| This article needs additional citations for verification. Please help improve this article by adding citations to reliable sources. Unsourced material may be challenged and removed. Find sources: "Inosinic acid" – news · newspapers · books · scholar · JSTOR (November 2014) (Learn how and when to remove this message) |

| |
| |
| Names | |
|---|---|
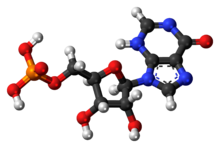
| IUPAC name 5'-Inosinic acid | |
Other names
| |
| Identifiers | |
| CAS Number | |
| 3D model (JSmol) | |
| ChEBI | |
| ChEMBL | |
| ChemSpider | |
| ECHA InfoCard | 100.004.588 |
| E number | E630 (flavour enhancer) |
| IUPHAR/BPS | |
| MeSH | Inosine+monophosphate |
| PubChem CID | |
| UNII | |
| CompTox Dashboard (EPA) | |
InChI
| |
SMILES
| |
| Properties | |
| Chemical formula | C10H13N4O8P |
| Molar mass | 348.208 g·mol |
| Except where otherwise noted, data are given for materials in their standard state (at 25 °C , 100 kPa).
| |
Inosinic acid or inosine monophosphate (IMP) is a nucleotide (that is, a nucleoside monophosphate). Widely used as a flavor enhancer, it is typically obtained from chicken byproducts or other meat industry waste. Inosinic acid is important in metabolism. It is the ribonucleotide of hypoxanthine and the first nucleotide formed during the synthesis of purine nucleotides. It can also be formed by the deamination of adenosine monophosphate by AMP deaminase. It can be hydrolysed to inosine.
The enzyme deoxyribonucleoside triphosphate pyrophosphohydrolase, encoded by YJR069C in Saccharomyces cerevisiae and containing (d)ITPase and (d)XTPase activities, hydrolyzes inosine triphosphate (ITP) releasing pyrophosphate and IMP.
Important derivatives of inosinic acid include the purine nucleotides found in nucleic acids and adenosine triphosphate, which is used to store chemical energy in muscle and other tissues.
In the food industry, inosinic acid and its salts such as disodium inosinate are used as flavor enhancers. It is known as E number reference E630.
Inosinate synthesis
Main article: Purine metabolism § BiosynthesisThe inosinate synthesis is complex, beginning with a 5-phosphoribosyl-1-pyrophosphate (PRPP). Enzymes taking part in IMP synthesis constitute a multienzyme complex in the cell. Evidence demonstrates that there are multifunctional enzymes, and some of them catalyze non-sequential steps in the pathway.

Synthesis of other purine nucleotides
Within a few steps inosinate becomes AMP or GMP. Both compounds are RNA nucleotides. AMP differs from inosinate by the replacement of IMP's carbon-6 carbonyl with an amino group. The interconversion of AMP and IMP occurs as part of the purine nucleotide cycle. GMP is formed by the inosinate oxidation to xanthylate (XMP), and afterwards adds an amino group on carbon 2. Hydrogen acceptor on inosinate oxidation is NAD+. Finally, carbon 2 gains the amino group by spending an ATP molecule (which becomes AMP+2Pi). While AMP synthesis requires GTP, GMP synthesis uses ATP. That difference offers an important regulation possibility.

Regulation of purine nucleotide biosynthesis
Inosinate and many other molecules inhibit the synthesis of 5-phosphoribosylamine from 5-phosphoribosyl-1-pyrophosphate (PRPP), disabling the enzyme that catalyzes the reaction: glutamine-5-phosphoribosyl-1-pyrophosphate-amidotransferase. In other words, when levels of inosinate are high, glutamine-5-phosphoribosyl-1-pyrophosphate-amidotransferase is inhibited, and, as a consequence, inosinate levels decrease. Also, as a result, adenylate and guanylate are not produced, which means that RNA synthesis cannot be completed because of the lack of these two important RNA nucleotides.
Applications
Inosinic acid can be converted into various salts including disodium inosinate (E631), dipotassium inosinate (E632), and calcium inosinate (E633). These three compounds are used as flavor enhancers for the basic taste umami or savoriness with a comparatively high effectiveness. They are mostly used in soups, sauces, and seasonings for the intensification and balance of the flavor of meat.
See also
- Thioinosinic acid
- Phosphatidyl inositol
- Glutamate flavoring
- Kikunae Ikeda
- Umami
- Ajinomoto
- Tien Chu Ve-Tsin
- Glutamic acid
- Disodium glutamate
- Monopotassium glutamate
- Disodium inosinate
- Guanosine monophosphate
- Adenosine monophosphate
References
- Davies O, Mendes P, Smallbone K, Malys N (2012). "Characterisation of multiple substrate-specific (d)ITP/(d)XTPase and modelling of deaminated purine nucleotide metabolism". BMB Reports. 45 (4): 259–64. doi:10.5483/BMBRep.2012.45.4.259. PMID 22531138.
- ^ Mader, M. M.; Henry, J. R. (2007-01-01), Taylor, John B.; Triggle, David J. (eds.), "7.03 - Antimetabolites", Comprehensive Medicinal Chemistry II, Oxford: Elsevier, pp. 55–79, doi:10.1016/b0-08-045044-x/00204-2, ISBN 978-0-08-045044-5, retrieved 2020-12-17
- Voet, D, Voet, J. G., Biochemistry (3rd Edition), John Wiley & Sons, Inc., 2004, pg 1095
Further reading
- Berg, Jeremy M.; Bioquímica; Editorial Reverté; 6ena edició; Barcelona 2007.
- Nelson, David L.; Principles of biochemistry; Editorial W.H Freeman and Company; 4th edition; New York 2005.
| Nucleic acid constituents | |||||||
|---|---|---|---|---|---|---|---|
| Nucleobase | |||||||
| Nucleoside |
| ||||||
| Nucleotide (Nucleoside monophosphate) |
| ||||||
| Nucleoside diphosphate | |||||||
| Nucleoside triphosphate | |||||||
| Nucleotide metabolic intermediates | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| purine metabolism |
| ||||||||||
| pyrimidine metabolism |
| ||||||||||