| This article is missing information about pfam box for the actual catalytic domain; member of breaking-rejoining enzyme (BRE) superfamily; link between BRE and AraC homeobox per ECOD. Please expand the article to include this information. Further details may exist on the talk page. (February 2021) |
| DNA topoisomerase I, N-terminal (non-catalytic), viral | |||||||||
|---|---|---|---|---|---|---|---|---|---|
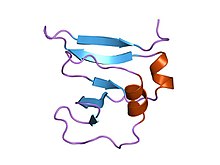 amino terminal 9kda domain of vaccinia virus dna topoisomerase i residues 1-77, experimental electron density for residues 1-77 amino terminal 9kda domain of vaccinia virus dna topoisomerase i residues 1-77, experimental electron density for residues 1-77 | |||||||||
| Identifiers | |||||||||
| Symbol | VirDNA-topo-I_N | ||||||||
| Pfam | PF09266 | ||||||||
| InterPro | IPR015346 | ||||||||
| SCOP2 | 1vcc / SCOPe / SUPFAM | ||||||||
| |||||||||
In molecular biology Type I topoisomerases are enzymes that cut one of the two strands of double-stranded DNA, relax the strand, and reanneal the strand. They are further subdivided into two structurally and mechanistically distinct topoisomerases: type IA and type IB.
- Type IA topoisomerases change the linking number of a circular DNA strand by units of strictly 1.
- Type IB topoisomerases change the linking number by multiples of 1 (n).
Historically, type IA topoisomerases are referred to as prokaryotic topo I, while type IB topoisomerases are referred to as eukaryotic topoisomerase. This distinction, however, no longer applies as type IA and type IB topoisomerases exist in all domains of life.
Functionally, these subclasses perform very specialized functions. Prokaryotic topoisomerase I (topo IA) can only relax negative supercoiled DNA, whereas eukaryotic topoisomerase I (topo IB) can introduce positive supercoils, separating the DNA of daughter chromosomes after DNA replication, and relax DNA.
Function
These enzymes have several functions: to remove DNA supercoils during transcription and DNA replication; for strand breakage during recombination; for chromosome condensation; and to disentangle intertwined DNA during mitosis.
Structure
This domain assumes a beta(2)-alpha-beta-alpha-beta(2) fold, with a left-handed crossover between strands beta2 and beta3. It has a four criss-crossed beta-strands surrounded by four alpha-helices that are arranged in a Rossmann fold
Mechanisms
Type I topoisomerases are ATP-independent enzymes (except for reverse gyrase), and can be subdivided according to their structure and reaction mechanisms: type IA (bacterial and archaeal topoisomerase I, topoisomerase III and reverse gyrase) and type IB (eukaryotic topoisomerase I and topoisomerase V). These enzymes are primarily responsible for relaxing positively and/or negatively supercoiled DNA, except for reverse gyrase, which can introduce positive supercoils into DNA.
DNA topoisomerases regulate the number of topological links between two DNA strands (i.e. change the number of superhelical turns) by catalysing transient single- or double-strand breaks, crossing the strands through one another, then resealing the breaks.
Classes
DNA topoisomerases are divided into two classes: type I enzymes (EC; topoisomerases I, III and V) break single-strand DNA, and type II enzymes (EC; topoisomerases II, IV and VI) break double-strand DNA.
Type IA topoisomerases
Introduction
Type IA topoisomerases, historically said to be found in prokaryotes, create a single break in DNA and pass a second strand or duplex through the break. This strand passage mechanism shares several features with type IIA topoisomerases. They both form a 5' phosphotyrosine intermediate, and require a divalent metal ion to perform its work. Unlike type II topoisomerases, type IA topoisomerases do not use energy to do its work (with the notable exception of reverse gyrase, see below).
Structure
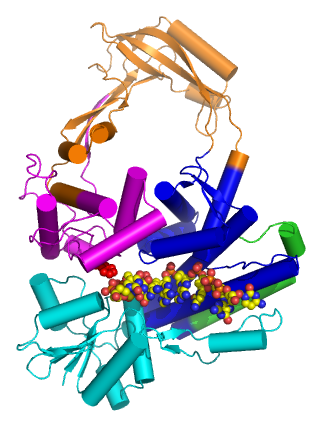
Type IA topoisomerases have several domains, often number Domain 1-4. Domain I contains a Toprim domain (a Rossman fold known to coordinate Magnesium ions), domain IV and domain III each consist of a helix-turn-helix (HTH) domain; the catalytic tyrosine resides on the HTH of domain III. Domain II is a flexible bridge between domains III and IV. The structure of type IA topoisomerase resembles a lock, with Domains I, III and IV lying on the bottom of the structure. The structure of topo III (see below) bound to single-stranded DNA (pdb id = 1I7D) shows how the HTH and Toprim domain are coordinated about the DNA.
Type IA topoisomerase variants
There are several variants of Type IA topoisomerases, differing by appendages attached to the main core (sometimes referred to as the "topo-fold"). Members of this subclass include topo I, topo III (which contain additional Zinc-binding motifs), and reverse gyrase. Reverse gyrase is particularly interesting because an ATPase domain, which resembles the helicase-like domain of the Rho transcription factor, is attached (the structure of reverse gyrase was solved by Rodriguez and Stock, EMBO J 2002). The enzyme uses the hydrolysis of ATP to introduce positive supercoils and overwinds DNA, a feature attractive in hyperthermophiles, in which reverse gyrase is known to exist. Rodriguez and Stock have done further work to identify a "latch" that is involved in communicating the hydrolysis of ATP to the introduction of positive supercoils.
The topo III variant is likewise very interesting because it has zinc-binding motifs that is thought to bind single-stranded DNA. Topo III has been identified to be associated with the BLM (for Bloom Syndrome) helicase during recombination.
Mechanism
Type IA topoisomerases operate through a strand-passage mechanism, using a single gate (in contrast with type II topoisomerases). First, the single-stranded DNA binds domain III and I. The catalytic tyrosine cleaves the DNA backbone, creating a transient 5' phosphotyrosine intermediate. The break is then separated, using domain II as a hinge, and a second duplex or strand of DNA is passed through. Domain III and I close and the DNA is re-annealed.
Type IB topoisomerases

Introduction
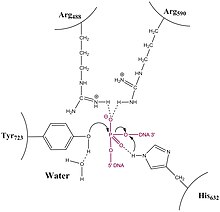
In contrast to type IA topoisomerases, type 1B Topoisomerase solves the problem of overwound and underwound (also referred to as positively or negatively supercoiled) DNA through a hindered rotary mechanism. Crystal structures, biochemistry, and single molecule experiments have contributed to a general mechanism. The enzyme first wraps around DNA and creates a single, 3' phosphotyrosine intermediate. The 5' end is then free to rotate, twisting it about the other strand, to relax DNA until the topoisomerase re-ligates the broken strands.
Structure
The structure of topo IB bound to DNA has been solved (pdb id = 1A36). Topo IB is composed of an NTD, a capping lobe, a catalytic lobe, and a C-terminal domain. The capping lobe and catalytic lobe wrap around the DNA.
Mechanism
Relaxation is not an active process and energy (in the form of ATP) is not spent during the nicking or ligation steps; this is because the reaction between the tyrosine residue at the active site of the enzyme with the phosphodiester DNA backbone simply replaces one phosphomonoester bond with another. The topoisomerase also does not use ATP during uncoiling of the DNA; rather, the torque present in the DNA drives the uncoiling and proceeds on average energetically downhill. Recent single molecule experiments have confirmed what bulk-plasmid relaxation experiments have proposed earlier, which is that uncoiling of the DNA is torque-driven and proceeds until religation occurs. No data suggest that Topo IB "controls" the swiveling insofar as that it has a mechanism in place that triggers religation after a specific number of supercoils removed. On the contrary, single-molecule experiments suggest that religation is a random process and has some probability of occurring each time the swiveling 5'-OH end comes in close proximity with the attachment site of the enzyme-linked 3'-end.
Type IB topoisomerases were originally identified in eukaryotes and in viruses. Viral topo I is unique because it binds DNA in a sequence-specific manner.
See the article TOP1 for further details on this well-studied type 1B topoisomerase.
Type IC topoisomerases
A third type of topoisomerase I was identified, topo V, in the archaeon Methanopyrus kandleri. Topo V is the founding member, and so far the only member, of the type IC topoisomerase, although some authors suggest it may have viral origins. The crystal structure of topo V was solved. Type IC topoisomerases work through a controlled rotary mechanism, much like type IB topoisomerases(pdb ID = 2CSB and 2CSD), but the fold is unique.
Intermediates
All topoisomerases form a phosphotyrosine intermediate between the catalytic tyrosine of the enzyme and the scissile phosphoryl of the DNA backbone.
- Type IA topoisomerases form a covalent linkage between the catalytic tyrosine and the 5'-phosphoryl.
- Type IB enzymes form a covalent 3'-phosphotyrosine intermediate.
- Type IC topoisomerases form a covalent 3'-phosphotyrosine intermediate.
This intermediate is isoenergetic, meaning that the forward cleavage reaction and the backward religation reaction are both energetically equal. As such, no outside energy source is necessary to conduct this reaction.
Inhibition
Main article: Topoisomerase inhibitorAs topoisomerases generate breaks in DNA, they are targets of small-molecule inhibitors that inhibit the enzyme. Type 1 topoisomerase is inhibited by irinotecan, topotecan, hexylresorcinol and camptothecin.
The human topoisomerase type IB enzyme forms a covalent 3'-phosphotyrosine intermediate, the topoisomerase 1-cleavage-complex (Top1cc). The active irinotecan metabolite, SN-38, acts by trapping (making a ternary complex with) a subset of Top1cc, those with a guanine +1 in the DNA sequence. One irinotecan-derived SN-38 molecule stacks against the base pairs flanking the topoisomerase-induced cleavage site and poisons (inactivates) the topoisomerase 1 enzyme.
Upon bacteriophage (phage) T4 infection of its bacterial host, Escherichia coli, the phage genome specifies a gene product (gp55.2) that inhibits the bacterial topoisomerase I. Gp55.2 binds DNA and specifically blocks the relaxation of negatively supercoiled DNA by topoisomerase I. This inhibition appears to be an adaptation to subtly modulate host topoisomerase I activity during infection to ensure optimal phage yield.
Synthetic lethality
Main article: TOP1 § Synthetic lethalitySynthetic lethality arises when a combination of deficiencies in the expression of two or more genes leads to cell death, whereas a deficiency in only one of these genes does not. The deficiencies can arise through mutation, epigenetic alteration or by inhibition of a gene's expression.
Topoisomerase 1 inhibition is synthetically lethal with deficiency of expression of certain DNA repair genes. In human patients the deficient DNA repair genes include WRN and MRE11. In pre-clinical studies related to cancer, the deficient DNA repair genes include ATM and NDRG1.
Autoantibodies
Main article: Anti-Scl-70 antibodiesAutoantibodies targeted against type I topoisomerase are called anti-Scl-70 antibodies, named by the association with scleroderma and the 70 kD extractable immunoreactive fragment that can be obtained from the otherwise larger (100-105 kD) target topoisomerase antigen (called the SCL-70 Antigen) of the antibodies.
References
- Wang JC (June 2002). "Cellular roles of DNA topoisomerases: a molecular perspective". Nat. Rev. Mol. Cell Biol. 3 (6): 430–40. doi:10.1038/nrm831. PMID 12042765. S2CID 205496065.
- Champoux JJ (2001). "DNA topoisomerases: structure, function, and mechanism". Annu. Rev. Biochem. 70: 369–413. doi:10.1146/annurev.biochem.70.1.369. PMID 11395412.
- Sharma A; Hanai R; Mondragón A (August 1994). "Crystal structure of the amino-terminal fragment of vaccinia virus DNA topoisomerase I at 1.6 A resolution". Structure. 2 (8): 767–77. doi:10.1016/s0969-2126(94)00077-8. PMID 7994576.
- Roca J (April 1995). "The mechanisms of DNA topoisomerases". Trends Biochem. Sci. 20 (4): 156–60. doi:10.1016/s0968-0004(00)88993-8. PMID 7770916.
- Gadelle D, Filée J, Buhler C, Forterre P (March 2003). "Phylogenomics of type II DNA topoisomerases". BioEssays. 25 (3): 232–42. doi:10.1002/bies.10245. PMID 12596227. S2CID 4642743.
- Lima, C. D.; Wang, J. C.; Mondragón, A (1994). "Three-dimensional structure of the 67K N-terminal fragment of E. Coli DNA topoisomerase I". Nature. 367 (6459): 138–46. Bibcode:1994Natur.367..138L. doi:10.1038/367138a0. PMID 8114910. S2CID 4314431.
- Changela, A; Digate, R. J.; Mondragón, A (2001). "Crystal structure of a complex of a type IA DNA topoisomerase with a single-stranded DNA molecule". Nature. 411 (6841): 1077–81. Bibcode:2001Natur.411.1077C. doi:10.1038/35082615. PMID 11429611. S2CID 4426078.
- Forterre P (June 2006). "DNA topoisomerase V: a new fold of mysterious origin". Trends Biotechnol. 24 (6): 245–7. doi:10.1016/j.tibtech.2006.04.006. PMID 16650908.
- Taneja B, Patel A, Slesarev A, Mondragón A (January 2006). "Structure of the N-terminal fragment of topoisomerase V reveals a new family of topoisomerases". EMBO J. 25 (2): 398–408. doi:10.1038/sj.emboj.7600922. PMC 1383508. PMID 16395333.
- Taneja, B; Schnurr, B; Slesarev, A; Marko, J. F.; Mondragón, A (2007). "Topoisomerase V relaxes supercoiled DNA by a constrained swiveling mechanism". Proceedings of the National Academy of Sciences. 104 (37): 14670–5. Bibcode:2007PNAS..10414670T. doi:10.1073/pnas.0701989104. PMC 1976220. PMID 17804808.
- ^ Pommier Y (2013). "Drugging topoisomerases: lessons and challenges". ACS Chem. Biol. 8 (1): 82–95. doi:10.1021/cb300648v. PMC 3549721. PMID 23259582.
- Mattenberger Y, Silva F, Belin D (2015). "55.2, a phage T4 ORFan gene, encodes an inhibitor of Escherichia coli topoisomerase I and increases phage fitness". PLoS One. 10 (4): e0124309. doi:10.1371/journal.pone.0124309. PMC 4396842. PMID 25875362.
- Agrelo R, Cheng WH, Setien F, Ropero S, Espada J, Fraga MF, Herranz M, Paz MF, Sanchez-Cespedes M, Artiga MJ, Guerrero D, Castells A, von Kobbe C, Bohr VA, Esteller M (2006). "Epigenetic inactivation of the premature aging Werner syndrome gene in human cancer". Proc. Natl. Acad. Sci. U.S.A. 103 (23): 8822–7. Bibcode:2006PNAS..103.8822A. doi:10.1073/pnas.0600645103. PMC 1466544. PMID 16723399.
- Pavelitz T, Renfro L, Foster NR, Caracol A, Welsch P, Lao VV, Grady WB, Niedzwiecki D, Saltz LB, Bertagnolli MM, Goldberg RM, Rabinovitch PS, Emond M, Monnat RJ, Maizels N (2014). "MRE11-deficiency associated with improved long-term disease free survival and overall survival in a subset of stage III colon cancer patients in randomized CALGB 89803 trial". PLOS ONE. 9 (10): e108483. Bibcode:2014PLoSO...9j8483P. doi:10.1371/journal.pone.0108483. PMC 4195600. PMID 25310185.
- Subhash VV, Tan SH, Yeo MS, Yan FL, Peethala PC, Liem N, Krishnan V, Yong WP (2016). "ATM expression predicts Veliparib and Irinotecan sensitivity in gastric cancer by mediating P53 independent regulation of cell cycle and apoptosis". Mol. Cancer Ther. 15 (12): 3087–96. doi:10.1158/1535-7163.MCT-15-1002. PMID 27638859.
- Wissing MD, Mendonca J, Kim E, Kim E, Shim JS, Kaelber NS, Kant H, Hammers H, Commes T, Van Diest PJ, Liu JO, Kachhap SK (2013). "Identification of cetrimonium bromide and irinotecan as compounds with synthetic lethality against NDRG1 deficient prostate cancer cells". Cancer Biol. Ther. 14 (5): 401–10. doi:10.4161/cbt.23759. PMC 3672184. PMID 23377825.
- Dominick G, Bowman J, Li X, Miller RA, Garcia GG (2016). "mTOR regulates the expression of DNA damage response enzymes in long-lived Snell dwarf, GHRKO, and PAPPA-KO mice". Aging Cell. 16 (1): 52–60. doi:10.1111/acel.12525. PMC 5242303. PMID 27618784.
- Product Name: SCL-70 Antigen Archived 2006-03-19 at the Wayback Machine at ImmunoVision.com, retrieved April 2011
External links
- DNA+Topoisomerases,+Type+I at the U.S. National Library of Medicine Medical Subject Headings (MeSH)
| Isomerase: topoisomerases (EC 5.6) | |
|---|---|
| 5.6.2 | |
| Enzymes | |
|---|---|
| Activity | |
| Regulation | |
| Classification | |
| Kinetics | |
| Types |
|