| Tat | |||||||||
|---|---|---|---|---|---|---|---|---|---|
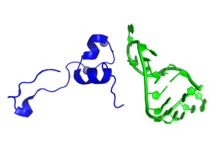 Representation of a fragment of the Tat protein (blue) from HIV along with two Zn molecules shown as grey spheres, in complex with HIV TAR RNA (green). Prepared using the data published under PDB code: 6CYT. Representation of a fragment of the Tat protein (blue) from HIV along with two Zn molecules shown as grey spheres, in complex with HIV TAR RNA (green). Prepared using the data published under PDB code: 6CYT. | |||||||||
| Identifiers | |||||||||
| Symbol | Tat | ||||||||
| Pfam | PF00539 | ||||||||
| InterPro | IPR001831 | ||||||||
| PROSITE | PDOC00836 | ||||||||
| SCOP2 | 1tvs / SCOPe / SUPFAM | ||||||||
| |||||||||
In molecular biology, Tat is a protein that is encoded for by the tat gene in HIV-1. Tat is a regulatory protein that drastically enhances the efficiency of viral transcription. Tat stands for "Trans-Activator of Transcription". The protein consists of between 86 and 101 amino acids depending on the subtype. Tat vastly increases the level of transcription of the HIV dsDNA. Before Tat is present, a small number of RNA transcripts will be made, which allow the Tat protein to be produced. Tat then binds to cellular factors and mediates their phosphorylation, resulting in increased transcription of all HIV genes, providing a positive feedback cycle. This in turn allows HIV to have an explosive response once a threshold amount of Tat is produced, a useful tool for defeating the body's response.
Tat also appears to play a more direct role in the HIV disease process. The protein is released by infected cells in culture, and is found in the blood of HIV-1 infected patients.
It can be absorbed by cells that are not infected with HIV, and can act directly as a toxin producing cell death via apoptosis in uninfected "bystander" T cells, assisting in progression toward AIDS.
By antagonizing the CXCR4 receptor, Tat also appears to selectively encourage the reproduction of less virulent M-tropic (macrophage-tropic) strains of HIV (which use the CCR5 receptor) early in the course of infection, allowing the more rapidly pathogenic T-tropic (T-cell-tropic) strains (which use the CXCR4 receptor) to emerge later after mutating from M-tropic strains.
Function and mechanism
Like other lentiviruses, HIV-1 encodes a trans-activating regulatory protein (Tat), which is essential for efficient transcription of the viral genome. Tat acts by binding to an RNA stem-loop structure, the trans-activation response element (TAR), found at the 5′ ends of nascent HIV-1 transcripts. In binding to TAR, Tat alters the properties of the transcription complex, recruits the positive transcription elongation complex (P-TEFb) of cellular CDK9 and cyclin T1, and hence increases the production of full-length viral RNA.
Tat protein also associates with RNA polymerase II complexes during early transcription elongation after the promoter clearance and before the synthesis of full-length TAR RNA transcript. This interaction of Tat with RNA polymerase II elongation complexes is P-TEFb-independent. There are two Tat binding sites on each transcription elongation complex; one is located on TAR RNA and the other one on RNA polymerase II near the exit site for nascent mRNA transcripts, which suggests the involvement of two Tat molecules in facilitating one round of HIV-1 mRNA synthesis.
The minimum Tat sequence that can mediate specific TAR binding in vitro has been mapped to a basic domain of 10 amino acids, comprising mostly Arg and Lys residues. Regulatory activity, however, also requires the 47 N-terminal residues, which interact with components of the transcription complex and function as a transcriptional activation domain.
Tat also uses an unusual transcellular transport pathway. Firstly, it binds with high affinity to Phosphatidylinositol 4,5-bisphosphate (PI(4,5)P2), found on the inner surface of the cell membrane, to facilitate Tat recruitment. Tat then crosses the plasma membrane to reach the extracellular space. Tat secretion by infected cells is highly active, and export is the major destination for HIV-1 Tat.
Structure
The basic region of HIV-Tat protein is suggested to form an alpha helix. The basic region is involved in RNA (TAR, trans-activation response element) binding and Tat proteins thus belong to the family of arginine-rich motif (ARM) RNA binding proteins.
Protein transduction domain
Tat contains a protein transduction domain (PTD), which is also known as a cell-penetrating peptide. Originally characterised by Frankel and Pabo (1988) and Green and Loewenstein (1988), this domain allows Tat to enter cells by crossing the cell membrane. The amino acid sequence of the PTD is YGRKKRRQRRR. This cationic eleven-amino-acid peptide is sufficient to target large molecules, including nanoparticles that display it, for transduction into cells. Unlike the heparinase-independent transduction of isolated PTD polypeptide, internalization of full-length Tat is mediated by interactions with negatively-charged heparan sulfate proteoglycans at the cell surface. This translocation is partly mediated by potocytosis.
The nuclear localisation signal found within the PTD, GRKKR, mediates further translocation of Tat into the cell nucleus. As of 2000 The biological role of this domain and exact mechanism of transfer is unknown.
Clinical significance
Inhibition of Tat has been investigated. It has been suggested that Tat antagonists may be of use in the treatment of HIV infections.
Biosantech has developed a novel vaccine called Tat Oyi, which aims at the Tat protein. The company's HIV vaccine candidate is not toxic to 48 HIV-positive patients enrolled in a double-blind study taking place in France. A Phase I/IIa study, published in 2016, shows a reduction in viral RNA for one of three doses tested. A dose-dependent response was not observed, raising questions about the robustness of the findings.
References
- Genes,+tat at the U.S. National Library of Medicine Medical Subject Headings (MeSH)
- ^ Debaisieux S, Rayne F, Yezid H, Beaumelle B (2012). "The ins and outs of HIV-1 Tat". Traffic. 13 (3): 355–63. doi:10.1111/j.1600-0854.2011.01286.x. PMID 21951552.
- Jeang KT (June 1997). "HIV-1 TAT: Structure and Function". In Myers G, Foley B, Korber B, Mellors J, Jeang K, Wain-Hobson S (eds.). Human Retroviruses and AIDS: A Compilation and Analysis of Nucleic Acid and Amino Acid Sequences. Los Alamos National Laboratory. pp. III-3–III-18. CiteSeerX 10.1.1.321.7430.
- Kim JB, Sharp PA (April 2001). "Positive transcription elongation factor B phosphorylates hSPT5 and RNA polymerase II carboxyl-terminal domain independently of cyclin-dependent kinase-activating kinase". J. Biol. Chem. 276 (15): 12317–23. doi:10.1074/jbc.M010908200. PMID 11145967.
- ^ Xiao H, Neuveut C, Tiffany HL, et al. (October 2000). "Selective CXCR4 antagonism by Tat: implications for in vivo expansion of coreceptor use by HIV-1". Proc. Natl. Acad. Sci. U.S.A. 97 (21): 11466–71. Bibcode:2000PNAS...9711466X. doi:10.1073/pnas.97.21.11466. PMC 17223. PMID 11027346.
- Campbell GR, Pasquier E, Watkins J, Bourgarel-Rey V, Peyrot V, Esquieu D, Barbier P, de Mareuil J, Braguer D, Kaleebu P, Yirrell DL, Loret EP (November 2004). "The glutamine-rich region of the HIV-1 Tat protein is involved in T-cell apoptosis". J. Biol. Chem. 279 (46): 48197–204. doi:10.1074/jbc.M406195200. PMID 15331610.
- Vaishnav YN, Wong-Staal F (1991). "The biochemistry of AIDS". Annu. Rev. Biochem. 60: 577–630. doi:10.1146/annurev.bi.60.070191.003045. PMID 1883204.
- ^ Mujeeb A, Bishop K, Peterlin BM, Turck C, Parslow TG, James TL (August 1994). "NMR structure of a biologically active peptide containing the RNA-binding domain of human immunodeficiency virus type 1 Tat". Proc. Natl. Acad. Sci. U.S.A. 91 (17): 8248–52. Bibcode:1994PNAS...91.8248M. doi:10.1073/pnas.91.17.8248. PMC 44583. PMID 8058789.
- Zhou C, Rana TM (July 2002). "A bimolecular mechanism of HIV-1 Tat protein interaction with RNA polymerase II transcription elongation complexes". J. Mol. Biol. 320 (5): 925–42. doi:10.1016/S0022-2836(02)00556-9. PMID 12126615.
- Selby MJ, Peterlin BM (August 1990). "Trans-activation by HIV-1 Tat via a heterologous RNA binding protein". Cell. 62 (4): 769–76. doi:10.1016/0092-8674(90)90121-T. PMID 2117500. S2CID 30833580.
- Kashanchi F, Piras G, Radonovich MF, Duvall JF, Fattaey A, Chiang CM, Roeder RG, Brady JN (January 1994). "Direct interaction of human TFIID with the HIV-1 transactivator Tat". Nature. 367 (6460): 295–9. Bibcode:1994Natur.367..295K. doi:10.1038/367295a0. PMID 8121496. S2CID 4362048.
- Tahirov TH, Babayeva ND, Varzavand K, Cooper JJ, Sedore SC, Price DH (June 2010). "Crystal structure of HIV-1 Tat complexed with human P-TEFb". Nature. 465 (7299): 747–51. Bibcode:2010Natur.465..747T. doi:10.1038/nature09131. PMC 2885016. PMID 20535204.
- ^ Schwarze SR, Hruska KA, Dowdy SF (July 2000). "Protein transduction: unrestricted delivery into all cells?". Trends Cell Biol. 10 (7): 290–5. doi:10.1016/S0962-8924(00)01771-2. PMID 10856932.
- Dietz GP, Bähr M (October 2004). "Delivery of bioactive molecules into the cell: the Trojan horse approach". Molecular and Cellular Neuroscience. 27 (2): 85–131. doi:10.1016/j.mcn.2004.03.005. PMID 15485768. S2CID 21373286.
- Frankel AD, Pabo CO (December 1988). "Cellular uptake of the tat protein from human immunodeficiency virus". Cell. 55 (6): 1189–93. doi:10.1016/0092-8674(88)90263-2. PMID 2849510.
- Green M, Loewenstein PM (December 1988). "Autonomous functional domains of chemically synthesized human immunodeficiency virus tat trans-activator protein". Cell. 55 (6): 1179–88. doi:10.1016/0092-8674(88)90262-0. PMID 2849509. S2CID 26846467.
- ^ MacKay, J. Andrew; Szoka, Jr., Francis C. (2003). "HIV TAT Protein Transduction Domain Mediated Cell Binding and Intracellular Delivery of Nanoparticles". Journal of Dispersion Science and Technology. 24 (3): 465–473. doi:10.1081/dis-120021802. PMC 2929673. PMID 20808712.
- Silhol, Michelle; Tyagi, Mudit; Giacca, Mauro; Lebleu, Bernard; Vivès, Eric (2002). "Different mechanisms for cellular internalization of the HIV-1 Tat-derived cell penetrating peptide and recombinant proteins fused to Tat". European Journal of Biochemistry. 269 (2): 494–501. doi:10.1046/j.0014-2956.2001.02671.x. PMID 11856307.
- ^ Eguchi, Akiko; Akuta, Teruo; Okuyama, Hajime; et al. (2001). "Protein Transduction Domain of HIV-1 Tat Protein Promotes Efficient Delivery of DNA into Mammalian Cells". Journal of Biological Chemistry. 276 (28): 26204–26210. doi:10.1074/jbc.M010625200. PMID 11346640.
- Ruben S, Perkins A, Purcell R, et al. (January 1989). "Structural and functional characterization of human immunodeficiency virus Tat protein". Journal of Virology. 63 (1): 1–8. doi:10.1128/JVI.63.1.1-8.1989. PMC 247650. PMID 2535718.
- Hauber J, Malim MH, Cullen BR (March 1989). "Mutational analysis of the conserved basic domain of human immunodeficiency virus tat protein". Journal of Virology. 63 (3): 1181–7. doi:10.1128/JVI.63.3.1181-1187.1989. PMC 247813. PMID 2536828.
- Cook JA, August A, Henderson AJ (July 2002). "Recruitment of phosphatidylinositol 3-kinase to CD28 inhibits HIV transcription by a Tat-dependent mechanism". J. Immunol. 169 (1): 254–60. doi:10.4049/jimmunol.169.1.254. PMID 12077252.
- Bedoya LM, Beltrán M, Sancho R, et al. (October 2005). "4-Phenylcoumarins as HIV transcription inhibitors". Bioorg. Med. Chem. Lett. 15 (20): 4447–50. doi:10.1016/j.bmcl.2005.07.041. PMID 16137881.
- Loret EP, et al. (April 2016). "Intradermal injection of a Tat Oyi-based therapeutic HIV vaccine reduces of 1.5 log copies/mL the HIV RNA rebound median and no HIV DNA rebound following cART interruption in a phase I/II randomized controlled clinical trial". Retrovirology. 13: 21. doi:10.1186/s12977-016-0251-3. PMC 4818470. PMID 27036656.
| Viral proteins (early and late) | |||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| DNA |
| ||||||||||||||||||||||||||||||||||||||||||||||||||
| RNA |
| ||||||||||||||||||||||||||||||||||||||||||||||||||
| RT |
| ||||||||||||||||||||||||||||||||||||||||||||||||||