| Coiled-coil domain-containing protein 142 | |||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Identifiers | |||||||||||||||||||||||||||||||||||||||||||||||||||
| Aliases | CCDC142IPR026700Coiled-Coil Domain Containing 142 | ||||||||||||||||||||||||||||||||||||||||||||||||||
| External IDs | GeneCards: ; OMA:- orthologs | ||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| Wikidata | |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
The coiled-coil domain containing 142 (CCDC142) is a gene which in humans encodes the CCDC142 protein. The CCDC142 gene is located on chromosome 2 (at 2p13), spans 4339 base pairs and contains 9 exons. The gene codes for the coiled-coil domain containing protein 142 (CCDC142), whose function is not yet well understood. There are two known isoforms of CCDC142. CCDC142 proteins produced from these transcripts range in size from 743 to 665 amino acids and contain signals suggesting protein movement between the cytosol and nucleus. Homologous CCDC142 genes are found in many animals including vertebrates and invertebrates but not fungus, plants, protists, archea, or bacteria. Although the function of this protein is not well understood, it contains a coiled-coil domain and a RINT1_TIP1 motif located within the coiled-coil domain.
Locus

CCDC142 is found on the – strand of chromosome 2 (2p13.1), with the genomic sequence spanning bases 74,472,832 to 74,483,230. The coding region is 8292 base pairs long, encoding for two protein isoforms 743 to 665 amino acids in length. On the telomeric side, CCDC142 is followed by the MOGS and MRPL53 genes. On the centromeric side, it is followed by the C31, LBX2, LBX2-AS1, and PCGF1 genes.
mRNA
In Homo sapiens, the CCDC142 gene encodes for two alternatively spliced isoforms of the mRNA, called isoform 1 and isoform 2. Both of these isoforms have 9 exons. Isoform 1 is the longer of the two, being 4339bp long, while isoform 2 is 2253bp long. The main difference between the isoforms that isoform 2 has a shorter exon 9 and 3' UTR. Isoform 1 is the longest variant of the gene and protein and is the subject of this article.
Conservation
Paralogs
CCDC142 has no paralogs in Homo sapiens.
Orthologs
Below is a table of a variety of orthologs of CCDC142 whose protein sequence identity was compared to the Homo sapiens protein amino acid sequence. CCDC142 has more than 73% amino acid similarity in mammals, but is less conserved in other vertebrates and in invertebrates.
| Genus and Species | Common Name | Date of Divergence from Human Lineage (MYA) | % identity |
|---|---|---|---|
| Homo sapiens | Human | 0 | 100 |
| Pan troglodytes | Chimpanzee | 6.6 | 96 |
| Gorilla gorilla gorilla | Gorilla | 8.9 | 98 |
| Jaculus jaculus | Lesser Egyptian jerboa | 90.9 | 73 |
| Bos mutus | Yak | 97.5 | 74 |
| Eptesicus fuscus | Big brown bat | 97.5 | 74 |
| Python bivittatus | Burmese python | 320.5 | 36 |
| Gallus gallus | Chicken | 320.5 | 35 |
| Haliaeetus leucocephalus | Bald eagle | 320.5 | 33 |
| Anolis carolinensis | Carolina anole (lizard) | 320.5 | 33 |
| Calidris pugnax | Ruff (bird) | 320.5 | 32 |
| Xenopus tropicalis | Western clawed frog | 355.7 | 33 |
| Callorhinchus milii | Australian ghostshark | 429.6 | 36 |
| Lepisosteus oculatus | Spotted gar | 429.6 | 34 |
| Esox lucius | Northern pike | 429.6 | 33 |
| Danio rerio | Zebrafish | 429.6 | 33 |
| Lingula anatina | Tailed mussel | 847 | 29 |
| Crassostrea gigas | Pacific oyster | 847 | 29 |
| Octopus bimaculoides | California two-spot octopus | 847 | 27 |
| Drosophila melanogaster | Fruit fly | 847 | 23 |
Phylogeny
CCDC142 is closely related in mammals, mollusks and amphibians, reptiles and birds, and in fish. The CCDC142 gene goes as far back as Drosophila melanogaster, which split from the human lineage 847 million years ago. CCDC142 has mutated at a greater rate than both Cytochrome C (a highly conserved protein) and Fibrinogen A (a rapidly mutating protein). This indicates that CCDC142 is a rapidly mutating gene with an increasing rate of mutation (that is, evolution) over time.

Protein

Primary Structure, Variants, and Isoforms
The main isoform of the CCDC142 protein is 743 amino acids in length and the second isoform is 665 amino acids long. The difference in length is made entirely by amino acids missing from the C-terminus of isoform 2.
Domains and Motifs
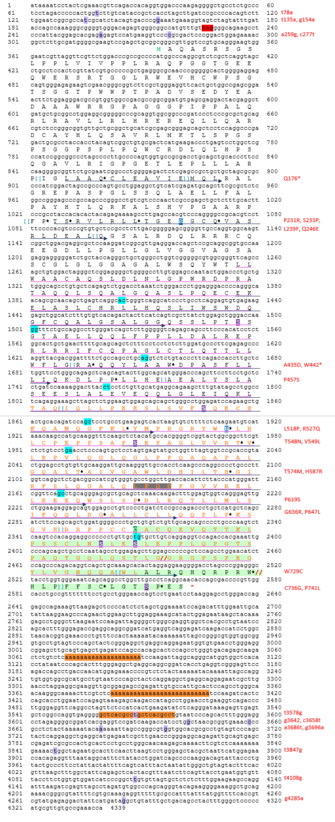 CCDC142 Conceptual Translation
CCDC142 Conceptual Translation CCDC142 Conceptual Translation Legend
CCDC142 Conceptual Translation Legend
The predicted coiled-coil domain of CCDC142 is from amino acids 308–719. A RINT1_TIP1 motif is also present from amino acids 490–621. RINT1_TIP1 is a family that includes RINT-1 (a protein involved in radiation-induced check point control) and TIP-1 (a yeast protein which is involved in Golgi transport). The extra ~250 amino acids found in the distant ortholog CCDC142 proteins are not found in the Homo sapiens genome the near CCDC142 gene.
Post-Translational Modifications
CCDC142 is predicted to have 6 phosphorylation sites, 4 methylation sites, 1 palmitoylation site, 1 sumoylation site, and 1 weak Nuclear Localization Signal. These modifications indicate that CCDC142 is localized to the nucleus and cytosol. Refer to the Conceptual Translation for annotations of these sites in the protein.
Structure prediction
Secondary structure of CCDC142 contains only α-helices as predicted by the Quick2D and Phyre2 programs . It is predicted that CCDC142 contains eight conserved α-helices, with six located in the coiled-coil region of the protein. The predicted tertiary structure of CCDC142 contains a large coiled-coil domain from amino acids 308–719.

Expression
Promoters and Regulatory Factors
The promoter region for CCDC142 was identified using the El Dorado program at Genomatix, it spans bases 74482896–74483908 in chromosome 2. This 1013bp region spans 1071–58bp upstream of the start codon of CCDC142. There is a region in the promoter which binds a large number of Krueppel-like transcription factors and BED zinc-finger proteins. This region has no single-nucleotide polymorphisms (SNPs) located in it. Many of the transcription factors that bind to the promoter region of CCDC142 have functions dealing with tumor suppression, neurogenesis, DNA damage, and photoreception. This promoter region also contains a mammalian C-type LTR TATA box which overlaps with the transcription start site of the gene.
RNA Binding Proteins
A number of possible RNA binding proteins bind to both the 3’ and 5’ untranslated regions (UTRs) of the CCDC142 mRNA. The PABPC1 and RBMX protein binding sites occur in high frequency in the 3’ UTR, with 49 and 21 sites respectively.
Expression
- Allen Human Brain Atlas Expression of CCDC142
-
 Lateral View
Lateral View
Red=Low Expression -
 Frontal View
Frontal View
Green=High Expression
Above are the Allen Human Brain Atlas expression data on CCDC142, with red indicating lower expression and green indicating higher expression. In the Homo sapiens brain, it was found that CCDC142 is lowly expressed in the cerebellar cortex, thalamus and hypothalamus. CCDC142 is also highly expressed in the substantia nigra, pons, claustrum, and mesencephalon. There is also relatively higher expression of CCDC142 in the mouth and thymus.
- NCBI GEO Expression Data
-
 MEKK 2/3 Knockout Experiment
MEKK 2/3 Knockout Experiment
-
 Myocardial Damage Experiment
Myocardial Damage Experiment
-
 SNAI Overexpression Experiment
SNAI Overexpression Experiment
The above experimental expression data shows many possible findings for CCDC142. Overexpression of SNAI1, a zinc finger protein, is correlated to the reduction of CCDC142 expression in Homo sapiens. A Mus musculus knockout of MEKK 2/3, which help regulate helper T cell differentiation, also showed lowered expression of CCDC142. Another Mus musculus experiment focusing on cardiomyopathy in mice showed lower levels of CCDC142 in mice with damaged myocardial cells.
Function and Biochemistry
Composition
CCDC142 has a relatively typical distribution of amino acids compared to other Homo sapiens proteins. However, some variations are noted across orthologs. Leucine is present in large amounts relative to other proteins (at over 15% of the protein) and asparagine is present in low amounts relative to other proteins (at less than 0.7% of the protein).
The coiled-coil domain and RINT1_TP1 motif of CCDC142 contain higher amounts of leucine relative to the rest of the protein (at over 16.6% of the region), higher amounts of glutamine (at over 8.4% of the region), and similarly low amounts of asparagine (at less than 0.7% of the region).
Interacting Proteins
No protein interactions have been found for CCDC142.
Clinical Significance
Pathology and Diseases
Copy number gain in the CCDC142 loci, including 25 other genes, showed a phenotype of developmental delay and significant developmental or morphological phenotypes. One result with a copy number loss in the CCDC142 loci, including 29 other genes, showed phenotypes of short stature, abnormal face shape, delayed speech and language development, overlapping toe, intrauterine growth retardation, patent ductus arteriosus, and delayed gross motor development. However, the effect of CCDC142 may have been confounded for these phenotypes since there were also abnormalities in many other genomic sections.
Mutations
There are a number of SNPs located in the CCDC142 gene. Some of these in the promoter region and 5’ UTR are within anchor sequences for transcription factors, and affect transcription factor binding if they are changed.
There are many SNPs in the protein's coding sequence which change CCDC142's amino acid composition. One SNP with a high prevalence rate in the population (1.8%) is notable for its change in chemistry, with a tyrosine to an asparagine shift at amino acid 548.
There are also numerous SNPs located in the large 3’ UTR of the gene, with many of these binding to areas containing stem loop structures in the mRNA. An SNP with a 7.7% prevalence rate (guanine to adenosine at bp4285) is in the 3’ UTR but not located in the conserved stem loop region.
These SNPs have been annotated in the Conceptual Translation located in the Protein section above.
Multiple Sequence Alignment
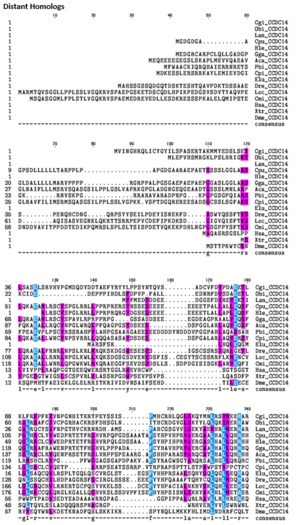
- Distant Ortholog Multiple Sequence Alignment of CCDC142
-

-

-
 Purple=Similar Amino Acid Chemistry
Purple=Similar Amino Acid Chemistry
Blue=Same Amino Acid -
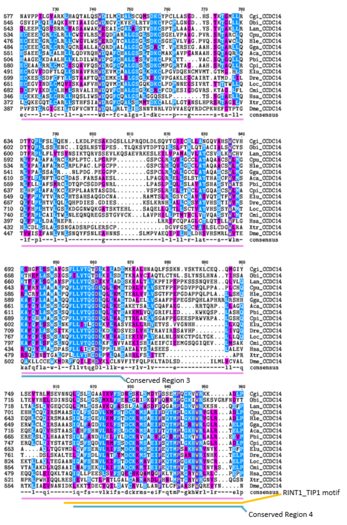
-

In the Multiple Sequence Alignment above (created using the CLUSTALW and TEXSHADE programs at SDSC Biology Workbench), organisms are labeled by the first letter of their genus and the first two letters of their species. The whole CCDC142 protein is highly conserved in mammals. The regions containing the Homo sapiens coiled-coil domain and the RINT1_TIP1 motif region are highly conserved in distant homologs. 12 of the 15 amino acids that match across all organisms in this region are nonpolar. Conserved Region 1 contains mostly nonpolar amino acids. Conserved Region 2 contains mostly nonpolar and basic amino acids. Conserved Region 3 contains both polar and nonpolar amino acids. Conserved Region 5 contains mostly nonpolar and basic amino acids.
Additional Transcription Factor Information
 Transcription Factor Binding Site Annotation
Transcription Factor Binding Site Annotation Transcription Factor Binding Site Legend
Transcription Factor Binding Site Legend
References
- ^ "CCDC142 coiled-coil domain containing 142 [Homo sapiens (human)] – Gene – NCBI". www.ncbi.nlm.nih.gov. Retrieved 2016-05-01.
- ^ "coiled-coil domain-containing protein 142 [Homo sapiens] – Protein – NCBI". www.ncbi.nlm.nih.gov. Retrieved 2016-05-01.
- ^ "CCDC142 – Coiled-coil domain-containing protein 142 – Homo sapiens (Human) – CCDC142 gene & protein". www.uniprot.org. Retrieved 2016-05-01.
- ^ "SSDB Motif Search Result: hsa:84865". www.kegg.jp. Retrieved 2016-05-01.
- ^ "SDSC Biology Workbench".
- "NetPhos 2.0 Server". www.cbs.dtu.dk. Retrieved 2016-05-01.
- "Memo:Protein Methylation Prediction". www.bioinfo.tsinghua.edu.cn. Archived from the original on 2016-03-14. Retrieved 2016-05-01.
- ":::NBA-Palm – Prediction of Palmitoylation Site Implemented In Naive Bayesian Algorithm:::". www.bioinfo.tsinghua.edu.cn. Archived from the original on 2016-06-09. Retrieved 2016-05-01.
- "SUMOplot™ Analysis Program | Abgent". www.abgent.com. Retrieved 2016-05-01.
- "NLS_Mapper". nls-mapper.iab.keio.ac.jp. Archived from the original on 2021-11-22. Retrieved 2016-05-01.
- ^ Kelley, Lawrence. "PHYRE2 Protein Fold Recognition Server". www.sbg.bio.ic.ac.uk. Retrieved 2016-05-01.
- ^ Remmert, Michael. "Quick2D". toolkit.tuebingen.mpg.de. Retrieved 2016-05-01.
- ^ "I-TASSER server for protein structure and function prediction". zhanglab.ccmb.med.umich.edu. Retrieved 2016-05-01.
- ^ "Genomatix – NGS Data Analysis & Personalized Medicine". www.genomatix.de. Archived from the original on 2001-02-24. Retrieved 2016-05-01.
- ^ snpdev. "SNP linked to Gene (geneID:84865) Via Contig Annotation". www.ncbi.nlm.nih.gov. Retrieved 2016-05-01.
- "RBPDB: The database of RNA-binding specificities". rbpdb.ccbr.utoronto.ca. Retrieved 2016-05-01.
- ^ "Microarray Data :: Allen Brain Atlas: Human Brain". human.brain-map.org. Retrieved 2016-05-01.
- "EST Profile – Hs.430199". www.ncbi.nlm.nih.gov. Retrieved 2016-05-01.
- geo. "Home – GEO – NCBI". www.ncbi.nlm.nih.gov. Retrieved 2016-05-01.
- ^ "GDS3596 / 1451178_at". www.ncbi.nlm.nih.gov. Retrieved 2016-05-01.
- "GDS4795 / ILMN_3023885". www.ncbi.nlm.nih.gov. Retrieved 2016-05-01.
- ^ ClinVar. "No items found – ClinVar – NCBI". www.ncbi.nlm.nih.gov. Retrieved 2016-05-05.
Further reading
- Orekhova AS, Sverdlova PS, Spirin PV, Leonova OG, Popenko VI, Prasolov VS, Rubtsov PM (2011-06-01). "". Molekuliarnaia Biologiia. 45 (3): 486–495. PMID 21790010.