| Revision as of 17:52, 9 May 2021 view sourceJtbobwaysf (talk | contribs)Extended confirmed users12,436 edits →Reservoir and zoonotic origin: UNDUETag: Reverted← Previous edit | Revision as of 20:29, 9 May 2021 view source RandomCanadian (talk | contribs)Extended confirmed users, Pending changes reviewers, Rollbackers36,695 editsm Rollback edit(s) by Jtbobwaysf (talk): WP:VAGUEWAVE. Unless you're objecting to the NYT as a WP:RS, I don't see how this is undue. (RW 16.1)Tag: RollbackNext edit → | ||
| Line 81: | Line 81: | ||
| Although the role of ] as an intermediate host was initially posited (a study published in July 2020 suggested that pangolins are an intermediate host of SARS‑CoV‑2-like coronaviruses<ref name="XiaoK2020July" /><ref name="ZhaoJ2020" />), subsequent studies have not substantiated their contribution to the spillover.<ref name="WHO_GlobalOrigin_China" /> Evidence against this hypothesis includes the fact that pangolin virus samples are too distant to SARS-CoV-2: isolates obtained from pangolins seized in ] were only 92% identical in sequence to the SARS‑CoV‑2 genome. In addition, despite similarities in a few critical amino acids,<ref name="HuB2020Oct" /> pangolin virus samples exhibit poor binding to the human ACE2 receptor.<ref name="GiovanettiM2020Nov" /> | Although the role of ] as an intermediate host was initially posited (a study published in July 2020 suggested that pangolins are an intermediate host of SARS‑CoV‑2-like coronaviruses<ref name="XiaoK2020July" /><ref name="ZhaoJ2020" />), subsequent studies have not substantiated their contribution to the spillover.<ref name="WHO_GlobalOrigin_China" /> Evidence against this hypothesis includes the fact that pangolin virus samples are too distant to SARS-CoV-2: isolates obtained from pangolins seized in ] were only 92% identical in sequence to the SARS‑CoV‑2 genome. In addition, despite similarities in a few critical amino acids,<ref name="HuB2020Oct" /> pangolin virus samples exhibit poor binding to the human ACE2 receptor.<ref name="GiovanettiM2020Nov" /> | ||
| All available evidence suggests that SARS‑CoV‑2 has a natural animal origin and is not ].<ref name="WHOorigins" /> Nevertheless, early in the pandemic, conspiracy theories spread on social media claiming that the virus was ] by China at the ] |
All available evidence suggests that SARS‑CoV‑2 has a natural animal origin and is not ].<ref name="WHOorigins" /> Nevertheless, early in the pandemic, conspiracy theories spread on social media claiming that the virus was ] by China at the ],<ref name=Zoumpourlis/> amplified by ] in the American far-right.<ref>{{cite news |last1=Qin |first1=Amy |last2=Wang |first2=Vivian |last3=Hakim |first3=Danny |title=How Steve Bannon and a Chinese Billionaire Created a Right-Wing Coronavirus Media Sensation |url=https://www.nytimes.com/2020/11/20/business/media/steve-bannon-china.html |work=The New York Times |date=20 November 2020 |archive-url=https://web.archive.org/web/20210430175006/https://www.nytimes.com/2020/11/20/business/media/steve-bannon-china.html |archive-date=30 April 2021}}</ref> A few individuals, including former CDC director ], have claimed, without evidence, that the virus may have been studied by and escaped from the Institute.<ref name=":0">{{Cite news|last=Gorman|first=James|last2=Barnes|first2=Julian E.|date=2021-03-26|title=The C.D.C.’s ex-director offers no evidence in favoring speculation that the coronavirus originated in a lab.|language=en-US|work=The New York Times|url=https://www.nytimes.com/2021/03/26/science/redfield-coronavirus-wuhan-lab.html|access-date=2021-04-29|issn=0362-4331}}</ref> Most virologists who have studied coronaviruses consider the possibility very remote,<ref>{{Cite journal|last=Hakim|first=Mohamad S.|date=14 February 2021|title=SARS-CoV-2, Covid-19, and the debunking of conspiracy theories|url=https://pubmed.ncbi.nlm.nih.gov/33586302/|journal=Reviews in Medical Virology|pages=e2222|doi=10.1002/rmv.2222|issn=1099-1654|pmid=33586302|pmc=7995093}}</ref><ref name="Graham2020immunity">{{Cite journal|last1=Graham|first1=Rachel L.|last2=Baric|first2=Ralph S.|date=19 May 2020|title=SARS-CoV-2: Combating Coronavirus Emergence|journal=Immunity|volume=52|issue=5|pages=734–736|doi=10.1016/j.immuni.2020.04.016|issn=1074-7613|pmc=7207110|pmid=32392464}}</ref> and the March 2021 WHO report on the joint WHO-China study stated that such an explanation is "extremely unlikely".<ref name=":0" /><ref name="WHO_GlobalOrigin_China" /> | ||
| ===Phylogenetics and taxonomy=== | ===Phylogenetics and taxonomy=== | ||
Revision as of 20:29, 9 May 2021
Virus that causes coronavirus disease 2019 (COVID-19) This article is about the virus that causes COVID-19. For the virus that causes SARS, see Severe acute respiratory syndrome coronavirus. For the species to which both viruses belong, see Severe acute respiratory syndrome–related coronavirus.Template:Use Commonwealth English
| Severe acute respiratory syndrome coronavirus 2 | |
|---|---|

| |
| Transmission electron micrograph of SARS-CoV-2 virions with visible coronae | |

| |
| Illustration of a SARS-CoV-2 virion Red protrusions: spike proteins (S) Grey coating: the envelope, composed mainly of lipids, which can be destroyed with alcohol or soap Yellow deposits: envelope proteins (E) Orange deposits: membrane proteins (M) | |
| Virus classification | |
| (unranked): | Virus |
| Realm: | Riboviria |
| Kingdom: | Orthornavirae |
| Phylum: | Pisuviricota |
| Class: | Pisoniviricetes |
| Order: | Nidovirales |
| Family: | Coronaviridae |
| Genus: | Betacoronavirus |
| Subgenus: | Sarbecovirus |
| Species: | Severe acute respiratory syndrome–related coronavirus |
| Virus: | Severe acute respiratory syndrome coronavirus 2 |
| Variants | |
| Synonyms | |
| |
Severe acute respiratory syndrome coronavirus 2 (SARS‑CoV‑2) is the virus that causes COVID-19 (coronavirus disease 2019), the respiratory illness responsible for the COVID-19 pandemic. Also colloquially known simply as the coronavirus, it was previously referred to by its provisional name, 2019 novel coronavirus (2019-nCoV), and has also been called human coronavirus 2019 (HCoV-19 or hCoV-19). The World Health Organization declared the outbreak a Public Health Emergency of International Concern on 30 January 2020, and a pandemic on 11 March 2020.
SARS‑CoV‑2 is a positive-sense single-stranded RNA virus (and hence Baltimore class IV) that is contagious in humans. As described by the US National Institutes of Health, it is the successor to SARS-CoV-1, the virus that caused the 2002–2004 SARS outbreak.
Taxonomically, SARS‑CoV‑2 is a virus of the species severe acute respiratory syndrome–related coronavirus (SARSr-CoV). It is believed to have zoonotic origins and has close genetic similarity to bat coronaviruses, suggesting it emerged from a bat-borne virus. Research is ongoing as to whether SARS‑CoV‑2 came directly from bats or indirectly through any intermediate hosts. The virus shows little genetic diversity, indicating that the spillover event introducing SARS‑CoV‑2 to humans is likely to have occurred in late 2019.
Epidemiological studies estimate that each infection results in an average of 2.39 to 3.44 new ones when no members of the community are immune and no preventive measures are taken. The virus primarily spreads between people through close contact and via respiratory droplets produced from coughs or sneezes. It mainly enters human cells by binding to the angiotensin converting enzyme 2 (ACE2).
Terminology

During the initial outbreak in Wuhan, China, various names were used for the virus; some names used by different sources included the "coronavirus" or "Wuhan coronavirus". In January 2020, the World Health Organisation recommended "2019 novel coronavirus" (2019-nCov) as the provisional name for the virus. This was in accordance with WHO's 2015 guidance against using geographical locations, animal species, or groups of people in disease and virus names.
On 11 February 2020, the International Committee on Taxonomy of Viruses adopted the official name "severe acute respiratory syndrome coronavirus 2" (SARS‑CoV‑2). To avoid confusion with the disease SARS, the WHO sometimes refers to SARS‑CoV‑2 as "the COVID-19 virus" in public health communications and the name HCoV-19 was included in some research articles.
The general public often calls both the virus and the disease it causes, "the coronavirus". U.S. President Donald Trump repeatedly referred to the virus as the "Chinese virus" in tweets, interviews, and White House press briefings, which drew some criticism that he was stigmatizing the disease with racial or nationalistic overtones.
Virology
Infection and transmission
Main article: Transmission of COVID-19Human-to-human transmission of SARS‑CoV‑2 was confirmed on 20 January 2020, during the COVID-19 pandemic. Transmission was initially assumed to occur primarily via respiratory droplets from coughs and sneezes within a range of about 1.8 metres (6 ft). Laser light scattering experiments suggest that speaking is an additional mode of transmission and a far-reaching and under-researched one, indoors, with little air flow. Other studies have suggested that the virus may be airborne as well, with aerosols potentially being able to transmit the virus. During human-to-human transmission, an average 1000 infectious SARS‑CoV‑2 virions are thought to initiate a new infection.
Indirect contact via contaminated surfaces is another possible cause of infection. Preliminary research indicates that the virus may remain viable on plastic (polypropylene) and stainless steel (AISI 304) for up to three days, but does not survive on cardboard for more than one day or on copper for more than four hours; the virus is inactivated by soap, which destabilises its lipid bilayer. Viral RNA has also been found in stool samples and semen from infected individuals.
The degree to which the virus is infectious during the incubation period is uncertain, but research has indicated that the pharynx reaches peak viral load approximately four days after infection or the first week of symptoms, and declines after.
A study by a team of researchers from the University of North Carolina found that the nasal cavity is seemingly the dominant initial site for infection with subsequent aspiration-mediated virus seeding into the lungs in SARS‑CoV‑2 pathogenesis. They found that there was an infection gradient from high in proximal towards low in distal pulmonary epithelial cultures, with a focal infection in ciliated cells and type 2 pneumocytes in the airway and alveolar regions respectively.
There is some evidence of human-to-animal transmission of SARS‑CoV‑2, including examples in felids. Some institutions have advised those infected with SARS‑CoV‑2 to restrict contact with animals.
Asymptomatic transmission
On 1 February 2020, the World Health Organization (WHO) indicated that "transmission from asymptomatic cases is likely not a major driver of transmission". One meta-analysis found that 17% of infections are asymptomatic, and asymptomatic individuals were 42% less likely to transmit the virus.
However, an epidemiological model of the beginning of the outbreak in China suggested that "pre-symptomatic shedding may be typical among documented infections" and that subclinical infections may have been the source of a majority of infections. That may explain how out of 217 onboard a cruise liner that docked at Montevideo, only 24 of 128 who tested positive for viral RNA showed symptoms. Similarly, a study of ninety-four patients hospitalized in January and February 2020 estimated patients shed the greatest amount of virus two to three days before symptoms appear and that "a substantial proportion of transmission probably occurred before first symptoms in the index case".
Reinfection
There is uncertainty about reinfection and long-term immunity. It is not known how common reinfection is, but reports have indicated that it is occurring with variable severity.
The first reported case of reinfection was a 33-year-old man from Hong Kong who first tested positive on 26 March 2020, was discharged on 15 April 2020 after two negative tests, and tested positive again on 15 August 2020 (142 days later), which was confirmed by whole-genome sequencing showing that the viral genomes between the episodes belong to different clades. The findings had the implications that herd immunity may not eliminate the virus if reinfection is not an uncommon occurrence and that vaccines may not be able to provide lifelong protection against the virus.
Another case study described a 25-year-old man from Nevada who tested positive for SARS‑CoV‑2 on 18 April 2020 and on 5 June 2020 (separated by two negative tests). Since genomic analyses showed significant genetic differences between the SARS‑CoV‑2 variant sampled on those two dates, the case study authors determined this was a reinfection. The man's second infection was symptomatically more severe than the first infection, but the mechanisms that could account for this are not known.
Reservoir and zoonotic origin

The first known infections from SARS‑CoV‑2 were discovered in Wuhan, China. The original source of viral transmission to humans remains unclear, as does whether the virus became pathogenic before or after the spillover event. Because many of the early infectees were workers at the Huanan Seafood Market, it has been suggested that the virus might have originated from the market. However, other research indicates that visitors may have introduced the virus to the market, which then facilitated rapid expansion of the infections. A March 2021 WHO report on a joint WHO-China study stated that human spillover via an intermediate animal host was the most likely explanation, with direct spillover from bats next most likely. Introduction through the food supply chain and the Huanan Seafood Market was considered another possible, but less likely, explanation.
The mutation rate estimated from early cases of SARS-CoV-2 was of 6.54 X 10 per site per year. A phylogenetic network analysis of 160 early coronavirus genomes sampled from December 2019 to February 2020 showed that the virus type most closely related to the bat coronavirus was most abundant in Guangdong, China, and designated type "A". The predominant type among samples from Wuhan, "B", is more distantly related to the bat coronavirus than the ancestral type "A".
Research into the natural reservoir of the virus that caused the 2002–2004 SARS outbreak has resulted in the discovery of many SARS-like bat coronaviruses, most originating in the Rhinolophus genus of horseshoe bats. Phylogenetic analysis indicates that samples taken from Rhinolophus sinicus show a resemblance of 80% to SARS‑CoV‑2. Phylogenetic analysis also indicates that a virus from Rhinolophus affinis, collected in Yunnan province and designated RaTG13, has a 96% resemblance to SARS‑CoV‑2. The RaTG13 virus sequence is the closest known sequence to SARS-CoV-2.

Bats are considered the most likely natural reservoir of SARS‑CoV‑2, but differences between the bat coronavirus and SARS‑CoV‑2 suggest that humans were infected via an intermediate host.
Although the role of pangolins as an intermediate host was initially posited (a study published in July 2020 suggested that pangolins are an intermediate host of SARS‑CoV‑2-like coronaviruses), subsequent studies have not substantiated their contribution to the spillover. Evidence against this hypothesis includes the fact that pangolin virus samples are too distant to SARS-CoV-2: isolates obtained from pangolins seized in Guangdong were only 92% identical in sequence to the SARS‑CoV‑2 genome. In addition, despite similarities in a few critical amino acids, pangolin virus samples exhibit poor binding to the human ACE2 receptor.
All available evidence suggests that SARS‑CoV‑2 has a natural animal origin and is not genetically engineered. Nevertheless, early in the pandemic, conspiracy theories spread on social media claiming that the virus was bio-engineered by China at the Wuhan Institute of Virology, amplified by echo chambers in the American far-right. A few individuals, including former CDC director Robert R. Redfield, have claimed, without evidence, that the virus may have been studied by and escaped from the Institute. Most virologists who have studied coronaviruses consider the possibility very remote, and the March 2021 WHO report on the joint WHO-China study stated that such an explanation is "extremely unlikely".
Phylogenetics and taxonomy
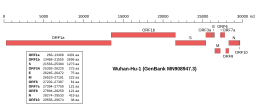 Genomic organisation of isolate Wuhan-Hu-1, the earliest sequenced sample of SARS-CoV-2 Genomic organisation of isolate Wuhan-Hu-1, the earliest sequenced sample of SARS-CoV-2 | |
| NCBI genome ID | 86693 |
|---|---|
| Genome size | 29,903 bases |
| Year of completion | 2020 |
| Genome browser (UCSC) | |
SARS‑CoV‑2 belongs to the broad family of viruses known as coronaviruses. It is a positive-sense single-stranded RNA (+ssRNA) virus, with a single linear RNA segment. Other coronaviruses are capable of causing illnesses ranging from the common cold to more severe diseases such as Middle East respiratory syndrome (MERS, fatality rate ~34%). It is the seventh known coronavirus to infect people, after 229E, NL63, OC43, HKU1, MERS-CoV, and the original SARS-CoV.
Like the SARS-related coronavirus implicated in the 2003 SARS outbreak, SARS‑CoV‑2 is a member of the subgenus Sarbecovirus (beta-CoV lineage B). Coronaviruses also undergo frequent recombination. Its RNA sequence is approximately 30,000 bases in length, relatively long for a coronavirus. SARS‑CoV‑2 is unique among known betacoronaviruses in its incorporation of a polybasic site cleaved by furin, a characteristic known to increase pathogenicity and transmissibility in other viruses.
With a sufficient number of sequenced genomes, it is possible to reconstruct a phylogenetic tree of the mutation history of a family of viruses. By 12 January 2020, five genomes of SARS‑CoV‑2 had been isolated from Wuhan and reported by the Chinese Center for Disease Control and Prevention (CCDC) and other institutions; the number of genomes increased to 42 by 30 January 2020. A phylogenetic analysis of those samples showed they were "highly related with at most seven mutations relative to a common ancestor", implying that the first human infection occurred in November or December 2019. As of 7 May 2020, 4,690 SARS‑CoV‑2 genomes sampled on six continents were publicly available.
On 11 February 2020, the International Committee on Taxonomy of Viruses announced that according to existing rules that compute hierarchical relationships among coronaviruses based on five conserved sequences of nucleic acids, the differences between what was then called 2019-nCoV and the virus from the 2003 SARS outbreak were insufficient to make them separate viral species. Therefore, they identified 2019-nCoV as a virus of Severe acute respiratory syndrome–related coronavirus.
In July 2020, scientists reported that a more infectious SARS‑CoV‑2 variant with spike protein variant G614 has replaced D614 as the dominant form in the pandemic. In October 2020 scientists reported in a preprint that a variant, 20A.EU1, was first observed in Spain in early summer and has become the most frequent variant in multiple European countries. They also illustrate the emergence and spread of other frequent clusters of sequences using Nextstrain.
In October 2020, researchers discovered a possible overlapping gene named ORF3d, in the SARS‑CoV‑2 genome. It is unknown if the protein produced by ORF3d has any function, but it provokes a strong immune response. ORF3d has been identified before, in a variant of coronavirus that infects pangolins.
Variants
Main article: Variants of SARS-CoV-2
There are many thousands of variants of SARS-CoV-2, which can be grouped into the much larger clades. Several different clade nomenclatures have been proposed. Nextstrain divides the variants into five clades (19A, 19B, 20A, 20B, and 20C), while GISAID divides them into seven (L, O, V, S, G, GH, and GR).
Several notable variants of SARS-CoV-2 emerged in late 2020.
- Lineage B.1.1.7 (formerly known as Variant of Concern 202012/01 (VOC 202012/01)) is believed to have emerged in the United Kingdom in September 2020. Epidemiological markers suggest that the variant is more transmissible and lethal. Among the variant's several mutations is one in the receptor-binding domain of the spike protein that changes the asparagine at position 501 to tyrosine (N501Y). This mutation may cause the virus to bind more tightly to the ACE2 receptor. It is currently spread globally.
- The 501.V2 variant, which has the same N501Y mutation, arose independently in South Africa. It was detected in patient specimens collected at the beginning of October 2020 and currently spread globally.
- The B.1.207 variant appeared in Nigeria. It has a mutation in the spike protein (P681H) that is also found in the VOC 202012/01 variant. P681H is located near the S1/S2 furin cleavage site. There is no evidence that the mutations increase the transmissibility of the variant.
- The Lineage B.1.525 first cases were detected in December 2020 in the United Kingdom and Nigeria. It is currently spread globally.
- The Cluster 5 variant emerged among minks and mink farmers in Denmark. It has a set of mutations that have not been observed in other variants, including four amino acid changes in the spike protein. The variant moderately resists neutralizing antibodies. After strict quarantines, a ban on mink farming, and a mink euthanasia campaign, it is believed to have been eradicated.
- Lineage P.1 first detected in Manaus, Brazil and already spread globally, the preprint works showed the variant to be more transmissible and have a higher rate of deaths than B.1.1.28 and B.1.195 lineages.
Structural biology

Each SARS-CoV-2 virion is 50–200 nanometres in diameter. Like other coronaviruses, SARS-CoV-2 has four structural proteins, known as the S (spike), E (envelope), M (membrane), and N (nucleocapsid) proteins; the N protein holds the RNA genome, and the S, E, and M proteins together create the viral envelope. The spike protein, which has been imaged at the atomic level using cryogenic electron microscopy, is the protein responsible for allowing the virus to attach to and fuse with the membrane of a host cell; specifically, its S1 subunit catalyzes attachment, the S2 subunit fusion.
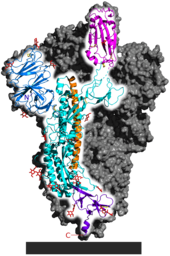
Protein modeling experiments on the spike protein of the virus soon suggested that SARS‑CoV‑2 has sufficient affinity to the receptor angiotensin converting enzyme 2 (ACE2) on human cells to use them as a mechanism of cell entry. By 22 January 2020, a group in China working with the full virus genome and a group in the United States using reverse genetics methods independently and experimentally demonstrated that ACE2 could act as the receptor for SARS‑CoV‑2. Studies have shown that SARS‑CoV‑2 has a higher affinity to human ACE2 than the original SARS virus. SARS‑CoV‑2 may also use basigin to assist in cell entry.
Initial spike protein priming by transmembrane protease, serine 2 (TMPRSS2) is essential for entry of SARS‑CoV‑2. The host protein neuropilin 1 (NRP1) may aid the virus in host cell entry using ACE2. After a SARS‑CoV‑2 virion attaches to a target cell, the cell's protease TMPRSS2 cuts open the spike protein of the virus, exposing a fusion peptide in the S2 subunit, and the host receptor ACE2. After fusion, an endosome forms around the virion, separating it from the rest of the host cell. The virion escapes when the pH of the endosome drops or when cathepsin, a host cysteine protease, cleaves it. The virion then releases RNA into the cell and forces the cell to produce and disseminate copies of the virus, which infect more cells.
SARS‑CoV‑2 produces at least three virulence factors that promote shedding of new virions from host cells and inhibit immune response. Whether they include downregulation of ACE2, as seen in similar coronaviruses, remains under investigation (as of May 2020).
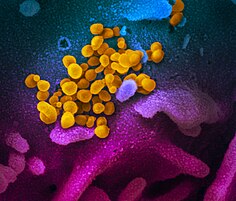
 Digitally colourised scanning electron micrographs of SARS-CoV-2 virions (yellow) emerging from human cells cultured in a laboratory
Digitally colourised scanning electron micrographs of SARS-CoV-2 virions (yellow) emerging from human cells cultured in a laboratory
Epidemiology
Main article: COVID-19 pandemic
Based on the low variability exhibited among known SARS‑CoV‑2 genomic sequences, health authorities likely detected the virus within weeks of its emergence among the human population in late 2019. The earliest case of infection currently known is dated to 1 December 2019, although an earlier case could have occurred on 17 November 2019. The pandemic onset was estimated by tMRCA analysis to have ocurred before the end of December 2019, but this statistical inference do not provide definitiv proof of time of origins. The virus subsequently spread to all provinces of China and to more than 150 other countries across the world. Human-to-human transmission of the virus has been confirmed in all these regions. On 30 January 2020, SARS‑CoV‑2 was designated a Public Health Emergency of International Concern by the WHO, and on 11 March 2020 the WHO declared it a pandemic.
Retrospective tests collected within the Chinese surveillance system revealed no clear indication of substantial unrecognized circulation of SARS‑CoV‑2 in Wuhan during the latter part of 2019.
The basic reproduction number () of the virus has been estimated to be around 5.7. This means each infection from the virus is expected to result in 5.7 new infections when no members of the community are immune and no preventive measures are taken. The reproduction number may be higher in densely populated conditions such as those found on cruise ships. Many forms of preventive efforts may be employed in specific circumstances to reduce the propagation of the virus.
There have been about 96,000 confirmed cases of infection in mainland China. While the proportion of infections that result in confirmed cases or progress to diagnosable disease remains unclear, one mathematical model estimated that 75,815 people were infected on 25 January 2020 in Wuhan alone, at a time when the number of confirmed cases worldwide was only 2,015. Before 24 February 2020, over 95% of all deaths from COVID-19 worldwide had occurred in Hubei province, where Wuhan is located. As of 10 March 2023, the percentage had decreased to 0.047%.
As of 10 March 2023, there have been 676,609,955 total confirmed cases of SARS‑CoV‑2 infection in the ongoing pandemic. The total number of deaths attributed to the virus is 6,881,955. Many recoveries from both confirmed and untested infections go unreported, since some countries do not collect this data, but at least people have recovered from confirmed infections.
References
- ^ Giaimo C (1 April 2020). "The Spiky Blob Seen Around the World". The New York Times. Archived from the original on 2 April 2020. Retrieved 6 April 2020.
- ^ Gorbalenya AE, Baker SC, Baric RS, de Groot RJ, Drosten C, Gulyaeva AA, et al. (March 2020). "The species Severe acute respiratory syndrome-related coronavirus: classifying 2019-nCoV and naming it SARS-CoV-2". Nature Microbiology. 5 (4): 536–544. doi:10.1038/s41564-020-0695-z. PMC 7095448. PMID 32123347.
- "Coronavirus disease named Covid-19". BBC News Online. 11 February 2020. Archived from the original on 15 February 2020. Retrieved 15 February 2020.
- Zimmer C (26 February 2021). "The Secret Life of a Coronavirus - An oily, 100-nanometer-wide bubble of genes has killed more than two million people and reshaped the world. Scientists don't quite know what to make of it". Retrieved 28 February 2021.
- Surveillance case definitions for human infection with novel coronavirus (nCoV): interim guidance v1, January 2020 (Report). World Health Organization. January 2020. hdl:10665/330376. WHO/2019-nCoV/Surveillance/v2020.1.
- ^ "Healthcare Professionals: Frequently Asked Questions and Answers". United States Centers for Disease Control and Prevention (CDC). 11 February 2020. Archived from the original on 14 February 2020. Retrieved 15 February 2020.
- "About Novel Coronavirus (2019-nCoV)". United States Centers for Disease Control and Prevention (CDC). 11 February 2020. Archived from the original on 11 February 2020. Retrieved 25 February 2020.
- Harmon A (4 March 2020). "We Spoke to Six Americans with Coronavirus". The New York Times. Archived from the original on 13 March 2020. Retrieved 16 March 2020.
- ^ Wong G, Bi YH, Wang QH, Chen XW, Zhang ZG, Yao YG (May 2020). "Zoonotic origins of human coronavirus 2019 (HCoV-19 / SARS-CoV-2): why is this work important?". Zoological Research. 41 (3): 213–219. doi:10.24272/j.issn.2095-8137.2020.031. PMC 7231470. PMID 32314559.
- ^ Andersen KG, Rambaut A, Lipkin WI, Holmes EC, Garry RF (17 March 2020). "Correspondence: The proximal origin of SARS-CoV-2". Nature Medicine. 26 (4): 450–452. doi:10.1038/s41591-020-0820-9. PMC 7095063. PMID 32284615.
- ^ van Doremalen N, Bushmaker T, Morris DH, Holbrook MG, Gamble A, Williamson BN, et al. (April 2020). "Aerosol and Surface Stability of SARS-CoV-2 as Compared with SARS-CoV-1". The New England Journal of Medicine. 382 (16): 1564–1567. doi:10.1056/NEJMc2004973. PMC 7121658. PMID 32182409.
- "hCoV-19 Database". China National GeneBank. Archived from the original on 17 June 2020. Retrieved 2 June 2020.
- ^ "Statement on the second meeting of the International Health Regulations (2005) Emergency Committee regarding the outbreak of novel coronavirus (2019-nCoV)". World Health Organization (WHO) (Press release). 30 January 2020. Archived from the original on 31 January 2020. Retrieved 30 January 2020.
- ^ "WHO Director-General's opening remarks at the media briefing on COVID-19 – 11 March 2020". World Health Organization (WHO) (Press release). 11 March 2020. Archived from the original on 11 March 2020. Retrieved 12 March 2020.
- ^ "CoV2020". GISAID EpifluDB. Archived from the original on 12 January 2020. Retrieved 12 January 2020.
- Machhi J, Herskovitz J, Senan AM, Dutta D, Nath B, Oleynikov MD, et al. (September 2020). "The Natural History, Pathobiology, and Clinical Manifestations of SARS-CoV-2 Infections". Journal of Neuroimmune Pharmacology. 15 (3): 359–386. doi:10.1007/s11481-020-09944-5. PMC 7373339. PMID 32696264.
- Sills ES, Wood SH (10 November 2020). "An Experimental Model for Peri-conceptual COVID-19 Pregnancy Loss and Proposed Interventions to Optimize Outcomes". International Journal of Molecular and Cellular Medicine. 9 (3): 180–187. doi:10.22088/IJMCM.BUMS.9.3.180. PMC 7703664. PMID 33274180..
For Baltimore's original paper, see Baltimore D (September 1971). "Expression of animal virus genomes". Bacteriological Reviews. 35 (3): 235–41. doi:10.1128/MMBR.35.3.235-241.1971. PMC 378387. PMID 4329869. - ^ Chan JF, Yuan S, Kok KH, To KK, Chu H, Yang J, et al. (February 2020). "A familial cluster of pneumonia associated with the 2019 novel coronavirus indicating person-to-person transmission: a study of a family cluster". The Lancet. 395 (10223): 514–523. doi:10.1016/S0140-6736(20)30154-9. PMC 7159286. PMID 31986261.
- "New coronavirus stable for hours on surfaces". National Institutes of Health (NIH). NIH.gov. 17 March 2020. Archived from the original on 23 March 2020. Retrieved 4 May 2020.
- ^ Zhou P, Yang XL, Wang XG, Hu B, Zhang L, Zhang W, et al. (February 2020). "A pneumonia outbreak associated with a new coronavirus of probable bat origin". Nature. 579 (7798): 270–273. Bibcode:2020Natur.579..270Z. doi:10.1038/s41586-020-2012-7. PMC 7095418. PMID 32015507.
- Perlman S (February 2020). "Another Decade, Another Coronavirus". The New England Journal of Medicine. 382 (8): 760–762. doi:10.1056/NEJMe2001126. PMC 7121143. PMID 31978944.
- ^ Benvenuto D, Giovanetti M, Ciccozzi A, Spoto S, Angeletti S, Ciccozzi M (April 2020). "The 2019-new coronavirus epidemic: Evidence for virus evolution". Journal of Medical Virology. 92 (4): 455–459. doi:10.1002/jmv.25688. PMC 7166400. PMID 31994738.
- Novel Coronavirus (2019-nCoV): situation report, 22 (Report). World Health Organization. 11 February 2020. hdl:10665/330991.
- Shield C (7 February 2020). "Coronavirus: From bats to pangolins, how do viruses reach us?". Deutsche Welle. Archived from the original on 4 June 2020. Retrieved 13 March 2020.
- ^ Cohen J (January 2020). "Wuhan seafood market may not be source of novel virus spreading globally". Science. doi:10.1126/science.abb0611.
- Billah MA, Miah MM, Khan MN (11 November 2020). "Reproductive number of coronavirus: A systematic review and meta-analysis based on global level evidence". PLOS ONE. 15 (11): e0242128. Bibcode:2020PLoSO..1542128B. doi:10.1371/journal.pone.0242128. PMC 7657547. PMID 33175914.
{{cite journal}}: CS1 maint: unflagged free DOI (link) - "Q&A on coronaviruses (COVID-19)". World Health Organization (WHO). 11 February 2020. Archived from the original on 20 January 2020. Retrieved 24 February 2020.
- ^ "How COVID-19 Spreads". U.S. Centers for Disease Control and Prevention (CDC). 27 January 2020. Archived from the original on 28 January 2020. Retrieved 29 January 2020.
- ^ Letko M, Marzi A, Munster V (February 2020). "Functional assessment of cell entry and receptor usage for SARS-CoV-2 and other lineage B betacoronaviruses". Nature Microbiology. 5 (4): 562–569. doi:10.1038/s41564-020-0688-y. PMC 7095430. PMID 32094589.
- ^ Hoffmann M, Kleine-Weber H, Schroeder S, Krüger N, Herrler T, Erichsen S, et al. (April 2020). "SARS-CoV-2 Cell Entry Depends on ACE2 and TMPRSS2 and Is Blocked by a Clinically Proven Protease Inhibitor". Cell. 181 (2): 271–280.e8. doi:10.1016/j.cell.2020.02.052. PMC 7102627. PMID 32142651.
- Wu KJ (15 April 2020). "There are more viruses than stars in the universe. Why do only some infect us? – More than a quadrillion quadrillion individual viruses exist on Earth, but most are not poised to hop into humans. Can we find the ones that are?". National Geographic Society. Archived from the original on 23 April 2020. Retrieved 18 May 2020.
- Huang P (22 January 2020). "How Does Wuhan Coronavirus Compare with MERS, SARS and the Common Cold?". NPR. Archived from the original on 2 February 2020. Retrieved 3 February 2020.
- ^ Fox D (January 2020). "What you need to know about the novel coronavirus". Nature. doi:10.1038/d41586-020-00209-y. PMID 33483684.
- World Health Organization (30 January 2020). Novel Coronavirus (2019-nCoV): situation report, 10 (Report). World Health Organization. hdl:10665/330775.
- "World Health Organization Best Practices for the Naming of New Human Infectious Diseases" (PDF). WHO. May 2015. Archived (PDF) from the original on 12 February 2020.
- "Novel coronavirus named 'Covid-19': WHO". TODAYonline. Archived from the original on 21 March 2020. Retrieved 11 February 2020.
- "The coronavirus spreads racism against—and among—ethnic Chinese". The Economist. 17 February 2020. Archived from the original on 17 February 2020. Retrieved 17 February 2020.
- "Naming the coronavirus disease (COVID-2019) and the virus that causes it". World Health Organization. Archived from the original on 28 February 2020. Retrieved 14 December 2020.
ICTV announced "severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)" as the name of the new virus on 11 February 2020. This name was chosen because the virus is genetically related to the coronavirus responsible for the SARS outbreak of 2003. While related, the two viruses are different.
- Hui M (18 March 2020). "Why won't the WHO call the coronavirus by its name, SARS-CoV-2?". Quartz. Archived from the original on 25 March 2020. Retrieved 26 March 2020.
- "Naming the coronavirus disease (COVID-2019) and the virus that causes it". World Health Organization. Archived from the original on 28 February 2020. Retrieved 14 December 2020.
From a risk communications perspective, using the name SARS can have unintended consequences in terms of creating unnecessary fear for some populations. ... For that reason and others, WHO has begun referring to the virus as "the virus responsible for COVID-19" or "the COVID-19 virus" when communicating with the public. Neither of these designations is intended as replacements for the official name of the virus as agreed by the ICTV.
- News, A. B. C. "Trump's 'Chinese Virus' tweet helped lead to rise in racist anti-Asian Twitter content: Study". ABC News. Retrieved 7 April 2021.
{{cite news}}:|last1=has generic name (help) - "Trump urges Americans to get COVID vaccine in Fox News exclusive". Fox News. Retrieved 7 April 2021.
We were the envy of the world and then when we got hit by the, as I call it, 'the China virus' — COVID — it obviously went down along with every other economy.
- "President Trump Holds News Conference | C-SPAN.org". www.c-span.org. Retrieved 7 April 2021.
We had tremendous growth until we got hit with the China virus.
- Gstalter M (19 March 2020). "WHO official warns against calling it 'Chinese virus', says 'there is no blame in this'". The Hill. Archived from the original on 18 April 2020. Retrieved 21 March 2020.
- Shinkman P (17 March 2020). "Trump Fires Back at Complaints He's Stigmatizing China Over Coronavirus". US News. Archived from the original on 29 March 2020. Retrieved 21 March 2020.
- Steakin W (20 June 2020). "Trump heads to Tulsa for return rally amid pandemic, despite mounting warnings from health experts". Archived from the original on 20 June 2020. Retrieved 20 June 2020.
- Li JY, You Z, Wang Q, Zhou ZJ, Qiu Y, Luo R, Ge XY (March 2020). "The epidemic of 2019-novel-coronavirus (2019-nCoV) pneumonia and insights for emerging infectious diseases in the future". Microbes and Infection. 22 (2): 80–85. doi:10.1016/j.micinf.2020.02.002. PMC 7079563. PMID 32087334.
- Kessler G (17 April 2020). "Trump's false claim that the WHO said the coronavirus was 'not communicable'". The Washington Post. Archived from the original on 17 April 2020. Retrieved 17 April 2020.
- Kuo L (21 January 2020). "China confirms human-to-human transmission of coronavirus". The Guardian. Archived from the original on 22 March 2020. Retrieved 18 April 2020.
- Edwards E (25 January 2020). "How does coronavirus spread?". NBC News. Archived from the original on 28 January 2020. Retrieved 13 March 2020.
- Anfinrud P, Stadnytskyi V, Bax CE, Bax A (May 2020). "Visualizing Speech-Generated Oral Fluid Droplets with Laser Light Scattering". The New England Journal of Medicine. 382 (21): 2061–2063. doi:10.1056/NEJMc2007800. PMC 7179962. PMID 32294341.
- Stadnytskyi V, Bax CE, Bax A, Anfinrud P (June 2020). "The airborne lifetime of small speech droplets and their potential importance in SARS-CoV-2 transmission". Proceedings of the National Academy of Sciences of the United States of America. 117 (22): 11875–11877. doi:10.1073/pnas.2006874117. PMC 7275719. PMID 32404416.
- Klompas M, Baker MA, Rhee C (4 August 2020). "Airborne Transmission of SARS-CoV-2". JAMA. 324 (5): 441. doi:10.1001/jama.2020.12458. S2CID 220500293. Retrieved 23 January 2021.
Investigators have demonstrated that speaking and coughing produce a mixture of both droplets and aerosols in a range of sizes, that these secretions can travel together for up to 27 feet, that it is feasible for SARS-CoV-2 to remain suspended in the air and viable for hours, that SARS-CoV-2 RNA can be recovered from air samples in hospitals, and that poor ventilation prolongs the amount of time that aerosols remain airborne.
- Asadi S, Bouvier N, Wexler AS, Ristenpart WD (2 June 2020). "The coronavirus pandemic and aerosols: Does COVID-19 transmit via expiratory particles?". Aerosol Science and Technology. 54 (6): 635–638. Bibcode:2020AerST..54..635A. doi:10.1080/02786826.2020.1749229. PMC 7157964. PMID 32308568.
It is unclear which of these mechanisms plays a key role in transmission of COVID-19. Much airborne disease research prior to the current pandemic has focused on 'violent' expiratory events like sneezing and coughing
- Rettner R (21 January 2021). "Talking is worse than coughing for spreading COVID-19 indoors". livescience.com. Retrieved 23 January 2021.
In one modeled scenario, the researchers found that after a short cough, the number of infectious particles in the air would quickly fall after 1 to 7 minutes; in contrast, after speaking for 30 seconds, only after 30 minutes would the number of infectious particles fall to similar levels; and a high number of particles were still suspended after one hour. In other words, a dose of virus particles capable of causing an infection would linger in the air much longer after speech than a cough. (In this modeled scenario, the same number of droplets were admitted during a 0.5-second cough as during the course of 30 seconds of speech.)
{{cite web}}: CS1 maint: url-status (link) - de Oliveira PM, Mesquita LC, Gkantonas S, Giusti A, Mastorakos E (January 2021). "Evolution of spray and aerosol from respiratory releases: theoretical estimates for insight on viral transmission". Proceedings of the Royal Society A: Mathematical, Physical and Engineering Sciences. 477 (2245): 20200584. Bibcode:2021RSPSA.47700584D. doi:10.1098/rspa.2020.0584. PMC 7897643. PMID 33633490. S2CID 231643585.
- Mandavilli A (4 July 2020). "239 Experts With One Big Claim: The Coronavirus Is Airborne – The W.H.O. has resisted mounting evidence that viral particles floating indoors are infectious, some scientists say. The agency maintains the research is still inconclusive". The New York Times. Archived from the original on 17 November 2020. Retrieved 5 July 2020.
- Tufekci Z (30 July 2020). "We Need to Talk About Ventilation". The Atlantic. Archived from the original on 17 November 2020. Retrieved 8 September 2020.
- Lewis D (July 2020). "Mounting evidence suggests coronavirus is airborne - but health advice has not caught up". Nature. 583 (7817): 510–513. Bibcode:2020Natur.583..510L. doi:10.1038/d41586-020-02058-1. PMID 32647382. S2CID 220470431. Retrieved 9 October 2020.
{{cite journal}}: CS1 maint: url-status (link) - Popa A, Genger JW, Nicholson MD, Penz T, Schmid D, Aberle SW, et al. (December 2020). "Genomic epidemiology of superspreading events in Austria reveals mutational dynamics and transmission properties of SARS-CoV-2". Science Translational Medicine. 12 (573): eabe2555. doi:10.1126/scitranslmed.abe2555. PMC 7857414. PMID 33229462.
- Prentiss MG, Chu A, Berggren KK (23 October 2020). "Superspreading Events Without Superspreaders: Using High Attack Rate Events to Estimate Nº for Airborne Transmission of COVID-19". medRxiv. doi:10.1101/2020.10.21.20216895. S2CID 225040713. Retrieved 1 December 2020.
- "Getting your workplace ready for COVID-19" (PDF). World Health Organization. 27 February 2020. Archived (PDF) from the original on 2 March 2020. Retrieved 3 March 2020.
- Yong E (20 March 2020). "Why the Coronavirus Has Been So Successful". The Atlantic. Archived from the original on 20 March 2020. Retrieved 20 March 2020.
- Gibbens S (18 March 2020). "Why soap is preferable to bleach in the fight against coronavirus". National Geographic. Archived from the original on 2 April 2020. Retrieved 2 April 2020.
- Holshue ML, DeBolt C, Lindquist S, Lofy KH, Wiesman J, Bruce H, et al. (March 2020). "First Case of 2019 Novel Coronavirus in the United States". The New England Journal of Medicine. 382 (10): 929–936. doi:10.1056/NEJMoa2001191. PMC 7092802. PMID 32004427.
- Li D, Jin M, Bao P, Zhao W, Zhang S (May 2020). "Clinical Characteristics and Results of Semen Tests Among Men With Coronavirus Disease 2019". JAMA Network Open. 3 (5): e208292. doi:10.1001/jamanetworkopen.2020.8292. PMC 7206502. PMID 32379329.
- Wölfel R, Corman VM, Guggemos W, Seilmaier M, Zange S, Müller MA, et al. (April 2020). "Virological assessment of hospitalized patients with COVID-2019". Nature. 581 (7809): 465–469. Bibcode:2020Natur.581..465W. doi:10.1038/s41586-020-2196-x. PMID 32235945.
- Kupferschmidt K (February 2020). "Study claiming new coronavirus can be transmitted by people without symptoms was flawed". Science. doi:10.1126/science.abb1524.
- To KK, Tsang OT, Leung WS, Tam AR, Wu TC, Lung DC, et al. (May 2020). "Temporal profiles of viral load in posterior oropharyngeal saliva samples and serum antibody responses during infection by SARS-CoV-2: an observational cohort study". The Lancet. Infectious Diseases. 20 (5): 565–574. doi:10.1016/S1473-3099(20)30196-1. PMC 7158907. PMID 32213337. Archived from the original on 17 April 2020. Retrieved 21 April 2020.
- ^ Hou YJ, Okuda K, Edwards CE, Martinez DR, Asakura T, Dinnon KH, et al. (July 2020). "SARS-CoV-2 Reverse Genetics Reveals a Variable Infection Gradient in the Respiratory Tract". Cell. 182 (2): 429–446.e14. doi:10.1016/j.cell.2020.05.042. PMC 7250779. PMID 32526206.
- "Questions and Answers on the COVID-19: OIE – World Organisation for Animal Health". www.oie.int. Archived from the original on 31 March 2020. Retrieved 16 April 2020.
- Goldstein J (6 April 2020). "Bronx Zoo Tiger Is Sick with the Coronavirus". The New York Times. Archived from the original on 9 April 2020. Retrieved 10 April 2020.
- "USDA Statement on the Confirmation of COVID-19 in a Tiger in New York". United States Department of Agriculture. 5 April 2020. Archived from the original on 15 April 2020. Retrieved 16 April 2020.
- "If You Have Animals—Coronavirus Disease 2019 (COVID-19)". Centers for Disease Control and Prevention (CDC). 13 April 2020. Archived from the original on 1 April 2020. Retrieved 16 April 2020.
- World Health Organization (1 February 2020). Novel Coronavirus (2019-nCoV): situation report, 12 (Report). World Health Organization. hdl:10665/330777.
- Nogrady B (18 November 2020). "What the data say about asymptomatic COVID infections". Nature. 587 (7835): 534–535. Bibcode:2020Natur.587..534N. doi:10.1038/d41586-020-03141-3. PMID 33214725.
- Li R, Pei S, Chen B, Song Y, Zhang T, Yang W, et al. (16 March 2020). "Substantial undocumented infection facilitates the rapid dissemination of novel coronavirus (SARS-CoV2)". Science. 368 (6490): 489–493. Bibcode:2020Sci...368..489L. doi:10.1126/science.abb3221. PMC 7164387. PMID 32179701.
- Daily Telegraph, Thursday 28 May 2020, page 2 column 1, which refers to the medical journal Thorax; Thorax May 2020 article COVID-19: in the footsteps of Ernest Shackleton Archived 30 May 2020 at the Wayback Machine
- He X, Lau EH, Wu P, Deng X, Wang J, Hao X, et al. (May 2020). "Temporal dynamics in viral shedding and transmissibility of COVID-19". Nature Medicine. 26 (5): 672–675. doi:10.1038/s41591-020-0869-5. PMID 32296168. Retrieved 21 April 2020.
{{cite journal}}: CS1 maint: url-status (link) - ^ Ledford H (September 2020). "Coronavirus reinfections: three questions scientists are asking". Nature. 585 (7824): 168–169. doi:10.1038/d41586-020-02506-y. PMID 32887957. S2CID 221501940. Archived from the original on 17 November 2020. Retrieved 9 October 2020.
- ^ To KK, Hung IF, Ip JD, Chu AW, Chan WM, Tam AR, et al. (August 2020). "COVID-19 re-infection by a phylogenetically distinct SARS-coronavirus-2 strain confirmed by whole genome sequencing". Clinical Infectious Diseases: ciaa1275. doi:10.1093/cid/ciaa1275. PMC 7499500. PMID 32840608. S2CID 221308584.
- ^ Tillett RL, Sevinsky JR, Hartley PD, Kerwin H, Crawford N, Gorzalski A, et al. (January 2021). "Genomic evidence for reinfection with SARS-CoV-2: a case study". The Lancet. Infectious Diseases. 21 (1): 52–58. doi:10.1016/S1473-3099(20)30764-7. PMC 7550103. PMID 33058797.
- Eschner K (28 January 2020). "We're still not sure where the Wuhan coronavirus really came from". Popular Science. Archived from the original on 30 January 2020. Retrieved 30 January 2020.
- Huang C, Wang Y, Li X, Ren L, Zhao J, Hu Y, et al. (15 February 2020). "Clinical features of patients infected with 2019 novel coronavirus in Wuhan, China". The Lancet. 395 (10223): 497–506. doi:10.1016/S0140-6736(20)30183-5. PMC 7159299. PMID 31986264. Archived from the original on 31 January 2020. Retrieved 26 March 2020.
- ^ Chen N, Zhou M, Dong X, Qu J, Gong F, Han Y, et al. (15 February 2020). "Epidemiological and clinical characteristics of 99 cases of 2019 novel coronavirus pneumonia in Wuhan, China: a descriptive study". The Lancet. 395 (10223): 507–513. doi:10.1016/S0140-6736(20)30211-7. PMC 7135076. PMID 32007143. Archived from the original on 31 January 2020. Retrieved 9 March 2020.
- ^ Cyranoski D (26 February 2020). "Mystery deepens over animal source of coronavirus". Nature. 579 (7797): 18–19. Bibcode:2020Natur.579...18C. doi:10.1038/d41586-020-00548-w. PMID 32127703.
- Yu WB, Tang GD, Zhang L, Corlett RT (21 February 2020). "Decoding evolution and transmissions of novel pneumonia coronavirus using the whole genomic data". ChinaXiv. doi:10.12074/202002.00033 (inactive 5 January 2021). Archived from the original on 23 February 2020. Retrieved 25 February 2020.
{{cite journal}}: CS1 maint: DOI inactive as of January 2021 (link) - ^ "WHO-convened global study of origins of SARS-CoV-2: China Part". www.who.int. Retrieved 31 March 2021.
- Forster P, Forster L, Renfrew C, Forster M (8 April 2020). "Phylogenetic network analysis of SARS-CoV-2 genomes" (PDF). PNAS. 117 (17): 9241–9243. doi:10.1073/pnas.2004999117. PMC 7196762. PMID 32269081. Archived (PDF) from the original on 16 April 2020. Retrieved 17 April 2020.
- "COVID-19: genetic network analysis provides 'snapshot' of pandemic origins". Cambridge University. 9 April 2020. Archived from the original on 16 April 2020. Retrieved 17 April 2020.
- "Bat SARS-like coronavirus isolate bat-SL-CoVZC45, complete genome". National Center for Biotechnology Information (NCBI). 15 February 2020. Archived from the original on 4 June 2020. Retrieved 15 February 2020.
- "Bat SARS-like coronavirus isolate bat-SL-CoVZXC21, complete genome". National Center for Biotechnology Information (NCBI). 15 February 2020. Archived from the original on 4 June 2020. Retrieved 15 February 2020.
- "Bat coronavirus isolate RaTG13, complete genome". National Center for Biotechnology Information (NCBI). 10 February 2020. Archived from the original on 15 May 2020. Retrieved 5 March 2020.
- Report of the WHO-China Joint Mission on Coronavirus Disease 2019 (COVID-19) (PDF) (Report). World Health Organization (WHO). 24 February 2020. Archived (PDF) from the original on 29 February 2020. Retrieved 5 March 2020.
- Lu R, Zhao X, Li J, Niu P, Yang B, Wu H, et al. (February 2020). "Genomic characterisation and epidemiology of 2019 novel coronavirus: implications for virus origins and receptor binding". The Lancet. 395 (10224): 565–574. doi:10.1016/S0140-6736(20)30251-8. PMC 7159086. PMID 32007145.
- Xiao K, Zhai J, Feng Y, Zhou N, Zhang X, Zou JJ, Li N, Guo Y, Li X, Shen X, Zhang Z, Shu F, Huang W, Li Y, Zhang Z, Chen RA, Wu YJ, Peng SM, Huang M, Xie WJ, Cai QH, Hou FH, Chen W, Xiao L, Shen Y (July 2020). "Isolation of SARS-CoV-2-related coronavirus from Malayan pangolins". Nature. 583 (7815): 286–289. Bibcode:2020Natur.583..286X. doi:10.1038/s41586-020-2313-x. PMID 32380510. S2CID 218557880.
- Zhao J, Cui W, Tian BP (2020). "The Potential Intermediate Hosts for SARS-CoV-2". Frontiers in Microbiology. 11: 580137. doi:10.3389/fmicb.2020.580137. PMC 7554366. PMID 33101254.
{{cite journal}}: CS1 maint: unflagged free DOI (link) - Hu B, Guo H, Zhou P, Shi ZL (October 2020). "Characteristics of SARS-CoV-2 and COVID-19". Nature Reviews Microbiology. 19 (3): 141–154. doi:10.1038/s41579-020-00459-7. PMC 7537588. PMID 33024307.
- Giovanetti M, Benedetti F, Campisi G, Ciccozzi A, Fabris S, Ceccarelli G, Tambone V, Caruso A, Angeletti S, Zella D, Ciccozzi M (November 2020). "Evolution patterns of SARS-CoV-2: Snapshot on its genome variants". Biochemical and Biophysical Research Communications. 538: 88–91. doi:10.1016/j.bbrc.2020.10.102. PMC 7836704. PMID 33199021. S2CID 226988090.
- "Origin of SARS-CoV-2". www.who.int. Archived from the original on 17 November 2020. Retrieved 14 October 2020.
- Zoumpourlis V, Goulielmaki M, Rizos E, Baliou S, Spandidos DA (October 2020). "The COVID-19 pandemic as a scientific and social challenge in the 21st century". Mol Med Rep (Review). 22 (4): 3035–3048. doi:10.3892/mmr.2020.11393. PMC 7453598. PMID 32945405.
- Qin, Amy; Wang, Vivian; Hakim, Danny (20 November 2020). "How Steve Bannon and a Chinese Billionaire Created a Right-Wing Coronavirus Media Sensation". The New York Times. Archived from the original on 30 April 2021.
- ^ Gorman, James; Barnes, Julian E. (26 March 2021). "The C.D.C.'s ex-director offers no evidence in favoring speculation that the coronavirus originated in a lab". The New York Times. ISSN 0362-4331. Retrieved 29 April 2021.
- Hakim, Mohamad S. (14 February 2021). "SARS-CoV-2, Covid-19, and the debunking of conspiracy theories". Reviews in Medical Virology: e2222. doi:10.1002/rmv.2222. ISSN 1099-1654. PMC 7995093. PMID 33586302.
- Graham, Rachel L.; Baric, Ralph S. (19 May 2020). "SARS-CoV-2: Combating Coronavirus Emergence". Immunity. 52 (5): 734–736. doi:10.1016/j.immuni.2020.04.016. ISSN 1074-7613. PMC 7207110. PMID 32392464.
- Zhu N, Zhang D, Wang W, Li X, Yang B, Song J, et al. (February 2020). "A Novel Coronavirus from Patients with Pneumonia in China, 2019". The New England Journal of Medicine. 382 (8): 727–733. doi:10.1056/NEJMoa2001017. PMC 7092803. PMID 31978945.
- "Phylogeny of SARS-like betacoronaviruses". nextstrain. Archived from the original on 20 January 2020. Retrieved 18 January 2020.
- Wong AC, Li X, Lau SK, Woo PC (February 2019). "Global Epidemiology of Bat Coronaviruses". Viruses. 11 (2): 174. doi:10.3390/v11020174. PMC 6409556. PMID 30791586.
{{cite journal}}: CS1 maint: unflagged free DOI (link) - Singh, Devika; Yi, Soojin V. (16 April 2021). "On the origin and evolution of SARS-CoV-2". Experimental & Molecular Medicine. doi:10.1038/s12276-021-00604-z.
- Andersen KG, Rambaut A, Lipkin WI, Holmes EC, Garry RF (April 2020). "The proximal origin of SARS-CoV-2". Nature Medicine. 26 (4): 450–452. doi:10.1038/s41591-020-0820-9. PMC 7095063. PMID 32284615.
- Walls AC, Park YJ, Tortorici MA, Wall A, McGuire AT, Veesler D (9 March 2020). "Structure, function and antigenicity of the SARS-CoV-2 spike glycoprotein". Cell. 181 (2): 281–292.e6. doi:10.1016/j.cell.2020.02.058. PMC 7102599. PMID 32155444.
- Coutard B, Valle C, de Lamballerie X, Canard B, Seidah NG, Decroly E (February 2020). "The spike glycoprotein of the new coronavirus 2019-nCoV contains a furin-like cleavage site absent in CoV of the same clade". Antiviral Research. 176: 104742. doi:10.1016/j.antiviral.2020.104742. PMC 7114094. PMID 32057769.
- "Initial genome release of novel coronavirus". Virological. 11 January 2020. Archived from the original on 12 January 2020. Retrieved 12 January 2020.
- ^ Bedford T, Neher R, Hadfield N, Hodcroft E, Ilcisin M, Müller N. "Genomic analysis of nCoV spread: Situation report 2020-01-30". nextstrain.org. Archived from the original on 15 March 2020. Retrieved 18 March 2020.
- "Genomic epidemiology of novel coronavirus - Global subsampling". Nextstrain. Archived from the original on 20 April 2020. Retrieved 7 May 2020.
- Gorbalenya AE, Baker SC, Baric RS, de Groot RJ, Drosten C, Gulyaeva AA, et al. (March 2020). "The species Severe acute respiratory syndrome-related coronavirus: classifying 2019-nCoV and naming it SARS-CoV-2". Nature Microbiology. 5 (4): 536–544. doi:10.1038/s41564-020-0695-z. PMC 7095448. PMID 32123347.
- "New, more infectious strain of COVID-19 now dominates global cases of virus: study". medicalxpress.com. Archived from the original on 17 November 2020. Retrieved 16 August 2020.
- Korber B, Fischer WM, Gnanakaran S, Yoon H, Theiler J, Abfalterer W, et al. (August 2020). "Tracking Changes in SARS-CoV-2 Spike: Evidence that D614G Increases Infectivity of the COVID-19 Virus". Cell. 182 (4): 812–827.e19. doi:10.1016/j.cell.2020.06.043. PMC 7332439. PMID 32697968.
- Meredith S (29 October 2020). "A new coronavirus variant is seen spreading across Europe, research says". CNBC. Retrieved 10 November 2020.
- Hodcroft EB, Zuber M, Nadeau S, Crawford KH, Bloom JD, Veesler D, et al. (November 2020). "Emergence and spread of a SARS-CoV-2 variant through Europe in the summer of 2020". MedRxiv: 2020.10.25.20219063. doi:10.1101/2020.10.25.20219063. PMC 7709189. PMID 33269368.
- Dockrill P (11 November 2020). "Scientists Just Found a Mysteriously Hidden 'Gene Within a Gene' in SARS-CoV-2". ScienceAlert. Archived from the original on 17 November 2020. Retrieved 11 November 2020.
- Nelson CW, et al. (1 October 2020). "Dynamically evolving novel overlapping gene as a factor in the SARS-CoV-2 pandemic". eLife. 9. doi:10.7554/eLife.59633. PMC 7655111. PMID 33001029. Archived from the original on 17 November 2020. Retrieved 11 November 2020.
{{cite journal}}: CS1 maint: unflagged free DOI (link) - Koyama T, Platt D, Parida L (July 2020). "Variant analysis of SARS-CoV-2 genomes". Bulletin of the World Health Organization. 98 (7): 495–504. doi:10.2471/BLT.20.253591. PMC 7375210. PMID 32742035.
We detected in total 65776 variants with 5775 distinct variants.
- Alm E, Broberg EK, Connor T, Hodcroft EB, Komissarov AB, Maurer-Stroh S, Melidou A, Neher RA, O'Toole Á, Pereyaslov D (August 2020). "Geographical and temporal distribution of SARS-CoV-2 clades in the WHO European Region, January to June 2020". Euro Surveillance. 25 (32). doi:10.2807/1560-7917.ES.2020.25.32.2001410. PMC 7427299. PMID 32794443.
- "Emerging SARS-CoV-2 Variants". Centers for Disease Control and Prevention. 30 December 2020. Retrieved 30 December 2020.
- "SARS-CoV-2 mink-associated variant strain – Denmark". WHO. 3 December 2020. Retrieved 30 December 2020.
- ^ Wu C, Liu Y, Yang Y, Zhang P, Zhong W, Wang Y, et al. (February 2020). "Analysis of therapeutic targets for SARS-CoV-2 and discovery of potential drugs by computational methods". Acta Pharmaceutica Sinica B. 10 (5): 766–788. doi:10.1016/j.apsb.2020.02.008. PMC 7102550. PMID 32292689.
- ^ Wrapp D, Wang N, Corbett KS, Goldsmith JA, Hsieh CL, Abiona O, et al. (February 2020). "Cryo-EM structure of the 2019-nCoV spike in the prefusion conformation". Science. 367 (6483): 1260–1263. Bibcode:2020Sci...367.1260W. doi:10.1126/science.abb2507. PMC 7164637. PMID 32075877.
- Mandelbaum RF (19 February 2020). "Scientists Create Atomic-Level Image of the New Coronavirus's Potential Achilles Heel". Gizmodo. Archived from the original on 8 March 2020. Retrieved 13 March 2020.
- ^ Aronson JK (25 March 2020). "Coronaviruses – a general introduction". Centre for Evidence-Based Medicine, Nuffield Department of Primary Care Health Sciences, University of Oxford. Archived from the original on 22 May 2020. Retrieved 24 May 2020.
- Xu X, Chen P, Wang J, Feng J, Zhou H, Li X, et al. (March 2020). "Evolution of the novel coronavirus from the ongoing Wuhan outbreak and modeling of its spike protein for risk of human transmission". Science China. Life Sciences. 63 (3): 457–460. doi:10.1007/s11427-020-1637-5. PMC 7089049. PMID 32009228.
- Letko M, Munster V (January 2020). "Functional assessment of cell entry and receptor usage for lineage B β-coronaviruses, including 2019-nCoV" (PDF). bioRxiv (preprint). doi:10.1101/2020.01.22.915660. PMC 7217099. PMID 32511294. Archived (PDF) from the original on 22 April 2020. Retrieved 5 May 2020.
- El Sahly HM. "Genomic Characterization of the 2019 Novel Coronavirus". The New England Journal of Medicine. Archived from the original on 17 February 2020. Retrieved 9 February 2020.
- "Novel coronavirus structure reveals targets for vaccines and treatments". National Institutes of Health (NIH). 2 March 2020. Archived from the original on 1 April 2020. Retrieved 3 April 2020.
- Wang K, Chen W, Zhou YS, Lian JQ, Zhang Z, Du P, et al. (14 March 2020). "SARS-CoV-2 invades host cells via a novel route: CD147-spike protein" (PDF). bioRxiv (preprint). doi:10.1101/2020.03.14.988345. S2CID 214725955. Archived (PDF) from the original on 11 May 2020. Retrieved 5 May 2020.
- Zamorano Cuervo N, Grandvaux N (November 2020). "ACE2: Evidence of role as entry receptor for SARS-CoV-2 and implications in comorbidities". eLife. 9. doi:10.7554/eLife.61390. PMC 7652413. PMID 33164751.
{{cite journal}}: CS1 maint: unflagged free DOI (link) - "Anatomy of a Killer: Understanding SARS-CoV-2 and the drugs that might lessen its power". The Economist. 12 March 2020. Archived from the original on 14 March 2020. Retrieved 14 March 2020.
- Beeching NJ, Fletcher TE, Fowler R (22 May 2020). "BMJ Best Practice: Coronavirus Disease 2019 (COVID-19)" (PDF). BMJ. Archived (PDF) from the original on 13 June 2020. Retrieved 25 May 2020.
- Oberholzer M, Febbo P (19 February 2020). "What We Know Today about Coronavirus SARS-CoV-2 and Where Do We Go from Here". Genetic Engineering and Biotechnology News. Archived from the original on 14 March 2020. Retrieved 13 March 2020.
- The First 50 days of COVID-19: A Detailed Chronological Timeline and Extensive Review of Literature Documenting the Pandemic. Elsevier. 2020. ISBN 978-0-12-824313-8.
- Ma J (13 March 2020). "Coronavirus: China's first confirmed Covid-19 case traced back to November 17". South China Morning Post. Archived from the original on 13 March 2020. Retrieved 16 March 2020.
- ^ "COVID-19 Dashboard by the Center for Systems Science and Engineering (CSSE) at Johns Hopkins University (JHU)". ArcGIS. Johns Hopkins University. Retrieved 10 March 2023.
- Coronavirus disease 2019 (COVID-19) Situation Report – 69 (Report). World Health Organization. 29 March 2020. hdl:10665/331615.
- Wee SL, McNeil Jr. DG, Hernández JC (30 January 2020). "W.H.O. Declares Global Emergency as Wuhan Coronavirus Spreads". The New York Times. Archived from the original on 30 January 2020. Retrieved 30 January 2020.
- McKay B, Calfas J, Ansari T (11 March 2020). "Coronavirus Declared Pandemic by World Health Organization". The Wall Street Journal. Archived from the original on 11 March 2020. Retrieved 12 March 2020.
- "WHO-convened Global Study of Origins of SARS-CoV-2: China Part. Joint Report" (PDF). World Health Organization. World Health Organization. Retrieved 30 March 2021.
- Sanche S, Lin YT, Xu C, Romero-Severson E, Hengartner N, Ke R (July 2020). "High Contagiousness and Rapid Spread of Severe Acute Respiratory Syndrome Coronavirus 2". Emerging Infectious Diseases. 26 (7): 1470–1477. doi:10.3201/eid2607.200282. PMC 7323562. PMID 32255761.
- Rocklöv J, Sjödin H, Wilder-Smith A (February 2020). "COVID-19 outbreak on the Diamond Princess cruise ship: estimating the epidemic potential and effectiveness of public health countermeasures". Journal of Travel Medicine. 27 (3). doi:10.1093/jtm/taaa030. PMC 7107563. PMID 32109273.
- Dhama K, Khan S, Tiwari R, Sircar S, Bhat S, Malik YS, et al. (September 2020). "Coronavirus Disease 2019-COVID-19". Clinical Microbiology Reviews. 33 (4). doi:10.1128/CMR.00028-20. PMC 7405836. PMID 32580969.
- Branswell H (30 January 2020). "Limited data on coronavirus may be skewing assumptions about severity". STAT. Archived from the original on 1 February 2020. Retrieved 13 March 2020.
- Wu JT, Leung K, Leung GM (February 2020). "Nowcasting and forecasting the potential domestic and international spread of the 2019-nCoV outbreak originating in Wuhan, China: a modelling study". The Lancet. 395 (10225): 689–697. doi:10.1016/S0140-6736(20)30260-9. PMC 7159271. PMID 32014114.
- Boseley S, McCurry J (30 January 2020). "Coronavirus deaths leap in China as countries struggle to evacuate citizens". The Guardian. Archived from the original on 6 February 2020. Retrieved 10 March 2020.
- Paulinus A (25 February 2020). "Coronavirus: China to repay Africa in safeguarding public health". The Sun. Archived from the original on 9 March 2020. Retrieved 10 March 2020.
Further reading
- Bar-On YM, Flamholz A, Phillips R, Milo R (31 March 2020). "SARS-CoV-2 (COVID-19) by the numbers". eLife. 9. arXiv:2003.12886. Bibcode:2020arXiv200312886B. doi:10.7554/eLife.57309. PMC 7224694. PMID 32228860.
{{cite journal}}: CS1 maint: unflagged free DOI (link) - Brüssow H (March 2020). "The Novel Coronavirus – A Snapshot of Current Knowledge". Microbial Biotechnology. 2020 (3): 607–612. doi:10.1111/1751-7915.13557. PMC 7111068. PMID 32144890.
- Cascella M, Rajnik M, Cuomo A, Dulebohn SC, Di Napoli R (January 2020). "Features, Evaluation and Treatment Coronavirus (COVID-19)". StatPearls. PMID 32150360. Archived from the original on 6 April 2020. Retrieved 4 April 2020.
- Laboratory testing for coronavirus disease 2019 (COVID-19) in suspected human cases (Report). World Health Organization. 2 March 2020. hdl:10665/331329.
- Zoumpourlis V, Goulielmaki M, Rizos E, Baliou S, Spandidos DA (October 2020). "The COVID-19 pandemic as a scientific and social challenge in the 21st century". Mol Med Rep (Review). 22 (4): 3035–3048. doi:10.3892/mmr.2020.11393. PMC 7453598. PMID 32945405.
External links
- "Coronavirus Disease 2019 (COVID-19)". Centers for Disease Control and Prevention (CDC). 11 February 2020.
- "Coronavirus disease (COVID-19) Pandemic". World Health Organization (WHO).
- "SARS-CoV-2 (Severe acute respiratory syndrome coronavirus 2) Sequences". National Center for Biotechnology Information (NCBI).
- "COVID-19 Resource Centre". The Lancet.
- "Coronavirus (Covid-19)". The New England Journal of Medicine.
- "Covid-19: Novel Coronavirus Outbreak". Wiley.
- "SARS-CoV-2". Virus Pathogen Database and Analysis Resource.
- "SARS-CoV-2 related protein structures". Protein Data Bank.
| Classification | D |
|---|
| Taxonomy of the Coronaviridae | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Higher taxonomy: Riboviria > Orthornavirae > Pisuviricota > Pisoniviricetes > Nidovirales > Cornidovirineae > Coronaviridae | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Source: ICTV –– | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Infectious diseases – viral systemic diseases | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Oncovirus | |||||||||
| Immune disorders | |||||||||
| Central nervous system |
| ||||||||
| Cardiovascular | |||||||||
| Respiratory system/ acute viral nasopharyngitis/ viral pneumonia |
| ||||||||
| Human digestive system |
| ||||||||
| Urogenital | |||||||||
| Zoonotic viral diseases (A80–B34, 042–079) | |||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Arthropod -borne |
| ||||||||||||||||||||||||||||||||
| Mammal -borne |
| ||||||||||||||||||||||||||||||||
 Definitions from Wiktionary
Definitions from Wiktionary Media from Commons
Media from Commons News from Wikinews
News from Wikinews Quotations from Wikiquote
Quotations from Wikiquote Texts from Wikisource
Texts from Wikisource Textbooks from Wikibooks
Textbooks from Wikibooks Resources from Wikiversity
Resources from Wikiversity Travel guides from Wikivoyage
Travel guides from Wikivoyage Taxa from Wikispecies
Taxa from Wikispecies Data from Wikidata
Data from Wikidata
| Taxon identifiers | |
|---|---|
| Severe acute respiratory syndrome-related coronavirus | |
 ) of the virus has been estimated to be around 5.7. This means each infection from the virus is expected to result in 5.7 new infections when no members of the community are
) of the virus has been estimated to be around 5.7. This means each infection from the virus is expected to result in 5.7 new infections when no members of the community are