scores". Nature Reviews. Genetics. 19 (9): 581–590. doi:10.1038/s41576-018-0018-x. PMID 29789686. S2CID 46893131.
- ^ Lambert SA, Abraham G, Inouye M (November 2019). "Towards clinical utility of polygenic risk scores". Human Molecular Genetics. 28 (R2): R133–R142. doi:10.1093/hmg/ddz187. PMID 31363735.
- Regalado A (8 March 2019). "23andMe thinks polygenic risk scores are ready for the masses, but experts aren't so sure". MIT Technology Review. Retrieved 2020-08-14.
- ^ Lello L, Raben TG, Yong SY, Tellier LC, Hsu SD (October 2019). "Genomic Prediction of 16 Complex Disease Risks Including Heart Attack, Diabetes, Breast and Prostate Cancer". Scientific Reports. 9 (1): 15286. Bibcode:2019NatSR...915286L. doi:10.1038/s41598-019-51258-x. PMC 6814833. PMID 31653892.
- Visscher PM, Wray NR, Zhang Q, Sklar P, McCarthy MI, Brown MA, Yang J (July 2017). "10 Years of GWAS Discovery: Biology, Function, and Translation". American Journal of Human Genetics. 101 (1): 5–22. doi:10.1016/j.ajhg.2017.06.005. PMC 5501872. PMID 28686856.
- ^ Spiliopoulou A, Nagy R, Bermingham ML, Huffman JE, Hayward C, Vitart V, et al. (July 2015). "Genomic prediction of complex human traits: relatedness, trait architecture and predictive meta-models". Human Molecular Genetics. 24 (14): 4167–4182. doi:10.1093/hmg/ddv145. PMC 4476450. PMID 25918167.
- Spindel JE, McCouch SR (December 2016). "When more is better: how data sharing would accelerate genomic selection of crop plants". The New Phytologist. 212 (4): 814–826. doi:10.1111/nph.14174. PMID 27716975.
- "Modern genetics will improve health and usher in "designer" children". The Economist. 2019-11-09. Retrieved 2021-04-12.
- "Test could predict risk of future heart disease for just £40". The Guardian. 2018-10-08. Retrieved 2021-04-12.
- ^ Raben TG, Lello L, Widen E, Hsu SD (2021-01-14). "From Genotype to Phenotype: polygenic prediction of complex human traits". arXiv:2101.05870 .
- "Big picture genetic scoring approach reliably predicts heart disease". Science Daily. 2019-06-11. Retrieved 2021-04-12.
- Weedon MN, McCarthy MI, Hitman G, Walker M, Groves CJ, Zeggini E, et al. (October 2006). "Combining information from common type 2 diabetes risk polymorphisms improves disease prediction". PLOS Medicine. 3 (10): e374. doi:10.1371/journal.pmed.0030374. PMC 1584415. PMID 17020404.
- Meuwissen TH, Hayes BJ, Goddard ME (April 2001). "Prediction of total genetic value using genome-wide dense marker maps". Genetics. 157 (4): 1819–1829. doi:10.1093/genetics/157.4.1819. PMC 1461589. PMID 11290733.
- Crouch DJ, Bodmer WF (August 2020). "Polygenic inheritance, GWAS, polygenic risk scores, and the search for functional variants". Proceedings of the National Academy of Sciences of the United States of America. 117 (32): 18924–18933. Bibcode:2020PNAS..11718924C. doi:10.1073/pnas.2005634117. PMC 7431089. PMID 32753378.
- Wray NR, Goddard ME, Visscher PM (October 2007). "Prediction of individual genetic risk to disease from genome-wide association studies". Genome Research. 17 (10): 1520–1528. doi:10.1101/gr.6665407. PMC 1987352. PMID 17785532.
- Purcell SM, Wray NR, Stone JL, Visscher PM, O'Donovan MC, Sullivan PF, Sklar P (August 2009). "Common polygenic variation contributes to risk of schizophrenia and bipolar disorder". Nature. 460 (7256): 748–752. Bibcode:2009Natur.460..748P. doi:10.1038/nature08185. PMC 3912837. PMID 19571811.
- Choi SW, Mak TS, O'Reilly PF (September 2020). "Tutorial: a guide to performing polygenic risk score analyses". Nature Protocols. 15 (9): 2759–2772. doi:10.1038/s41596-020-0353-1. PMC 7612115. PMID 32709988. S2CID 220732490.
- Wand H, Lambert SA, Tamburro C, Iacocca MA, O'Sullivan JW, Sillari C, et al. (March 2021). "Improving reporting standards for polygenic scores in risk prediction studies". Nature. 591 (7849): 211–219. Bibcode:2021Natur.591..211W. doi:10.1038/s41586-021-03243-6. PMC 8609771. PMID 33692554.
- ^ Lewis CM, Vassos E (May 2020). "Polygenic risk scores: from research tools to clinical instruments". Genome Medicine. 12 (1): 44. doi:10.1186/s13073-020-00742-5. PMC 7236300. PMID 32423490.
- Vattikuti S, Lee JJ, Chang CC, Hsu SD, Chow CC (2014). "Applying compressed sensing to genome-wide association studies". GigaScience. 3 (1): 10. doi:10.1186/2047-217X-3-10. PMC 4078394. PMID 25002967.
- Ge T, Chen CY, Ni Y, Feng YA, Smoller JW (April 2019). "Polygenic prediction via Bayesian regression and continuous shrinkage priors". Nature Communications. 10 (1): 1776. Bibcode:2019NatCo..10.1776G. doi:10.1038/s41467-019-09718-5. PMC 6467998. PMID 30992449.
- Meuwissen TH, Hayes BJ, Goddard ME (April 2001). "Prediction of total genetic value using genome-wide dense marker maps". Genetics. 157 (4): 1819–1829. doi:10.1093/genetics/157.4.1819. PMC 1461589. PMID 11290733.
- Privé F, Arbel J, Vilhjálmsson BJ (December 2020). "LDpred2: better, faster, stronger". Bioinformatics. 36 (22–23): 5424–5431. doi:10.1093/bioinformatics/btaa1029. PMC 8016455. PMID 33326037.
- Márquez-Luna C, Loh PR, Price AL (December 2017). "Multiethnic polygenic risk scores improve risk prediction in diverse populations". Genetic Epidemiology. 41 (8): 811–823. doi:10.1002/gepi.22083. PMC 5726434. PMID 29110330.
- Hu Y, Lu Q, Powles R, Yao X, Yang C, Fang F, et al. (June 2017). Rigoutsos I (ed.). "Leveraging functional annotations in genetic risk prediction for human complex diseases". PLOS Computational Biology. 13 (6): e1005589. Bibcode:2017PLSCB..13E5589H. doi:10.1371/journal.pcbi.1005589. PMC 5481142. PMID 28594818.
- Ni G, Zeng J, Revez JA, Wang Y, Zheng Z, Ge T, et al. (November 2021). "A Comparison of Ten Polygenic Score Methods for Psychiatric Disorders Applied Across Multiple Cohorts". Biological Psychiatry. 90 (9): 611–620. doi:10.1016/j.biopsych.2021.04.018. PMC 8500913. PMID 34304866.
- Duncan L, Shen H, Gelaye B, Meijsen J, Ressler K, Feldman M, et al. (July 2019). "Analysis of polygenic risk score usage and performance in diverse human populations". Nature Communications. 10 (1): 3328. Bibcode:2019NatCo..10.3328D. doi:10.1038/s41467-019-11112-0. PMC 6658471. PMID 31346163.
- Wang Y, Guo J, Ni G, Yang J, Visscher PM, Yengo L (July 2020). "Theoretical and empirical quantification of the accuracy of polygenic scores in ancestry divergent populations". Nature Communications. 11 (1): 3865. Bibcode:2020NatCo..11.3865W. doi:10.1038/s41467-020-17719-y. PMC 7395791. PMID 32737319.
- Li Z, Chen J, Yu H, He L, Xu Y, Zhang D, et al. (November 2017). "Genome-wide association analysis identifies 30 new susceptibility loci for schizophrenia" (PDF). Nature Genetics. 49 (11): 1576–1583. doi:10.1038/ng.3973. PMID 28991256. S2CID 205355668.
- ^ Martin AR, Kanai M, Kamatani Y, Okada Y, Neale BM, Daly MJ (April 2019). "Clinical use of current polygenic risk scores may exacerbate health disparities". Nature Genetics. 51 (4): 584–591. doi:10.1038/s41588-019-0379-x. PMC 6563838. PMID 30926966.
- "Diversity and Inclusion". All of Us Research Program | NIH. 2020-06-24. Retrieved 2023-05-24.
- Zeevi DA, Backenroth D, Hakam-Spector E, Renbaum P, Mann T, Zahdeh F, et al. (July 2021). "Expanded clinical validation of Haploseek for comprehensive preimplantation genetic testing". Genetics in Medicine. 23 (7): 1334–1340. doi:10.1038/s41436-021-01145-6. PMID 33772222. S2CID 232377300.
- Treff NR, Zimmerman R, Bechor E, Hsu J, Rana B, Jensen J, et al. (August 2019). "Validation of concurrent preimplantation genetic testing for polygenic and monogenic disorders, structural rearrangements, and whole and segmental chromosome aneuploidy with a single universal platform". European Journal of Medical Genetics. 62 (8): 103647. doi:10.1016/j.ejmg.2019.04.004. PMID 31026593.
- Carey Goldberg (2021-09-17). "Picking Embryos With Best Health Odds Sparks New DNA Debate". Bloomberg News. Archived from the original on 2021-09-17.
Aurea appears to be the first child born after a new type of DNA testing that gave her a "polygenic risk score." ... Her parents underwent fertility treatment in 2019 ...
- Birney E (11 November 2019). "Why using genetic risk scores on embryos is wrong". ewanbirney.com. Retrieved 2020-12-16.
- Karavani E, Zuk O, Zeevi D, Barzilai N, Stefanis NC, Hatzimanolis A, et al. (November 2019). "Screening Human Embryos for Polygenic Traits Has Limited Utility". Cell. 179 (6): 1424–1435.e8. doi:10.1016/j.cell.2019.10.033. PMC 6957074. PMID 31761530.
- Lázaro-Muñoz G, Pereira S, Carmi S, Lencz T (March 2021). "Screening embryos for polygenic conditions and traits: ethical considerations for an emerging technology". Genetics in Medicine. 23 (3): 432–434. doi:10.1038/s41436-020-01019-3. PMC 7936952. PMID 33106616.
- Treff NR, Eccles J, Marin D, Messick E, Lello L, Gerber J, et al. (June 2020). "Preimplantation Genetic Testing for Polygenic Disease Relative Risk Reduction: Evaluation of Genomic Index Performance in 11,883 Adult Sibling Pairs". Genes. 11 (6): 648. doi:10.3390/genes11060648. PMC 7349610. PMID 32545548.
- Treff NR, Eccles J, Lello L, Bechor E, Hsu J, Plunkett K, et al. (2019-12-04). "Utility and First Clinical Application of Screening Embryos for Polygenic Disease Risk Reduction". Frontiers in Endocrinology. 10 (845): 845. doi:10.3389/fendo.2019.00845. PMC 6915076. PMID 31920964.
- Munday S, Savulescu J (January 2021). "Three models for the regulation of polygenic scores in reproduction". Journal of Medical Ethics. 47 (12): e91. doi:10.1136/medethics-2020-106588. PMC 8639919. PMID 33462079.
- Kemper JM, Gyngell C, Savulescu J (September 2019). "Subsidizing PGD: The Moral Case for Funding Genetic Selection". Journal of Bioethical Inquiry. 16 (3): 405–414. doi:10.1007/s11673-019-09932-2. PMC 6831526. PMID 31418161.
- Savulescu J (October 2001). Francis L (ed.). "Procreative beneficence: why we should select the best children". Bioethics. 15 (5–6): 413–426. doi:10.1093/oxfordhb/9780199981878.013.26. ISBN 978-0-19-998187-8. PMID 12058767.
- "The Polygenic Score (PGS) Catalog". Polygenic Score (PGS) Catalog. Retrieved 29 April 2020.
An open database of polygenic scores and the relevant metadata required for accurate application and evaluation
- Richardson TG, Harrison S, Hemani G, Davey Smith G (March 2019). "An atlas of polygenic risk score associations to highlight putative causal relationships across the human phenome". eLife. 8: e43657. doi:10.7554/eLife.43657. PMC 6400585. PMID 30835202.
- Khera A, Chaggin M, Aragam KG, Emdin CA, Klarin D, Haas ME, Roselli C, Natarajan P, Kathiresan S (2017-11-15). "Genome-wide polygenic score to identify a monogenic risk-equivalent for coronary disease". bioRxiv 10.1101/218388.
- Mavaddat N, Michailidou K, Dennis J, Lush M, Fachal L, Lee A, et al. (January 2019). "Polygenic Risk Scores for Prediction of Breast Cancer and Breast Cancer Subtypes". American Journal of Human Genetics. 104 (1): 21–34. doi:10.1016/j.ajhg.2018.11.002. PMC 6323553. PMID 30554720.
- Ripke S, Walters JT, O'Donovan MC, et al. (Schizophrenia Working Group of the Psychiatric Genomics Consortium) (2020-09-12). "Mapping genomic loci prioritises genes and implicates synaptic biology in schizophrenia". medRxiv 10.1101/2020.09.12.20192922.
- Wray NR, Lin T, Austin J, McGrath JJ, Hickie IB, Murray GK, Visscher PM (January 2021). "From Basic Science to Clinical Application of Polygenic Risk Scores: A Primer". JAMA Psychiatry. 78 (1): 101–109. doi:10.1001/jamapsychiatry.2020.3049. ISSN 2168-622X. PMID 32997097. S2CID 222169651.
- ^ Wray NR, Lin T, Austin J, McGrath JJ, Hickie IB, Murray GK, Visscher PM (January 2021). "From Basic Science to Clinical Application of Polygenic Risk Scores: A Primer". JAMA Psychiatry. 78 (1): 101–109. doi:10.1001/jamapsychiatry.2020.3049. ISSN 2168-622X. PMID 32997097. S2CID 222169651.
- Brockman DG, Petronio L, Dron JS, Kwon BC, Vosburg T, Nip L, et al. (October 2021). "Design and user experience testing of a polygenic score report: a qualitative study of prospective users". BMC Medical Genomics. 14 (1): 238. doi:10.1186/s12920-021-01056-0. PMC 8485114. PMID 34598685.
- Peck L, Borle K, Folkersen L, Austin J (July 2021). "Why do people seek out polygenic risk scores for complex disorders, and how do they understand and react to results?". European Journal of Human Genetics. 30 (1): 81–87. doi:10.1038/s41431-021-00929-3. PMC 8738734. PMID 34276054. S2CID 236090477.
- Millward M, Tiller J, Bogwitz M, Kincaid H, Taylor S, Trainer AH, Lacaze P (September 2020). "Impact of direct-to-consumer genetic testing on Australian clinical genetics services". European Journal of Medical Genetics. 63 (9): 103968. doi:10.1016/j.ejmg.2020.103968. PMID 32502649.
- Sirugo G, Williams SM, Tishkoff SA (March 2019). "The Missing Diversity in Human Genetic Studies". Cell. 177 (1): 26–31. doi:10.1016/j.cell.2019.02.048. PMC 7380073. PMID 30901543.
- ^ Adeyemo, Adebowale; Balaconis, Mary K.; Darnes, Deanna R.; Fatumo, Segun; Granados Moreno, Palmira; Hodonsky, Chani J.; Inouye, Michael; Kanai, Masahiro; Kato, Kazuto; Knoppers, Bartha M.; Lewis, Anna C. F.; Martin, Alicia R.; McCarthy, Mark I.; Meyer, Michelle N.; Okada, Yukinori; Richards, J. Brent; Richter, Lucas; Ripatti, Samuli; Rotimi, Charles N.; Sanderson, Saskia C.; Sturm, Amy C.; Verdugo, Ricardo A.; Widen, Elisabeth; Willer, Cristen J.; Wojcik, Genevieve L.; Zhou, Alicia (November 2021). "Responsible use of polygenic risk scores in the clinic: potential benefits, risks and gaps". Nature Medicine. 27 (11): 1876–1884. doi:10.1038/s41591-021-01549-6. PMID 34782789. S2CID 244131258.
- Brockman DG, Petronio L, Dron JS, Kwon BC, Vosburg T, Nip L, et al. (January 2021). "Design and user experience testing of a polygenic score report: a qualitative study of prospective users". p. 238. medRxiv 10.1101/2021.04.14.21255397.
- Lineweaver TT, Bondi MW, Galasko D, Salmon DP (February 2014). "Effect of knowledge of APOE genotype on subjective and objective memory performance in healthy older adults". The American Journal of Psychiatry. 171 (2): 201–208. doi:10.1176/appi.ajp.2013.12121590. PMC 4037144. PMID 24170170.
- Parens E, Appelbaum PS (May 2019). "On What We Have Learned and Still Need to Learn about the Psychosocial Impacts of Genetic Testing". The Hastings Center Report. 49 (Suppl 1): S2–S9. doi:10.1002/hast.1011. PMC 6640636. PMID 31268574. S2CID 195786850.
- Varghese MJ (May 2014). "Familial hypercholesterolemia: A review". Annals of Pediatric Cardiology. 7 (2): 107–117. doi:10.4103/0974-2069.132478. PMC 4070199. PMID 24987256.
- Ganna A, Magnusson PK, Pedersen NL, de Faire U, Reilly M, Arnlöv J, et al. (September 2013). "Multilocus genetic risk scores for coronary heart disease prediction". Arteriosclerosis, Thrombosis, and Vascular Biology. 33 (9): 2267–2272. doi:10.1161/atvbaha.113.301218. PMID 23685553. S2CID 182105.
- Merry AH (2010). Coronary heart disease in the Netherlands : incidence, etiology and risk prediction (Thesis). University of Maastricht. doi:10.26481/dis.20101210am. ISBN 9789052789873.
- Nadir MA, Struthers AD (April 2011). "Family history of premature coronary heart disease and risk prediction". Heart. 97 (8): 684, author reply 684. doi:10.1136/hrt.2011.222265. PMID 21367743. S2CID 36394971.
- Neale B (2019-11-26). "Faculty Opinions recommendation of Genomic risk prediction of coronary artery disease in 480,000 adults: implications for primary prevention". Journal of the American College of Cardiology. doi:10.3410/f.734171897.793567857. S2CID 212766426.
- Oram RA, Patel K, Hill A, Shields B, McDonald TJ, Jones A, et al. (March 2016). "A Type 1 Diabetes Genetic Risk Score Can Aid Discrimination Between Type 1 and Type 2 Diabetes in Young Adults". Diabetes Care. 39 (3): 337–344. doi:10.2337/dc15-1111. PMC 5642867. PMID 26577414.
- Sharp SA, Jones SE, Kimmitt RA, Weedon MN, Halpin AM, Wood AR, et al. (October 2020). "A single nucleotide polymorphism genetic risk score to aid diagnosis of coeliac disease: a pilot study in clinical care". Alimentary Pharmacology & Therapeutics. 52 (7): 1165–1173. doi:10.1111/apt.15826. PMID 32790217. S2CID 221127365.
- McBride CM, Koehly LM, Sanderson SC, Kaphingst KA (2010-03-01). "The behavioral response to personalized genetic information: will genetic risk profiles motivate individuals and families to choose more healthful behaviors?". Annual Review of Public Health. 31 (1): 89–103. doi:10.1146/annurev.publhealth.012809.103532. PMID 20070198.
- Saya S, McIntosh JG, Winship IM, Clendenning M, Milton S, Oberoi J, et al. (2020). "A Genomic Test for Colorectal Cancer Risk: Is This Acceptable and Feasible in Primary Care?". Public Health Genomics. 23 (3–4): 110–121. doi:10.1159/000508963. PMID 32688362. S2CID 220669421.
- Turnwald BP, Goyer JP, Boles DZ, Silder A, Delp SL, Crum AJ (January 2019). "Learning one's genetic risk changes physiology independent of actual genetic risk". Nature Human Behaviour. 3 (1): 48–56. doi:10.1038/s41562-018-0483-4. PMC 6874306. PMID 30932047.
- Singh Y (November 2016). "Effectiveness of Screening Programmes for Detection of Major Congenital Heart Diseases". European Journal of Pediatrics. 175 (11): 1582. doi:10.26226/morressier.57d034ced462b80292382f40.
- Esserman LJ (2017-09-13). "The WISDOM Study: breaking the deadlock in the breast cancer screening debate". npj Breast Cancer. 3 (1): 34. doi:10.1038/s41523-017-0035-5. PMC 5597574. PMID 28944288. S2CID 3313995.
- Farooq V, Serruys PW (July 2018). "Risk stratification and risk scores". In Wijns W (ed.). ESC CardioMed. Oxford University Press. pp. 1371–1384. doi:10.1093/med/9780198784906.003.0335. ISBN 978-0-19-878490-6.
- Mbatchou, J; Barnard, L; Backman, J; Marcketta, A; Kosmicki, JA; Ziyatdinov, A; Benner, C; O'Dushlaine, C; Barber, M; Boutkov, B; Habegger, L; Ferreira, M; Baras, A; Reid, J; Abecasis, G; Maxwell, E; Marchini, J (July 2021). "Computationally efficient whole-genome regression for quantitative and binary traits". Nature Genetics. 53 (7): 1097–1103. doi:10.1038/s41588-021-00870-7. PMID 34017140. S2CID 220044465.
- Jurgens, SJ; Pirruccello, JP; Choi, SH; Morrill, VN; Chaffin, M; Lubitz, SA; Lunetta, KL; Ellinor, PT (23 March 2023). "Adjusting for common variant polygenic scores improves yield in rare variant association analyses". Nature Genetics. 55 (4): 544–548. doi:10.1038/s41588-023-01342-w. PMC 11078202. PMID 36959364. S2CID 239054376.
- Bennett, D; O'Shea, D; Ferguson, J; Morris, D; Seoighe, C (1 October 2021). "Controlling for background genetic effects using polygenic scores improves the power of genome-wide association studies". Scientific Reports. 11 (1): 19571. Bibcode:2021NatSR..1119571B. doi:10.1038/s41598-021-99031-3. PMC 8486788. PMID 34599249.
- ^ Wray NR, Kemper KE, Hayes BJ, Goddard ME, Visscher PM (April 2019). "Complex Trait Prediction from Genome Data: Contrasting EBV in Livestock to PRS in Humans: Genomic Prediction". Genetics. 211 (4): 1131–1141. doi:10.1534/genetics.119.301859. PMC 6456317. PMID 30967442.
Further reading
- Francesca Forzano, Olga Antonova, Angus Clarke, Guido de Wert, Sabine Hentze, Yalda Jamshidi, Yves Moreau, Markus Perola, Inga Prokopenko, Andrew Read, Alexandre Reymond, Vigdis Stefansdottir, Carla van El (2022). "The use of polygenic risk scores in pre-implantation genetic testing: an unproven, unethical practice." Nature 30, pages 493–495.
 is obtained as the sum across m number of SNPs with risk-increasing alleles weighted by their weights, i.e.,
is obtained as the sum across m number of SNPs with risk-increasing alleles weighted by their weights, i.e.,  .
.
